The Past, Present, and Future of Biomarkers: A Need for Molecular Beacons for the Clinical Management of Pancreatic Cancer
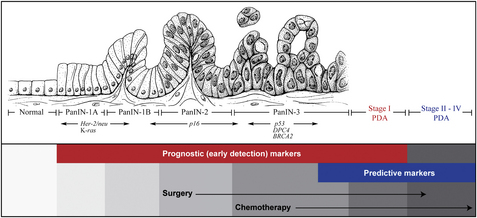
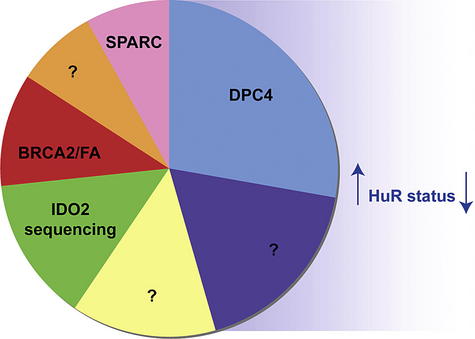
Pancreatic ductal adenocarcinoma (PDA) is one of the most lethal cancers. It is the fourth leading cause of cancer-related death in the United States [1]. Nearly 40,000 Americans are affected by this disease every year and more than half of these individuals succumb to cancer-related complications [1]. Even with cases that are identified early and undergo surgical resection, the diagnosis of PDA is associated with an overall 5-year survival rate of only 6% to 25% [2]. Although many resources and large genome-profiling studies have been completed (Fig. 1) [3,4], the clinical management of this disease has still made only modest strides in the past 2 decades.
Treatment options other than surgical resection (only 20% of patients meet the criteria for resection) include chemotherapy protocols that routinely use gemcitabine, and often include radiotherapy. Since 1997, gemcitabine has been used as the standard of care for the treatment of PDA in both the postoperative adjuvant and metastatic settings [5]. Moreover, numerous clinical trials have compared multiple drugs in combination with gemcitabine, including 5-fluorouracil (5-FU), capecitabine (Xeloda), and erlotinib (Tarceva), but all these drug combinations have failed despite costly clinical trials involving thousands of patients. Dr Roland Schmid [6], in an editorial in Gastroenterology, stated: “In fact, pancreatic cancer is the number 1 killer in phase III trials.”
For PDA, there have been 2 areas of recent promise. First, in medical oncology, therapies targeted against DNA repair pathways (eg, BRCA2 and Fanconi anemia deficiencies) have attracted attention. Second, a less targeted and selected therapy, but one that showed a modest improvement in patient outcomes in the metastatic setting, was presented at the 2010 American Society of Clinical Oncology (ASCO) meeting, with the regimen of FOLFIRINOX (5-FU/leucovorin [LV], irinotecan [I], and oxaliplatin [O]), which increased survival in the metastatic setting by roughly 4 months [7]. Other intriguing therapies that have received attention in the national landscape include paclitaxel (Abraxane), which has validated the concept of targeting the stromal portion (ie, the SPARC protein) of a tumor mass [8]. Abraxane is currently indicated for chemoresistant breast cancers, and it is debatable whether this drug will be useful (based on SPARC positivity) for a select group of patients who have pancreatic cancer in the phase III setting. Other than a half-dozen promising, theoretic strategies, most physicians (particularly oncologists) treating patients with PDA agree that they are largely focused on treating their patients in the palliative setting.
These modest improvements in patient outcomes are not caused by a lack of understanding or interest in the genetic basis of this disease [3,4]. A plethora of work was recently added to by a genome-wide survey of multiple pancreatic cancers. These data were coordinated with gene expression analyses of 24 pancreatic tumors to define 12 core signaling pathways involved in the development of PDA [4]. Although many novel targets (including membrane proteins) were newly identified, these studies primarily discovered and validated pathways and mutated genes that are disrupted in PDA [9]. An additional survey of familial pancreatic cancer genomes revealed an additional gene mutated in the Fanconi anemia DNA repair pathway: PALB2 [3]. Identifying other genes disrupted in this pathway may have important predictive biomarker implications for the treatment of PDA that harbor such PALB2 mutations and related pathway mutations (discussed later).
