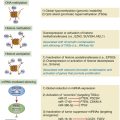
Figure 59-1 Immunohistochemistry (A) (1) An antibody specific for a particular protein, phosphate, sulfate, carbohydrate, or other moiety is allowed to bind to the tissue section. (2) A secondary “anti-antibody” is added, bearing numerous biotin tags. (3) A complex of avidin and horseradish peroxidase (HRP) is added. Avidin binds tightly to the biotin on the secondary antibody, while HRP reacts with a chromogen in the presence of H2O2, producing a visible precipitate on the tissue. (B) Microscopic images of a gastrointestinal stromal tumor. The brown staining in the right image represents immunohistochemical detection of the KIT tyrosine kinase (CD117) in the tumor.
Although immunohistochemistry is a mainstay in modern pathology, it does have limitations. Not all antibodies are perfectly specific; background issues and cross reactivity can affect interpretation. Further, the technique is at best only semiquantitative and can be influenced by the length of formaldehyde fixation and many other parameters. Some of these limitations can be mitigated by automation of the staining, but tumor necrosis, intraoperative ischemia, and variation in specimen processing are all factors that can complicate the reading of immunohistochemical stains.
Immunofluorescence
Because formaldehyde fixation greatly distorts or even destroys many epitopes in tissue samples, there are numerous antibodies that will not work on FFPE sections. Such antibodies may still be useful, however, on cryostat sections, which can be prepared if fresh-frozen tissue is available. In these circumstances, fluorescence rather than histochemistry is the preferred method of detection, as it has greater sensitivity and allows for several antibodies to be stained at once (using different fluorophores).
9 Immunofluorescence is routinely used in evaluating medical diseases of the kidney and skin, but is rarely employed in cancer diagnosis.
In Situ Hybridization
Expression of particular mRNAs in tumor tissue can be measured by in situ hybridization (ISH).
10 In this approach, complementary oligonucleotide probes (either DNA or RNA) bearing a chemical tag such as digoxigenin are allowed to hybridize to their target mRNA. Once hybridized, the tag is detected with a secondary antibody (e.g., anti-digoxigenin), which is linked to an enzyme that produces a visible chemical deposit on the cells. Horseradish peroxidase and alkaline phosphatase of the two most commonly used enzymes in this powerful approach, which is widely employed in cancer research. However, ISH is technically challenging and works best on mRNAs that are expressed in high abundance. For these reasons, the use of ISH in clinical laboratories is generally restricted to the assessment of kappa and lambda light-chain expression in cases of suspected myeloma, or to detect the presence of Epstein-Barr virus in cases of lymphoproliferative disease driven by this infection.
Fluorescence in Situ Hybridization (FISH)
Nucleic acid probes are used not only to detect mRNA, but to assess interphase chromosomes in tumor cells. The probes may consist of either DNA or RNA, and they vary in length from short oligonucleotides to multigenic chromosomal segments cloned into bacteria (so-called bacterial artificial chromosomes, or BACs). When directly labeled with a fluorophore, they can be detected using a microscope equipped for immunofluorescence. This approach is routinely used in clinical cytogenetic laboratories to assess gene copy number as well as gene translocations. For example, FISH is commonly used to evaluate breast carcinomas for amplification of the ERBB2 (HER2) locus and to screen for increased copies of NMYC in neuroblastoma and MET in non–small-cell lung carcinoma. Loss of tumor suppressors such as PTEN, CDKN2A, and TP53 can also be assessed by this approach.
