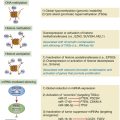
AML, Acute myeloid leukemia; BL, Burkitt’s lymphoma; CLL, chronic lymphocytic leukemia; DLBCL, diffuse large B-cell lymphoma; ES, Ewing sarcoma.
Main Types of Therapeutic RNA Molecules
Table 55-2
Glossary of Terms in RNA-Inhibition Strategies

Ribozymes
siRNAs and shRNAs
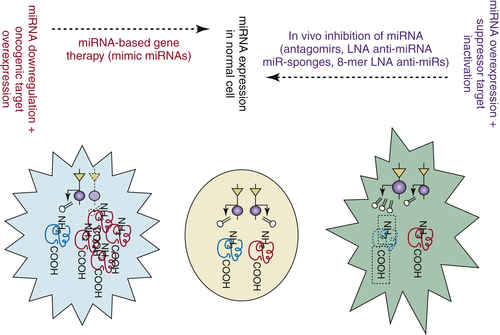
ASOs/AMOs Anti-miRNAs, LNAS Anti-miRNAs, and Antagomirs
miRNA-Mimic Agents
In Search of the Right Way and the Right Type of Delivery
A Strategy for Using RNA as Therapeutic Molecules
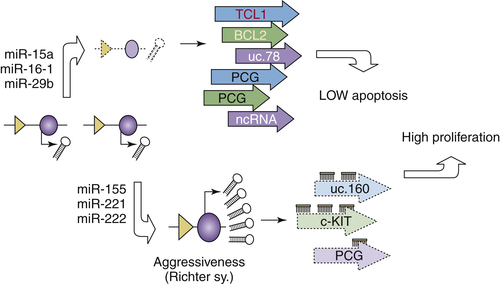
Table 55-3
Delivery Methods for RNA-Interference–Based Therapeutics
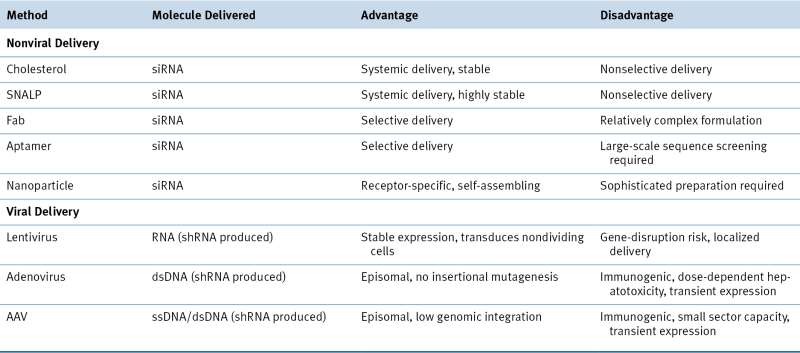
dsDNA, Double-strand DNA; SNALP, stable nucleic acid-lipid particle; ssDNA, single-strand DNA.
Modified with permission from Kim DH, Rossi JJ. Strategies for silencing human disease using RNA interference. Nat Rev Genet. 2007;8:173-184.
Acknowledgments
1. Cancer statistics . CA Cancer J Clin . 2012 ; 62 : 10 – 29 .
2. Antisense therapy for cancer . Nat Rev Cancer . 2005 ; 5 : 468 – 479 .
3. Phase I to II multicenter study of oblimersen sodium, a Bcl.2 antisense oligonucleotide, in patients with advanced chronic lymphocytic leukemia . J Clin Oncol . 2005 ; 23 : 7697 – 7702 .
4. Randomized phase III trial of fludarabine plus cyclophosphamide with or without oblimersen sodium (Bcl-2 antisense) in patients with relapsed or refractory chronic lymphocytic leukemia . J Clin Oncol . 2007 ; 25 : 1114 – 1120 .
5. Randomized phase II evaluation of aprinocarsen in combination with gemcitabine and cisplatin for patients with advanced/metastatic non small cell lung cancer . Invest New Drugs . 2005 ; 23 : 263 – 269 .
6. Phase II study of PKC alpha antisense oligonucleotide aprinocarsen in combination with gemcitabine and carboplatin in patients with advanced non small cell lung cancer . Lung Cancer . 2006 ; 52 : 173 – 180 .
7. Phase III study of gemcitabine and cisplatin with or without aprinocarsen, a protein kinase C-alpha antisense oligonucleotide, in patients with advanced-stage non-small-cell lung cancer . J Clin Oncol . 2006 ; 24 : 1428 – 1434 .