Current biomarkers used for the clinical management of PDA
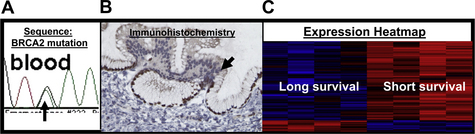
Tumor biomarkers (prognostic and predictive) are molecules, typically proteins, that are produced by the body in response to the presence of tumor cells and that may be detected in blood, urine, other bodily secretions, or tissue samples. The definition of biomarkers can be stratified into those with (1) prognostic and (2) predictive value. Simplistically, prognostic markers can be considered as those that identify patients with different risks of outcome (eg, for recurrence of disease) and can be helpful in up-front therapeutic decision making. A predictive marker can be defined as a clinically useful marker for prediction of a response to a specific therapy or treatment regimen. Prognostic and predictive markers can be (1) a defined genetic sequence (eg, BRCA2); (2) a cutoff level of a specific protein expression (eg, SPARC expression); (3) detection of a specific level of mRNA expression (eg, epidermal growth factor receptor [EGFR] mRNA levels); (4) any identifiable pathologic feature of a tumor (eg, lymph node positivity); (5) subcellular localization of a protein in a tumor or stromal cell (eg, HuR); and/or (6) any relevant molecular signature (mRNA, DNA, protein, or any combination of these aspects of a tumor); for example, as is currently used in breast cancer. Typically, a biomarker is described in the literature as 1 gene (either a DNA mutation or polymorphism on the genetic level; or the quantitative expression on the mRNA level with the use of quantitative polymerase chain reaction [PCR] technology; or immunohistochemistry on the protein level) (Fig. 2). A marker can have both prognostic and predictive value [10].
The limitations and usefulness of the available conventional markers
Unlike other tumor types, such as prostate cancer (eg, prostate-specific antigen), no simple blood test has been developed to detect the presence of a premalignant or malignant pancreatic cell (see Fig. 1). The most commonly used serum markers for pancreatic cancer are cancer antigen 19-9 (CA19-9), carcinoembryonic antigen (CEA), and rarely cancer antigen 125 (CA-125). The sensitivity and specificity of serum markers depend on the determination of cutoff values derived from different studies. For example, 1 study used different cutoff values of CA19-9 (37, 100, 200, and 300 U/mL) yielding different sensitivities (68%, 41%, 24%, and 15%, respectively) and specificities (70%, 86%, 96%, and 100%, respectively) for the same marker [11]. This study shows the lack of specificity and reproducibility of these tests as well as the subjectivity of using a desired cutoff value. With that stated, these markers are currently the most widely used clinically, and thus this article briefly describes the background and importance of each of these markers in the context of PDA.
CA19-9
The carbohydrate antigen CA19-9 is a tumor-associated antigen released into the serum of patients with gastrointestinal (GI) cancers [12]. CA19-9 was originally identified and developed in the late 1970s and early 1980s, when researchers were in search of a better and more specific diagnostic tool than CEA levels for GI cancers [13–15]. CA19-9 is a monosialoganglioside first isolated from a colorectal cell line, and since its discovery it has rapidly become the most widely used biomarker for pancreatic cancer [13,14]. As a biomarker in patients with PDA, CA19-9 is primarily used clinically to follow the response to therapy (both before and after surgical resection and chemoradiotherapy). CA19-9 is not specific for PDA, because it increased in 21% to 42% of cases of gastric cancer and 20% to 40% of cases of colon cancer. Overall, CA19-9 is increased in 71% to 93% of pancreatic cancer, and it has proved useful to differentiate benign from malignant pancreatic disease. However, most clinicians would gladly eliminate CA19-9 for a more reliable biomarker.
As a prognostic marker, it has been shown that CA19-9 can correlate with tumor resectability and pathologic features such as tumor stage [11]. More recently, in patients who had undergone pancreaticoduodenectomy for right-sided PDA, it was shown that postoperative levels of CA19-9 (at a cutoff value of 120U/ml) can stratify for both overall and disease-free survival, but does not correlate with margin status or lymph node positivity. In this study, Berger and colleagues [14] performed a prospective analysis of postoperative CA19-9 levels in patients treated with pancreatic resection and adjuvant chemoradiotherapy. This study confirmed the prognostic value of postresection CA19-9 levels in predicting outcome in patients with PDA.