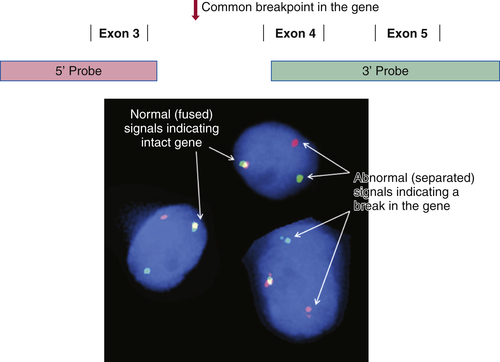
There is a long and growing list of gene translocation events that are linked to cancer. Whether the result of intra- or interchromosomal exchanges, these translocations commonly involve genes encoding a kinase or a transcription factor. The resulting fusion genes are often the principal drivers of tumorigenesis and therefore serve as diagnostic markers and/or targets for specific therapies (e.g., kinase inhibitors). Fusion mRNAs from translocation events can be detected by highly sensitive methodologies based on polymerase chain reaction (PCR); however, these approaches can be frustrated by the fact that a particular target gene may be fused to any of more than a dozen different partner genes, requiring numerous primers to cover all possible fusion events. In contrast, FISH can detect a translocation-related break in a target gene irrespective of which partner gene has been fused to it. This is done by labeling two pools of probes with different fluorophores; for example, one pool may be labeled red and hybridizes to the 5′ end of the gene, while the other is labeled green and hybridizes to the 3′ end of the gene. If the gene is intact, the red and green signals are close together and merge into yellow, but if the gene has been split apart by a translocation event, the red and green signals become well separated (
Figure 59-2 ). A further advantage of FISH is that only a few tumor cells are required to make a diagnosis, which is helpful in biopsies containing low tumor content.
Preparing Nucleic Acids from Cancer Specimens
Many of the assays that are used to molecularly subtype cancers require the extraction of either RNA or DNA from the tumor tissue. Two important issues are often overlooked in this regard. The first is that the nature of the tissue dictates the quality of
the available nucleic acid. Whereas fresh-frozen tissue will yield RNA or DNA that is largely intact, the nucleic acids derived from FFPE tissue are highly degraded. Thus, assays meant for use on FFPE-derived RNA or DNA must take into account the short, fragmented nature of these derivatives.
Figure 59-2 Fluorescence in situ hybridization (FISH) for identifying gene translocation events This approach is commonly used to look for evidence of gene translocations in interphase nuclei. Two probes are designed to hybridize to the 5′ and 3′ ends of a gene, flanking the region that is commonly “broken” during translocation. The probes are labeled with fluorescent tags of differing color, often creating a third color when they bind subjacently (e.g., green and red fusing into yellow). Wide separation of the probes indicates a break in the gene consistent with a translocation event. Note that this approach does not identify the partner gene in the translocation.
The second important issue is that tumor content and quality are highly variable from one specimen to the next, and even among different parts of the same specimen. For example, one paraffin block from a cancer resection specimen may consist of 90% tumor cells, while an adjacent block is dominated by stromal and inflammatory cells, with only 10% tumor cells being present. A third block from the same specimen may contain only necrotic tumor, as geographic necrosis is common at the center of large, high-grade malignancies. Thus, it is critical that all tumor material being considered for nucleic acid extraction first be evaluated by a qualified pathologist, who can assess whether the tumor sample is suitable for the proposed testing.
Tumor Enrichment by Macrodissection
The sensitivity of molecular diagnostic assays for mutations and other alterations in tumor DNA is dependent in part on the ratio of tumor DNA to normal cellular DNA present within a specimen. If 90% of the cells within a tissue sample are cancerous, mutation detection is straightforward, but if only 10% of the cells represent the tumor, DNA from the normal cells will effectively dilute out the mutation and make it harder to detect. For this reason, it has become standard practice to dissect out the tumor-rich areas of a specimen. This can be performed under a dissection microscope, but is more commonly done by simply scraping the tumor from unstained paraffin sections using a scalpel blade, or by taking a small core directly out of the paraffin block. Comparison with an H&E-stained section serves to guide the dissection.
Tumor Enrichment by Laser Capture Microdissection
Some tumor samples have too little tumor to allow macrodissection. In these cases, a microscope equipped with a laser can be used to isolate small clusters of tumor cells, or even single cells, from adjacent normal tissue elements. Although this is labor intensive and yields relatively little material for further testing, it can be used to salvage cases when no other tumor material from a patient is available.