8. A phase II trial of ISIS 3521 in patients with metastatic colorectal cancer . Clin Colorectal Cancer . 2004 ; 4 : 268 – 274 .
9. Phase II study of ISIS 3521, an antisense oligodeoxynucleotide to protein kinase C alpha, in patients with previously treated low grade non Hodgkin’s lymphoma . Ann Oncol . 2004 ; 15 : 1413 – 1418 .
10. A phase I/II study of LY900003, an antisense inhibitor of protein kinase C alpha, in combination with cisplatin and gemcitabine in patients with advanced non small cell lung cancer . Clin Cancer Res . 2004 ; 10 : 6086 – 6093 .
11. MicroRNAs in stress signaling and human disease . Cell . 2012 ; 148 : 1172 – 1187 .
12. Oncomirs—microRNAs with a role in cancer . Nat Rev Cancer . 2006 ; 6 : 259 – 269 .
13. MicroRNA signatures in human cancers . Nat Rev Cancer . 2006 ; 6 : 857 – 866 .
14. MicroRNA pathways in flies and worms: growth, death, fat, stress, and timing . Cell . 2003 ; 113 : 673 – 676 .
15. MicroRNAs: genomics, biogenesis, mechanism, and function . Cell . 2004 ; 116 : 281 – 297 .
16. MicroRNAs: small RNAs with a big role in gene regulation . Nat Rev Genet . 2004 ; 5 : 522 – 531 .
17. Transcription and processing of human microRNA precursors . Mol Cell . 2004 ; 16 : 861 – 865 .
18. MicroRNA cancer connection: the beginning of a new tale . Cancer Res . 2006 ; 66 : 7390 – 7394 .
19. An oligonucleotide microchip for genome-wide microRNA profiling in human and mouse tissues . Proc Natl Acad Sci U S A . 2004 ; 101 : 9740 – 9744 .
20. microRNA detection comes of age . Nat Methods . 2006 ; 3 : 12 – 13 .
21. A microRNA signature associated with prognosis and progression in chronic lymphocytic leukemia . N Engl J Med . 2005 ; 353 : 1793 – 1801 .
22. The role of microRNA genes in papillary thyroid carcinoma . Proc Natl Acad Sci U S A . 2005 ; 102 : 19075 – 19080 .
23. Long non-coding RNAs and cancer: a new frontier of translational research? Oncogene . 2012 ; 31 : 4577 – 4587 .
24. Conserved non-genic sequences: an unexpected feature of mammalian genomes . Nat Rev Genet . 2005 ; 6 : 151 – 157 .
25. Ultraconserved elements in the human genome . Science . 2004 ; 304 : 1321 – 1325 .
26. Ultraconserved regions encoding ncRNAs are altered in human leukemias and carcinomas . Cancer Cell . 2007 ; 12 : 215 – 229 .
27. Nucleic-acid therapeutics: basic principles and recent applications . Nat Rev Drug Discov . 2002 ; 1 : 503 – 514 .
28. Self-splicing RNA: autoexcision and autocyclization of the ribosomal RNA intervening sequence of . Tetrahymena. Cell . 1982 ; 31 : 147 – 157 .
29. Nucleotide sequence of the gene encoding the RNA subunit (M1 RNA) of ribonuclease P from . Escherichia coli. Cell . 1982 ; 30 : 627 – 636 .
30. Ribozyme catalysis: not different, just worse . Nat Struct Mol Biol . 2005 ; 12 : 395 – 402 .
31. Destroying RNA as a therapeutic approach . Curr Med Chem . 2006 ; 13 : 863 – 881 .
32. A phase I clinical trial of a ribozyme-based angiogenesis inhibitor targeting vascular endothelial growth factor receptor-1 for patients with refractory solid tumors . Mol Cancer Ther . 2006 ; 4 : 948 – 955 .
33. Safety and pharmacokinetic study of RPI.4610 (ANGIOZYME), an anti-VEGFR-1 ribozyme, in combination with carboplatin and paclitaxel in patients with advanced solid tumors . Cancer Chemother Pharmacol . 2005 ; 56 : 329 – 336 .
34. Pharmacokinetics and tolerability of an antiangiogenic ribozyme (ANGIOZYME) in healthy volunteers . J Clin Pharmacol . 2000 ; 40 ( Pt 2 ) : 1462 – 1469 .
35. Vascular and haematopoietic stem cells: novel targets for anti-angiogenesis therapy? Nat Rev Cancer . 2002 ; 2 : 826 – 835 .