Despite its widespread use, CA19-9 falls short as an informative marker for several reasons. First, a high percentage of patients with pancreatic cancer do not have any detectable CA19-9 in their sera, which rules out the use of this marker for use in early diagnosis and the further treatment of this cohort of patients. Second, CA19-9 does not accurately and reproducibly correlate with the clinical manifestations of this disease. For instance, in a pancreatic cancer registry in Japan, only 48% of patients with PDA smaller than 2 cm had increased CA19-9 [15]. Third, it has been observed that patients who are Lewis blood type negative (Le ab−) do not express the CA19-9 antigen, and, thus, a Lewis ab− blood type needs to be considered in patients with pancreatic cancer who lack an increased serum CA19-9 level (estimated to be 5%–10% of the population) [16,17]. Fourth, CA19-9 is also increased in inflammatory lesions of the pancreas, such as pancreatitis, which may interfere with the absolute diagnosis of PDA based on CA19-9 alone [18].
The variability in CA19-9 serum levels described earlier is not only detected in the patient population but can be variable throughout the course of treatment of an individual. However, this lack of reproducible quantitative measurement of this marker gives patients a false sense of understanding of their disease course. In many clinical instances, such as post-pancreatic resection, patients rely too heavily on their CA19-9 value as a marker for curable or silent disease. These numbers can give a false sense of hope to patients who get excited about a dramatic drop in their CA19-9 levels, only to be faced with the finding of a high CA19-9 value on follow-up visits. Therefore, CA19-9 cannot be used as a stand-alone marker, and the use of this marker should be assessed in conjunction with other markers (eg, CEA; see later discussion), imaging, and patient symptoms regarding clinical decision making. The primary use of CA19-9 as a tumor marker is to vaguely monitor a patient’s response to pancreatic cancer treatment and/or cancer progression [19].
CEA
CEA is a 180-kDa glycoprotein that was first discovered more than 45 years ago [20]. As a biomarker, CEA is primarily used in decision making in the course of treatment of colorectal cancers, but is also used in other cancers such as stomach, breast, and pancreatic cancers [21]. Similar to a CA19-9 blood test, a CEA blood test measures the specific levels of CEA protein in a patient’s serum. As with CA19-9, the specific link to pancreatic cancer is limited because of the lack of specificity (ie, many tumor types produce CEA) and because some patients never produce CEA at all. Thus, this marker cannot be used as a reliable early detection marker or a relevant biomarker throughout a patient’s disease course [22]. Distinct from levels in the serum, CEA levels in pancreatic cyst fluids may have some value in detecting pancreatic mucinous, premalignant, or malignant cysts [23].
A warning of caution is that serum CEA levels can be increased (falsely positive for cancer) in patients who have certain infections, pancreatitis, and cirrhosis of the liver. In addition, individuals who smoke cigarettes can have increased CEA levels compared with nonsmokers [24–26], in the absence of malignancy. Therefore, the realistic clinical usefulness of CEA may lie in its ability to be included as part of a multipanel biomarker panel for early detection screening purposes.
CA-125 and final thoughts on conventional markers
CA-125, or carbohydrate antigen 125, was first identified by immunizing ovarian cancer cells to mice [24]. CA-125 was later characterized and labeled as the human gene encoded by the MUC16 gene [27]. MUC16 was thus identified as a new member of the mucin family [27]. CA-125 is an antigen present on 80% of nonmucinous ovarian carcinomas [27] and has therefore been investigated for possible use as a biomarker in patients with ovarian carcinoma. Subsequently, CA-125 has been found to be increased in other cancers including pancreatic cancer and other gynecologic and nongynecologic conditions [23]. CA19-9 can be more useful in patients who do not present with jaundice, whereas CA-125 provides a limited contribution in jaundiced patients [25]. In current clinical practice, CA-125 is not sufficiently accurate to be useful in patients with pancreatic cancer.
The use of these markers (CA19-9, CEA, and to some extent CA-125) has already helped guide and monitor the treatment of many patients with PDA. However, taking into account the high, but still limited, sensitivity and specificity of the CA19-9 and CEA tests, their results in differential diagnosis of distinct pancreatic tumors should be interpreted cautiously, and in reference to imaging results (eg, ultrasonography, computed tomography, and magnetic resonance imaging) [26]. Together, these markers may be useful in the future of therapy for patients with PDA. Realistically, 1 or all these markers may be used in the future in a multiplexed panel with novel, presently undiscovered markers for this disease [28]. The molecular understanding of the dysregulation of these markers (eg, hypomethylation of a specific gene or overexpression of a transcriptional factor that regulates a specific gene) [29] in PDA cells may provide a useful insight into the molecular cause of this disease.