Assays for Single Genes or Single Mutations
Over the past two decades, a long and growing list of genes that play a significant role in cancer has been identified. For mutations in those genes that are of particular prognostic importance or serve as important predictors of therapeutic response, a variety of assays have been developed for use in clinical laboratories. Most of these assays are focused on a single mutation or a set of mutations occurring in a single gene exon. Important factors in the design of such an assay include the amount of input DNA or RNA needed, the ease and rapidity with which the assay can be completed, and its
sensitivity and specificity. Of course, assay cost is another important factor. There are many different platforms used to support these types of assays, and each represents a significant investment for the laboratory.
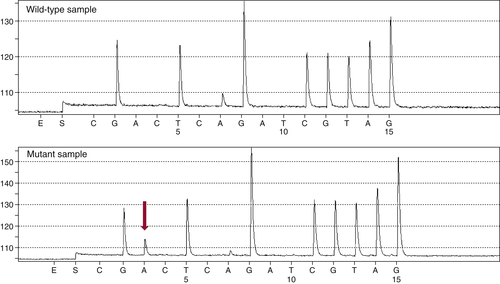
Figure 59-3 Pyrosequencing Two pyrograms are shown from tumor samples subjected to pyrosequencing for common mutations in the KRAS gene. The presence of an extra A peak in the lower panel is evidence for the presence of a mutation; the preceding G peak has a correspondingly lower signal, indicating a G→A substitution.
Sanger Sequencing
The single most common approach to screening mutations in cancer genes is Sanger sequencing. First described by Fred Sanger and colleagues in 1976, this method mixes nonextendable, fluorescently labeled dideoxy nucleotides together with standard nucleotides to generate a set of fragments of varying length as the original template DNA is copied.
11 When separated by capillary electrophoresis, the sequence of the DNA can be interpreted from the color of the last incorporated (dideoxy) nucleotide on each successive fragment. Highly reliable and reproducible, Sanger sequencing was used to generate the first complete sequence of the human genome.
12 Nevertheless, it has several significant drawbacks. First, it can only detect mutations that are present in at least 15% to 20% of the input DNA molecules. Second, it takes approximately 24 hours to perform and is relatively labor intensive. Third, it can only cover mutations across approximately 200 bp of FFPE DNA. This means that many genes require multiple, overlapping sequencing reactions in order to ensure that a mutation can be identified, adding to the overall cost. For this reason, many laboratories rely on other approaches for screening common cancer-related mutations.
Pyrosequencing
Developed as a rapid and sensitive approach to sequencing very short regions of DNA (up to approximately 30 bp), pyrosequencing is used by many laboratories to screen for common mutations in tumor oncogenes such as KRAS, EGFR, and BRAF.
13 This technology differs from Sanger sequencing in that it does not require the synthesis and electrophoretic separation of DNA fragments. Instead, the incorporation of nucleotides during DNA synthesis is measured through a secondary reaction: each pyrophosphate released during nucleotide incorporation results in the generation of a light signal by the enzyme luciferase. As dATP, dCTP, dGTP, and dTTP are sequentially added, their incorporation is detected by the luciferase reaction, allowing the DNA sequence to be interpreted (
Figure 59-3 ). The principal advantages of pyrosequencing are speed (sequencing can be completed in 2 hours) and sensitivity (as low as 5% mutant allele). A modification of the method is commonly used to assess DNA methylation.
High-Resolution Melting Curve Analysis
First developed in the late 1990s by Wittwer and colleagues, high-resolution melting curve analysis is now the backbone of a variety of assays designed to quickly and cost-effectively identify the presence of a mutation.