36. Construction of an allosteric trans-maxizyme targeting for two distinct oncogenes . Nucleic Acids Res suppl . 2002 ; 20 : 115 – 116 .
37. Potent and specific genetic interference by double-stranded RNA in . Caenorhabditis elegans. Nature . 1998 ; 391 : 806 – 811 .
38. Strategies for silencing human disease using RNA interference . Nat Rev Genet . 2007 ; 8 : 173 – 184 .
39. Gene silencing in mammals by small interfering RNAs . Nat Rev Genet . 2002 ; 3 : 737 – 747 .
40. shRNA libraries and their use in cancer genetics . Nat Methods . 2006 ; 3 : 701 – 706 .
41. Overcoming the classical multidrug resistance phenotype by adenoviral delivery of anti-MDR1 short hairpin RNAs and ribozymes . Int J Oncol . 2007 ; 31 : 419 – 430 .
42. Intraperitoneal delivery of liposomal siRNA for therapy of advanced ovarian cancer . Cancer Biol Ther . 2006 ; 5 : 1708 – 1713 .
43. Knockdown of mutant K-ras expression by adenovirus-mediated siRNA inhibits the in vitro and in vivo growth of lung cancer cells . Cancer Biol Ther . 2006 ; 5 : 1481 – 1486 .
44. , et al. RNAi therapeutics: a potential new class of pharmaceutical drugs . Nat Chem Biol . 2006 ; 2 : 711 – 719 .
45. miRNAs and their potential for use against cancer and other diseases . Future Oncol . 2007 ; 3 : 521 – 534 .
46. Chromosomal rearrangements and microRNAs: a new cancer link with clinical implications . J Clin Invest . 2007 ; 117 : 2059 – 2066 .
47. miR-122 regulation of lipid metabolism revealed by in vivo antisense targeting . Cell Metab . 2006 ; 3 : 87 – 98 .
48. MicroRNA-21 is an antiapoptotic factor in human glioblastoma cells . Cancer Res . 2005 ; 65 : 6029 – 6033 .
49. Silencing of microRNAs in vivo with “antagomirs.” . Nature . 2005 ; 438 : 685 – 689 .
50. Specificity, duplex degradation and subcellular localization of antagomirs . Nucleic Acids Res . 2007 ; 35 : 2885 – 2892 .
51. LNA-mediated anti-microRNA-155 silencing in low-grade B cell lymphomas . Blood . 2012 ; 120 : 1678 – 1686 .
52. MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells . Nat Methods . 2007 ; 4 : 721 – 726 .
53. Novel approaches for gene-specific interference via manipulating actions of microRNAs: examination on the pacemaker channel genes HCN2 and HCN4 . J Cell Physiol . 2007 ; 212 : 285 – 292 .
54. The let-7 microRNA reduces tumor growth in mouse models of lung cancer . Cell Cycle . 2008 ; 7 : 759 – 764 .
55. Effects of miR-34a on cell growth and chemoresistance in prostate cancer PC3 cells . Biochem Biophys Res Commun . 2008 ; 377 : 114 – 119 .
56. Therapeutic microRNA delivery suppresses tumorigenesis in a murine liver cancer model . Cell . 2009 ; 137 : 1005 – 1017 .
57. Fatality in mice due to oversaturation of cellular microRNA/short hairpin RNA pathways . Nature . 2006 ; 441 : 537 – 541 .
58. Nonviral delivery vehicles for use in short hairpin RNA-based cancer therapies . Expert Rev Anticancer Ther . 2007 ; 7 : 373 – 382 .
59. Gene silencing through RNA interference (RNAi) in vivo: strategies based on the direct application of siRNAs . J Biotechnol . 2006 ; 124 : 12 – 25 .
60. Chronic lymphocytic leukemia . N Engl J Med . 2005 ; 352 : 804 – 815 .
61. miR-15 and miR-16 induce apoptosis by targeting BCL2 . Proc Natl Acad Sci U S A . 2005 ; 102 : 13944 – 13949 .
62. Tcl1 expression in CLL is regulated by miR-29 and miR-181 . Cancer Res . 2006 ; 66 : 11590 – 11593 .
63. A ceRNA hypothesis: the Rosetta Stone of a hidden RNA language? Cell . 2011 ; 146 : 353 – 358 .
64. Noncoding RNA gas5 is a growth arrest- and starvation-associated repressor of the glucocorticoid receptor . Sci Signal . 2010 ; 3 : ra8 .