Novel markers (including early detection) of pancreatic cancer
The prognosis of PDA is poor, not only because of its aggressive biologic behavior, but because its commonly missed or late clinical diagnosis often prevents initiation of established curative therapies such as surgery. Therefore, one of the field’s major goals is to find molecular markers, specific and sensitive enough to make an early and correct diagnosis of early stage PDA and/or pancreatic cancer precursor lesions, before the tumor becomes unresectable and while patients are still clinically asymptomatic (see Fig. 1) [30]. Although the molecular markers (CA19-9 and CEA) described earlier have been used for the early diagnosis of PDA, these markers fail in many circumstances to differentiate benign from malignant processes, and also fail to differentiate locally confined resectable disease from widespread metastatic disease.
Currently, several notable and useful molecular early detection markers are genetic based and only have clinical significance for the familial form of PDA (familial pancreatic cancer [FPC]). The genome-maintenance and DNA repair genes, which include BRCA2 (see Fig. 2A), and the Fanconi anemia genes (FANCC, FANCG, and PALB2/FANCN) have received considerable attention in the past decade because of (1) the high frequency of inherited mutations in this pathway found in patients with FPC [3,31], and (2) the recent work on platinum-based and PARP inhibitor–based therapies [32], which have shown that tumors defective in this pathway should be exquisitely sensitive to these agents [3,33–35]. Thus, as a biomarker, a DNA sequence (detection of a germline mutation in these genes from constitutional DNA) in individuals composing a family that has a high presence of pancreatic and other related cancers (ovarian and breast) can be useful as a diagnostic and a predictive marker [33]. Myriad Genetics (Salt Lake City, UT, USA) offers a genetic survey of these genes based on a simple blood test to aid physicians and families in their decision making. The results may be used both for preventive surgical options such as prophylactic resection, or treatment options such as targeted chemotherapy (if the disease is present in family members who test positive for a germline mutation in one of the genes mentioned earlier).
Canto and colleagues [36,37] have been leaders in this field of screening high-risk families, trying to explore the importance of invasive screening tools (such as endoscopic ultrasound) to monitor high-risk individuals identified with a family history of pancreatic cancer and a positive genetic test for a BRCA2 or related gene mutation [31,36,38]. However, the debate continues as to the best overall strategy [37]. Further, a recent publication warns that individuals who carry a germline BRCA2 mutation may not harbor a loss of the corresponding allele or a second hit in the pancreatic cancer cells [39]. Therefore, if the 2-hit hypothesis does not apply here, the theoretic therapeutic window (ie, an Achilles heel of the tumor) for platinum-based or PARP inhibitor–based therapies may be negated [39], along with the predictive value of the BRCA2 gene status in these patients. Larger and more extensive genome-wide survey studies may find that certain carriers of a BRCA2 or relate gene mutation may require another genetic aberration (eg, a single nucleotide polymorphism [SNP] or mutation) to have clinical significance for cancer development and/or targeted treatment.
Recently, another landmark study was the discovery that the DPC4 (SMAD4) status of pancreatic cancers at the time of autopsy differentiates those patients who died with locally advanced and locally destructive pancreatic cancer from those patients who died with widespread metastatic disease [40]. Thus, it seems that DPC4 gene silencing [41] can stratify pancreatic tumors into 2 biologically distinct tumor phenotypes [40,41]. These findings underscore the involvement of DPC4 inactivation in pancreatic tumorigenesis and suggest a role for DPC4 status as a biomarker for specific local therapies (eg, radiotherapy for individuals with predicted locally aggressive tumors) or specific systemic therapies (eg, chemotherapy for individuals with predicted high risk of metastatic dissemination).
Besides genetic-based and immunohistochemistry-based tests, more accurate serum markers of pancreatic cancer may improve the early detection and prognosis of this deadly disease. Many other candidate PDA serum markers have been evaluated using specific assays, but none are superior to CA19-9 [42]. These markers include amylin (islet amyloid polypeptide), DUPAN-2 [45], CA242 [17], CAM 17.1, TPS, CA72-4, SPan-1, CA50, CA195, TATI, POA, YKL-40, and TUM2-PK [43–49]. Other peptides identified by surface-enhanced laser desorption and ionization [50], and markers such as MUC4 and synuclein [51], may prove useful but await confirmatory studies and assays. Koopmann and colleagues [52] reported that, in the stratification of patients with resectable pancreatic cancer from controls, serum macrophage inhibitory cytokine 1 (MIC-1) outperforms other markers including CA19-9. Most of the studies mentioned earlier have generated little validation in independent studies or ongoing citations, despite some being placed in prestigious journals related to pancreatic cancer.