14 After the DNA of interest is amplified by standard PCR, the melting analysis
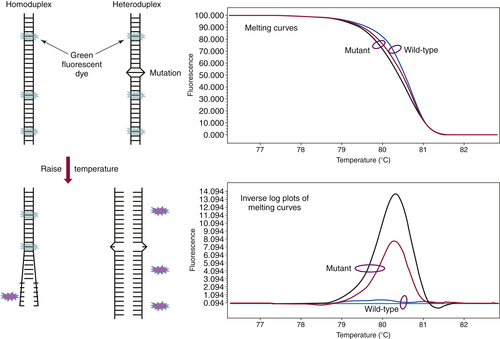
can be performed in approximately 20 minutes without the need to add reagents or switch to a different instrument. The analysis begins by melting apart the DNA strands of the final DNA product at 95° C and then allowing them to re-anneal as they cool. When a mixture of wild-type and mutant DNA is present in the sample, heteroduplexes are formed, and these can be detected by slowly ramping the temperature back up to 95° C in the presence of a fluorescent dye that binds preferentially to double-stranded DNA. As the temperature increases, the heteroduplexes unravel at a slightly lower temperature than the homoduplexes of wild-type/wild-type or mutant/mutant DNA (
Figure 59-4 ). Thus, the presence of a mutation shifts the melting point of the overall population, and this shift is readily detected by a decrease in fluorescence as the DNA strands separate.
Figure 59-4 High-resolution melting curve analysis This method takes advantage of the fact that DNA heteroduplexes formed when a mutation is present have a lower melting temperature than do homoduplexes. The resulting shift in the melting curve can be detected by loss of fluorescent signal as a dye intercalated into the double-stranded DNA is released. Plotting the inverse log of the slope of the melting curve highlights the differences between samples that are wild-type and those containing a mutation. The approach can be used to detect point mutations, insertions, and deletions within amplicons of up to several hundred base pairs.
A number of variations of this assay type have been developed. Some will allow exact identification of a specific mutation, whereas others are optimized to detect intragenic deletions or insertions, the sequence of which requires confirmatory Sanger sequencing. In general, the sensitivity of these assays is approximately 10% mutant allele.
Allele-Specific PCR
A number of approaches have been developed that promote selective PCR amplification of a mutant allele over a wild-type allele.
15 Each depends on a primer that selectively binds to a site of mutation, the specificity being determined by the degree to which the mutant allele is favored over the wild type. Modifications to the primer, such as inclusion of peptide nucleic acids or locked nucleic acids, are generally necessary to achieve the selectivity that is optimal for clinical use.
16 The best assays have detection levels below 1%, with virtually no background contributed by false priming of the wild-type allele.
Multiplexed Approaches to Cancer Genotyping
The past decade has witnessed an explosion in the number of genes identified as playing a causal role in tumor
development. This growing list of genes presents a challenge to clinical laboratories, which have traditionally used single-gene or single-exon approaches to genotyping. An alternative is to screen multiple genes/exons simultaneously using a multiplexed method, thereby saving on the labor and cost of performing multiple Sanger sequencing reactions. Two platforms have been developed to rapidly and cost-effectively screen for up to several hundred cancer gene mutations simultaneously. Both of these platforms—SnaPshot assays and mass spectroscopy–based assays—are based on so-called primer extension reactions.
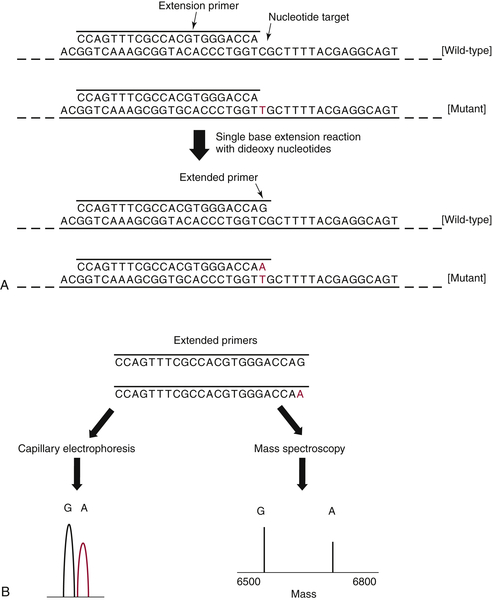
17 In brief, each gene/exon of interest is amplified by standard PCR and then allowed to anneal to an oligonucleotide primer that sits immediately adjacent to a potential site of mutation. Dideoxy nucleotides are then added, and the primer is extended by a single base (essentially a single-base sequencing reaction), following which the product is interrogated to determine whether the added base represents wild-type DNA, a mutation, or both (
Figure 59-5 ). The power of this approach is that the analysis can be performed in a multiplex manner, allowing many mutation sites to be screened at the same time.