Other innovative approaches to the early diagnosis of pancreatic cancer have included screening for early detection proteins via a proteomic screen of pancreatic juice [53]. The investigators identified a protein they labeled as PAP-2 and they claim that proteins found in this analysis could potentially be used as biomarkers in an immunoassay for the detection of suspicious pancreatic lesions. A follow-up paper to this initial publication by a separate group showed that a combined human screen with a screen of a genetically engineered mouse model of pancreatic cancer could potentially discover novel, useful biomarkers [54]. More recent approaches, such as differential proteomic screening approaches [55] and a Luminex platform using 3-gene signatures [28], have yielded some candidate biomarkers and panels that are pending validation studies. In the clinical setting of better identifying patients who have pancreatic cancer, the superior 3-marker signature included CA19-9, CEA, and TIMP1, with a sensitivity-specificity of 71% to 90%, compared with CA19-9 alone with a lower sensitivity (51%–90%).
Another promising line of investigation evaluates circulating tumor cells (CTCs) as a noninvasive means of detecting and categorizing a suspicious pancreatic mass or premalignant lesion. Strategies using CTCs are currently being developed for the detection and monitoring of therapy for breast and prostate cancers as well as pancreatic cancer [48,56,57]. The key aspects in the development of these strategies is to (1) adequately isolate CTCs and, at the same time, establish a cutoff value, and (2) define the appropriate marker or markers to determine the aggressiveness of asymptomatic disease.
These and other research studies [58] represent a widespread effort that includes multiple centers, a large number of patients, high-throughput data analyses, validation studies, antibody validation assays, and recruitment of healthy and disease-appropriate control groups [34]. More emphasis and funds need to be put into similar studies and further validation efforts, to define markers for individuals who are asymptomatic (ie, who harbor PanIN lesions or stage I disease) (see Fig. 1), so that viable treatment strategies like surgery can be provided to patients in a timely fashion [28,30].
Emerging and alternative molecular (predictive) biomarkers for the treatment of PDA
Biomarker studies rely heavily on the central dogma of molecular biology; that is, that DNA is made into RNA, which is then translated into protein. Most attention and focus on the processes that regulate the central dogma in cancer cells has been on alterations in DNA sequences and dysregulation of RNA expression levels. However, until recently, little attention has been focused on post-transcriptional gene regulation, a powerful and highly efficient mechanism that involves micro-RNAs (miRNA) and/or RNA-binding proteins (RBPs) that regulate protein expression levels. These molecules work through a series of highly regulated events that include specific sequence recognition motifs and enzymes. Abnormal levels of certain miRNAs and RBPs are detected in pancreatic cancer cells compared with normal cells [59,60]. The authors strongly believe these molecules (ie, miRNA and RBPs), that are involved in post-transcriptional gene regulation, are critical for driving pancreatic tumorigenesis and may also be the basis for chemotherapeutic resistance mechanisms [61–63]. Therefore, exploring the functional aspects of these molecules should help generate novel, and perhaps reliable, biomarkers for this disease.
HuR as a biomarker in pancreatic cancer
HuR is an RBP that regulates genes involved in the normal cellular response to several stressors. Increased cytoplasmic levels of HuR (ie, activated HuR) support cancer cell growth and survival [61–64]. HuR is referred to as activated when, in response to a stressor, it traffics to the cytoplasm and regulates the expression of specific HuR target genes. HuR is part of the embryonic lethal, abnormal vision, Drosophila-like, mRNA stability protein family [59]. When triggered by pancreatic-associated stressors (eg, hypoxia from the tumor microenvironment or specific drugs), HuR traffics to the cytoplasm where it potently influences the stabilization and translation of key survival and growth-related mRNAs [65–67]. This process defines HuR as a critical molecule involved in posttranscriptional gene regulation. As an example, more than a decade ago, it was reported that HuR increases the protein levels of the angiogenic factor vascular endothelial growth factor (VEGF) after a cell is in a hypoxic environment [68]. VEGF has been well established as a molecule important in pancreatic tumorigenesis.
Mechanistically, HuR regulates specific mRNA cargos that contain a U-rich or AU-rich sequence, typically residing in the 3’-untranslated region (UTR) of survival transcripts [69]. In relation to the pancreatic tumorigenesis process, some of these cargos include VEGF, p21, cyclins A and B1, COX-2, HIF-1, IGF-IR, and p53 [66,67,69]. To date, known HuR mRNA targets have expanded in number because of more global analyses being reported from array experiments from different laboratories [60]. In brief, HuR typically upregulates its targets by either affecting mRNA stability or protein translation of these target mRNAs [52]. There are examples and a precedent for HuR to also downregulate the protein expression of select target mRNAs, most notably IGF-1R, Wnt-5a, c-Myc, p27, and Death Receptor 5 [70–73] (Ritenhouse and colleagues, unpublished data). Accordingly, HuR seems to downregulate or upregulate mRNA targets, with the main objective of providing a pro-survival advantage to a pancreatic cancer cell under stress.
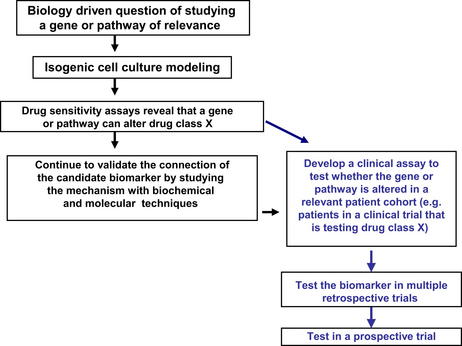
For PDA tumorigenesis, 12 core signaling pathways have recently been identified as being critical for the development of pancreatic cancer [4]. Nearly all (99%) of the identified tenfold overexpressed genes (identified via serial analysis of gene expression [SAGE] analysis) in pancreatic cancers were not linked to a specific genetic mutation [4]. Instead, we have discovered that these genes are potentially regulated, in part, by HuR (our unpublished data). Using bioinformatic software described previously in the literature, we identified putative HuR targets from the 540 listed overexpressed genes in PDA [4,69,74]. By PCR-based analysis using ribonucleoprotein immunoprecipitation assays (RNP-IP) of HuR bound mRNAs, we identified 60 putative HuR-regulated target genes of the 540 overexpressed genes (which accounts for 11.1% of the genes overexpressed by tenfold or more compared with normal cells) in PDA [4]. In comparison, it is estimated that genetic and epigenetic alterations contribute only 1% and 1.8% respectively to the possible mechanisms by which these 540 overexpressed genes are disrupted in PDA. Ten genes, including K-Ras [75], were experimentally validated as HuR targets involved in 5 of the 12 core signaling pathways in PDA. This work suggests that HuR is an unprecedented protein predicted to be involved in at least 5 of the 12 core signaling pathways of pancreatic cancer cell development and survival [4]. In relation to biomarker discovery, this work is mentioned here because biomarker discovery may likely come from these exploratory studies that complement cancer genome-wide profiling studies of DNA alterations and RNA expression levels [4]. Thus, focused research efforts on potent regulators like HuR may help identify novel prognostic and predictive markers [75,76] and targets for the treatment of PDA [66] (Fig. 3).
Based on the molecular work described previously, it has been shown that high cytoplasmic HuR expression (as a prognostic marker of adverse outcome) directly correlates with worse pathologic prognostic features of pancreatic cancers [77,78]. These data correlate with multiple studies of HuR in clinical samples that show that high abundance cytoplasmic HuR correlates with worse pathologically prognostic features (including high T stage and a greater positive lymph node ratio) [78]. In contrast with genetic events that may take decades to evolve and exert biologic influence, the regulation of post-transcription by HuR is an efficient, rapid, and potent mechanism by which cancer cells can orchestrate changes in multiple signaling pathways and thus thrive in many tumor-specific environments including hypoxic conditions and the exposure to stressful chemotherapeutic agents [66].
Previously reported clinical correlates and predictors of standard of care therapy for PDA (gemcitabine)
Gemcitabine (2’,2’-difluorodeoxycytidine; Gemzar) is the primary chemotherapeutic agent used against pancreatic cancer because it has the best evidence of clinical benefit [5]. Since its approval in 1996 for use as single-agent therapy, gemcitabine has been the reference therapeutic drug used against pancreatic cancer. However, gemcitabine therapy only modestly increases the overall survival of molecularly unselected patients with pancreatic cancer.