Figure 59-5 Primer extension assays These assays are used to interrogate specific “hotspot” sites of mutation occurring in oncogenes. (A) After an initial PCR amplification of the target sequence, an extension primer is added that sits immediately adjacent to the nucleotide of interest. (B) A single base extension reaction is then performed, and the products are analyzed either by capillary electrophoresis or mass spectroscopy. Both readouts are quantitative and have similar levels of sensitivity (5% to 10% mutant allele). PCR, Polymerase chain reaction.
SNaPshot Assays
Iafrate and colleagues were among the first to adapt this technology for screening oncogenic mutations in FFPE tumor DNA.
17 The initial PCR and subsequent primer extension reactions are multiplexed to the level of 5 to 10 reactions per tube; extended primers are then separated by capillary
electrophoresis and visualized via fluorescent tags present on the incorporated dideoxy nucleotides. Primers interrogating approximately 10 different mutation sites can be analyzed simultaneously on a single capillary. By running several multiplexes, more than 100 mutations can be screened across one or two dozen genes in approximately 24 hours. SNaPshot assays have excellent sensitivity (5% mutant allele) and are cost effective, making them popular in clinical diagnostic laboratories.
Mass Spectroscopy–Based Assays
These assays follow the same approach as SNaPshot assays, multiplexing PCR and primer extension reactions to the level of 10 to 30 per tube, but the readout on the extended primers is performed by mass spectroscopy instead of capillary electrophoresis. Although the sensitivity of the mass spectrometer is a bit lower (10% mutant allele), the throughput is significantly greater, as the instrument can read out 384 multiplexes (more than 4000 mutation sites) in 90 minutes.
18,19
Next-Generation DNA Sequencing
The development of so-called next-generation sequencing has transformed the study of cancer by making it possible, and affordable, to generate large amounts of sequencing data in a reasonably short period of time. There are a number of different “next-gen” platforms in use, but they all share a common approach, which is to generate sequence from single molecules of genomic DNA or cDNA in a massively parallel fashion—hence the alternative name massively parallel sequencing (MPSS). This differs from traditional Sanger sequencing, which generates data from a population of DNA molecules that are all co-amplified during the initial PCR step.
How do MPSS systems determine the sequence of single fragments of DNA? A number of approaches are being used, and more are in development. For example, a platform developed by Pacific Biosciences is capable of visualizing individual incorporation events as nucleotides are added to a single molecule of template DNA. This extraordinary feat allows the sequencing of very long stretches of DNA (up to 10,000 bp) but is subject to relatively high error rates. Other companies are developing nanopores that will allow a single DNA molecule to pass through under an electric field. Sequence is then interpreted as the DNA molecule (either intact or as serially degraded single nucleotides) moves through the narrow channel or pore. These technologies hold the promise of extremely high throughput, but they are still in early development.
Most next-gen sequencing is currently being performed on platforms that determine the sequence from clones rather than single molecules of DNA. Clones generate more signal during the actual sequencing reaction and are therefore easier to work with. There are two common methods for generating libraries of clones from original DNA molecules. One is to seed the single DNA molecules across the surface of a glass slide and then PCR-amplify each in its place, creating microcolonies of identical DNAs.
20 These colonies, or clones, provide the substrate for subsequent sequencing reactions. The other approach is to mix a single DNA molecule together with a small sphere, or particle, within a tiny droplet of water suspended in oil (an emulsion).