Several genes involved in gemcitabine (a prodrug) metabolism have been described as potential predictive biomarkers for gemcitabine response (Table 1) [78]. A large, multi-institutional trial (RTOG 9704) banked pancreatic tissues to stratify the population of patients with pancreatic cancer who received either gemcitabine or another antimetabolite, 5-fluorouracil. One group was able to use a tissue microarray generated by this trial to define, in a retrospective manner, human equilibrative nucleoside transporter-1 (hENT) protein expression as a predictive marker of gemcitabine response [79]. Another proposed biomarker, RRM1, the gene that encodes the M1 regulatory subunit of ribonucleotide reductase, has been shown to play a role in gemcitabine resistance in experimental preclinical cancer models and non–small cell lung cancer [76,80,81]. In clinical specimens, Akita and colleagues [82] showed a correlation between low RRM1 expression and gemcitabine response in patients with pancreatic cancer. ERCC1 was also described as a biomarker relevant to gemcitabine response [83,84]; in a testing set from our institution, we were unable to validate this work (our unpublished data). In collaboration with Baylor College of Medicine, we sequenced the REQL gene in more than 150 patient samples and only found a correlation between an REQL gene polymorphism (the AA genotype) and patient response in a neoadjuvant setting (ie, the group treated with preoperative chemotherapy) [85].
Table 1 Updated protein and genetic predictive biomarker evaluation of several candidate genes for gemcitabine response in pancreatic cancer
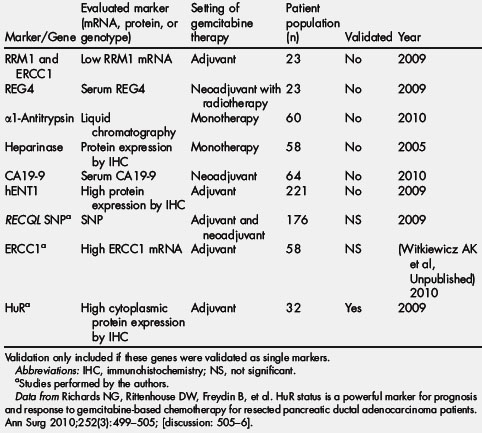
Although several candidate markers predictive of gemcitabine response have been described, they seem to lack the specificity and sensitivity required for guiding therapy in the clinical setting. Further, these predictive markers have either never been validated in independent, prospective trials or have not been tested in a pancreatic cancer–specific patient population (see Table 1) [78].
HuR is an unprecedented predictive marker for gemcitabine response in patients with PDA
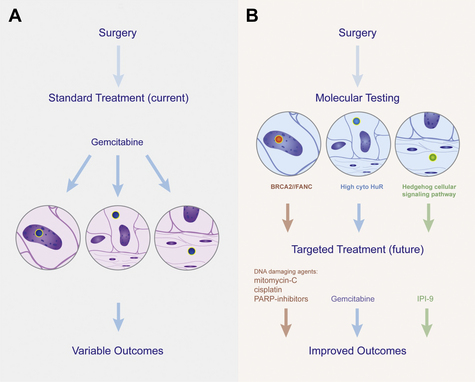
Given the role of HuR as a marker for poor prognosis in many cancers [78], a previous study on the consequences of modulating HuR levels in pancreatic cancer cells in vitro (see Fig. 3) [86] found that HuR is essential for growth and survival properties in pancreatic cancer cells (Late Breaking Abstract, AACR, 2011, Orlando, FL, USA). These data correlate with activated HuR status as a poor prognostic marker for patients with PDA [78,86]. In addition, it was determined that HuR is a powerful predictive marker of gemcitabine response. The study and validation of HuR as a predictive marker started with a defined drug screen [86]. This screening process discovered that PDA cells with exogenous overexpression of HuR had no discernable differential growth when untreated or treated with most anticancer agents (eg, 5-fluorouracil and others). However, our drug screen identified that HuR levels directly affected gemcitabine efficacy [86]. Based on this wet laboratory work, we directly tested this finding in our population of patients with pancreatic cancer (see Fig. 3). In these complementary, translational studies, we found a significant association between overall survival and activated HuR high cytoplasmic status, with the overall survival being significantly shorter for the patients having low HuR levels (a sevenfold difference) compared with those patients having high levels of HuR cytoplasmic expression [78,86]. Currently, a larger retrospective analysis is being performed to validate HuR as a clinical tool for predicting whether a PDA patient will benefit from gemcitabine-based therapy. This work exemplifies combining basic molecular biology techniques with translational pathologic analyses to define an unprecedented predictive marker [66,78,86,87] (see Fig. 3).