21 Also included in the droplet are DNA polymerase and dNTPs, leading to amplification of the DNA across the surface of the sphere as the emulsion is subjected to PCR thermocycling. Both approaches generate a library consisting of thousands to millions of clonal DNA fragments, which are then sequenced all at the same time in a parallel fashion.
Depending on how the library of clones is generated, there may be anywhere from a few dozen to several thousand clones that represent copies of a particular segment of the original DNA. The resulting sequence from each clone is called a
read. The sensitivity of MPSS for sequence alterations is directly determined by the number of reads that is generated for each segment of the original DNA. When sequencing the whole genome, 30 reads (or 30-fold coverage) is considered a minimum amount of data. For studies focused on the coding regions of the genome (the
exome, representing approximately 1.8% of the total genome), read depths are generally greater than 100, whereas assays that are limited to a few hundred genes or less will often have read depths greater than 500, supporting mutation sensitivities below 5% mutant allele.
22 An important consideration when evaluating MPSS data is the quality of the sequence reads. This can be influenced by many factors, including the source of the DNA or RNA/cDNA (e.g., FFPE versus fresh-frozen), the length of the reads, and whether the sequencing was performed in just one or both directions on the same piece of DNA. Another factor is the level of noise in MPSS datasets. Random errors occur during the initial clonal amplifications, as well as during the subsequent sequencing steps, creating variations in the sequence that must be distinguished from real changes. Finally, when MPSS is used to search for possible mutations in tumor DNA, it is best to compare the results with the sequence of germline DNA from the same individual, thereby defining which alterations are somatic. Without this comparison to the germline, it can be difficult to distinguish mutations from natural polymorphisms, which are abundant in all genomes.
Next-Generation Sequencing Based on Fluorescence Detection
There are several next-generation sequencing platforms that are based on the detection of fluorescence events during sequencing. The 454 system (Roche) is based on combination of emulsion PCR and pyrosequencing, using the same chemistry as described under single-gene assays.
23,24 The SOLiD system (Life Technologies) performs sequencing on glass slides by measuring the binding and ligation of short, fluorescently tagged oligonucleotides as they match up to the template DNA.
20 The most widely used systems, from Illumina Corp., also perform sequencing on glass slides, but measure the incorporation of individual fluorescently labeled nucleotides, similar to those used in Sanger sequencing.
25 The throughput on the Illumina systems has been expanded to the point where several whole-genome sequences can be generated simultaneously, while the cost per sample has decreased dramatically.
Next-Generation Sequencing Based on Semiconductors
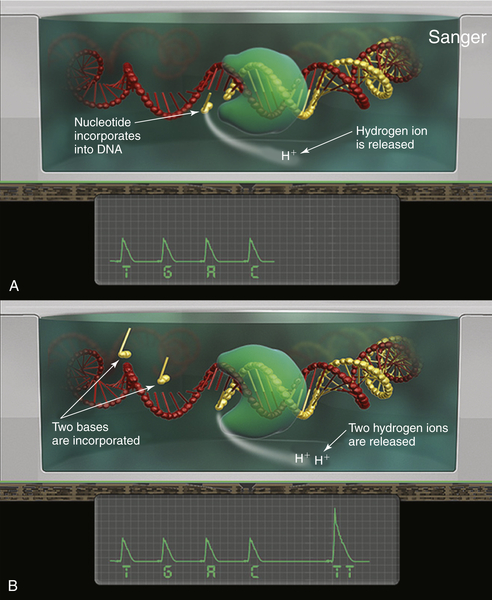
One drawback to the fluorescence-based next-generation sequencing platforms is that they generate large numbers of very high-resolution image files, requiring terabytes of storage and considerable computing power to analyze. An alternative to the use of fluorescence for sequencing has been developed by Ion Torrent (Life Technologies). Their platform looks for the pH change generated as a nucleotide is incorporated into a nascent strand of DNA (a hydrogen ion is released).