Another current example: The development of growth factor receptors in PDA as relevant biomarkers
EGFR has been extensively studied in PDA. Erlotinib, a small molecule inhibitor targeted against EGFR, was recently approved in combination with gemcitabine for metastatic pancreatic cancer [88]. EGFR gene amplifications and mutations are rare, but EGFR overexpression was reported in between 9% and 65% of cases, with a variable correlation with survival. Moreover, insulinlike growth factor 1 receptor (IGF-1R) was also reported to be overexpressed in 38% to 64% of the patients with PDA and its expression has been associated with a higher tumor grade, an antiapoptotic effect, and higher rates of cellular proliferation and angiogenesis [89,90]. Both receptors can heterodimerize in other tumor types, and, accordingly, IGF-1R has been identified as a potential source of resistance to EGFR-directed therapies, suggesting that dual blockage may provide greater therapeutic effects [91]. Thus, multiple phase I to II clinical trials are currently ongoing, testing IGF-1R inhibitors in a diverse range of epithelial cancers, including pancreatic cancer. Further retrospective molecular analyses from patient specimens acquired during these trials will be able to decipher whether IGF-1R and EGFR status, as well as pathway members, will be useful predictive markers for these targeted therapies.
Future exploratory work: the need for the integration of emerging technologies and systems biology to help create roadmaps for diagnosis and prediction of therapy
Large amounts of money and resources are being invested into direct sequencing and genome-wide expression analyses of pancreatic cancer genomes and transcriptomes. The promise of this work is that commonly found genetic alterations (eg, mutations and polymorphisms) in pancreatic cancer genomes or overexpressed genes will, in part, aid in the discovery of novel targets and biomarkers that are susceptible to drugs. The integration of RNA and DNA sequencing of tumor cells extracted from pathologic samples may provide a breakthrough in discovering realistic biomarkers. For instance, purifying RNA from intact tumor cells may be critical to identifying disrupted early detection candidates in their normal milieu, and not in an artificial setting as has been done with xenografted human tumors. Genome-wide expression profiling and sequencing, along with the use of laser capture microdissection techniques, are currently available and have recently provided us with landmark data [92] and should be incorporated in these high-throughput studies.
In the near future, sequencing a patient’s genome could become routine practice on a visit to the clinic by a patient with pancreatic cancer. As technologies advance and sequencing becomes less expensive, major medical centers will most likely be able to integrate and use direct sequencing of patient samples. Currently, the common goal of molecular biologists and pathologists should be to prove the validity of gene mutations, SNPs, and altered gene expression patterns as guides for the clinical management of patients with pancreatic cancer. Presently, high-throughput, genome-wide analyses of cancer genomes are taking place in multiple institutions and continents. These studies should incorporate innovative molecular techniques from the bench, with novel techniques in pathology to search for and validate robust biomarkers for this disease (see Fig. 3). Combining these results with computational programming should enhance the pace of biomarker discovery and expose uncharted regions and patterns of the genome that are important for the clinical management of PDA [93]. For example, an interdisciplinary and systems biology approach of combining biochemical techniques such as ribonucleoprotein immunoprecipitations with RNA sequencing of laser capture microdissected materials, along with cutting-edge, automated image analysis allowing for the objective quantitative analysis of protein expression in subcellular compartments in tumor samples, could identify biomarkers that conventional screening methods would be unable to detect (see Fig. 3).
Final thoughts
Arguably, the greatest resources supporting pancreatic cancer research should go into the identification of early detection strategies and predictive biomarkers. The best scenario would be to identify biomarkers that are both prognostic and predictive in value (eg, BRCA2). The challenge is to find other multifaceted markers, like BRCA2, that facilitate tumorigenesis and at the same time become that tumor’s Achilles heel (ie, a target that responds to drugs). However, the paucity of funds available for pancreatic cancer research leaves only a small amount for the validation and large-scale studies that are necessary for the success of this line of work. However, we are hopeful, and strongly believe, that the integration of previous studies with high-throughput technologies and more sophisticated biobanking should create a landscape in which realistic and reproducible biomarkers can be generated by the pancreatic cancer research community [94]. To realistically cure pancreatic cancer, 2 specific types of biomarkers are needed: (1) early detection markers that provide a timely opportunity for curative therapies to be performed (ie, surgery); and (2) predictive markers that select complementary, optimized adjuvant therapies (see Fig. 1). The development of these biomarkers will be a step toward progress (Figs. 4 and 5) that will enhance patient outcomes for this devastating disease.
References
[1] Society, AC. American Cancer Society: cancer facts and figures. Available at: http://www.cancer.org/downloads/STT/500809web.pdf, 2009. 2009
[6] R. Schmid. Pancreatologists: an endangered species? Gastroenterology. 2010;138(4):1236.
[17] R. Lamerz. Role of tumour markers, cytogenetics. Ann Oncol. 1999;10(Suppl 4):145-149.
[93] I. Rigoutsos. Short RNAs: how big is this iceberg? Curr Biol. 2010;20(3):R110-R113.