26 Sequencing is performed on modified semiconductor chips containing between 1.2 million and 1.1 billion nanowells, each serving as a micro pH meter (
Figure 59-6 ). Nanospheres that have been clonally loaded with DNA by emulsion PCR are seeded into the nanowells on the chip, and sequencing commences as nucleotides are serially flowed across the chip surface. Because pH changes are converted directly into electrical signals, this approach is very
fast and generates sequence files that are smaller than those from fluorescence-based platforms.
Figure 59-6 Semiconductor-based sequencing This method sequences DNA that has been clonally amplified onto nanospheres (0.5 to 3 μm) by measuring a drop in pH when hydrogen ions are released during nucleotide incorporation. (A) The nanospheres are distributed across a series of microwells, numbering from 1 to 660 million on a chip, each serving as an individual micro-pH meter. (B) As nucleotides are serially diffused across the chip surface, they are incorporated onto the nanosphere template DNA, and the resulting pH changes are interpreted to derive the template sequence.
Future Directions
The diagnosis of cancer has been, and will continue to be, a process that depends heavily on microscopic examination of the tissue. Identifying areas of neoplasia, determining their relationship with surrounding normal tissue elements, and assessing the extent of the disease are all key elements in defining the characteristics of a malignancy. However, the field of pathology is quickly moving from routine histological assessments to increasingly sophisticated assays for molecular markers that have prognostic and/or predictive importance. The introduction of next-generation sequencing is serving to uncover the true diversity of cancers as well as to define recurring mutations that can be targeted with new therapies. Such genomic-level analyses will continue to have an impact for many years. Some have even predicted that sequencing will make microscopy moot; however, this view ignores the growing importance of assessing the relationship of cancer cells to surrounding stromal cells and to immune cells—whose influence is critical to cancer cell survival. Thus, imaging will also play a critical role in coming years, as new in situ approaches to measuring molecular events are developed and used to help further the field.
References
1. Reid A.H. , Fanning T.G. , Hultin J.V. , Taubenberger J.K. Origin and evolution of the 1918 “Spanish” influenza virus hemagglutinin gene . Proc Natl Acad Sci U S A . 1999 ; 96 : 1651 – 1656 .
2. Archer F.L. , Kao V.C. Immunohistochemical identification of actomyosin in myoepithelium of human tissues . Lab Invest . 1968 ; 18 : 669 – 674 .
3. Fabbro D. , Di Loreto C. , Stamerra O. , Beltrami C.A. , Lonigro R. , Damante G. TTF-1 gene expression in human lung tumours . Eur J Cancer . 1996 ; 32A : 512 – 517 .
4. Werling R.W. , Yaziji H. , Bacchi C.E. , Gown A.M. CDX2, a highly sensitive and specific marker of adenocarcinomas of intestinal origin: an immunohistochemical survey of 476 primary and metastatic carcinomas . Am J Surg Pathol . 2003 ; 27 : 303 – 310 .
5. Jones T.D. , Ulbright T.M. , Eble J.N. , Baldridge L.A. , Cheng L. OCT4 staining in testicular tumors: a sensitive and specific marker for seminoma and embryonal carcinoma . Am J Surg Pathol . 2004 ; 28 : 935 – 940 .
6. Mottolese M. , Venturo I. , Benevolo M. et al. Immunocytochemical diagnosis of amelanotic metastatic melanoma using monoclonal antibodies HMB-45 and Ep1-3 . Melanoma Res . 1994 ; 4 : 53 – 58 .
7. Wikstrand C.J. , Hale L.P. , Batra S.K. et al. Monoclonal antibodies against EGFRvIII are tumor specific and react with breast and lung carcinomas and malignant gliomas . Cancer Res . 1995 ; 55 : 3140 – 3148 .
8. Capper D. , Preusser M. , Habel A. et al. Assessment of BRAF V600E mutation status by immunohistochemistry with a mutation-specific monoclonal antibody . Acta Neuropathol . 2011 ; 122 : 11 – 19 .
9. Mellors R.C. The application of labeled antibody technics in studying cell antigens . Cancer Res . 1968 ; 28 : 1372 – 1381 .
10. Varndell I.M. , Polak J.M. , Sikri K.L. , Minth C.D. , Bloom S.R. , Dixon J.E. Visualisation of messenger RNA directing peptide synthesis by in situ hybridisation using a novel single-stranded cDNA probe. Potential for the investigation of gene expression and endocrine cell activity . Histochemistry . 1984 ; 81 : 597 – 601 .
11. Sanger F. , Coulson A.R. A rapid method for determining sequences in DNA by primed synthesis with DNA polymerase . J Mol Biol . 1975 ; 94 : 441 – 448 .
12. International Human Genome Sequencing Consortium . Initial sequencing and analysis of the human genome . Nature . 2001 ; 409 : 860 – 921 .
13. Ronaghi M. , Uhlén M. , Nyrén P. A sequencing method based on real-time pyrophosphate . Science . 1998 ; 281 363, 365 .
14. Ririe K.M. , Rasmussen R.P. , Wittwer C.T. Product differentiation by analysis of DNA melting curves during the polymerase chain reaction . Anal Biochem . 1997 ; 245 : 154 – 160 .
15. Sommer S.S. , Cassady J.D. , Sobell J.L. , Bottema C.D. A novel method for detecting point mutations or polymorphisms and its application to population screening for carriers of phenylketonuria . Mayo Clin Proc . 1989 ; 64 : 1361 – 1372 .
16. Strerath M. , Detmer I. , Gaster J. , Marx A. Modified oligonucleotides as tools for allele-specific amplification . Methods Mol Biol . 2007 ; 402 : 317 – 328 .
17. Su Z. , Dias-Santagata D. , Duke M. et al. A platform for rapid detection of multiple oncogenic mutations with relevance to targeted therapy in non-small-cell lung cancer . J Mol Diagn . 2011 ; 13 : 74 – 84 .
18. MacConaill L.E. , Campbell C.D. , Kehoe S.M. et al. Profiling critical cancer gene mutations in clinical tumor samples . PLoS One . 2009 ; 4 : e7887 .
19. Beadling C. , Heinrich M.C. , Warrick A. et al. Multiplex mutation screening by mass spectrometry evaluation of 820 cases from a personalized cancer medicine registry . J Mol Diagn . 2011 ; 13 : 504 – 513 .
20. Shendure J. , Porreca G.J. , Reppas N.B. et al. Accurate multiplex polony sequencing of an evolved bacterial genome . Science . 2005 ; 309 : 1728 – 1732 .
21. Nakano M. , Komatsu J. , Matsuura S. , Takashima K. , Katsura S. , Mizuno A. Single-molecule PCR using water-in-oil emulsion . J Biotechnol . 2003 ; 102 : 117 – 124 .
22. Wagle N. , Berger M.F. , Davis M.J. et al. High-throughput detection of actionable genomic alterations in clinical tumor samples by targeted, massively parallel sequencing . Cancer Discov . 2012 ; 2 : 82 – 93 .
23. Margulies M. , Egholm M. , Altman W.E. et al. Genome sequencing in microfabricated high-density picolitre reactors . Nature . 2005 ; 437 : 376 – 380 .
24. Wheeler D.A. , Srinivasan M. , Egholm M. et al. The complete genome of an individual by massively parallel DNA sequencing . Nature . 2008 ; 452 : 872 – 876 .
25. Hillier L.W. , Marth G.T. , Quinlan A.R. et al. Whole-genome sequencing and variant discovery in . C. elegans. Nat Methods . 2008 ; 5 : 183 – 188 .
26. Rothberg J.M. , Hinz W. , Rearick T.M. et al. An integrated semiconductor device enabling non-optical genome sequencing . Nature . 2011 ; 475 : 348 – 352 .