[level-membership-for-basic-science-category]
CHAPTER 19 Mitochondria, Chloroplasts, Peroxisomes
This chapter considers three organelles formed by posttranslational import of proteins synthesized in the cytoplasm. Mitochondria and chloroplasts both arose from endosymbiotic bacteria, two singular events that occurred about one billion years apart. Both mitochondria and chloroplasts retain remnants of those prokaryotic genomes but depend largely on genes that were transferred to the nucleus of the host eukaryote. Both organelles brought biochemical mechanisms that allow their eukaryotic hosts to acquire and utilize energy more efficiently. In oxidative phosphorylation by mitochondria and photosynthesis by chloroplasts, energy from the breakdown of nutrients or from absorption of photons is used to energize electrons. As these electrons tunnel through transmembrane proteins, energy is partitioned off to create proton gradients. These proton gradients drive the rotary ATP synthase (see Fig. 8-5) to make adenosine triphosphate (ATP), which is used as energy currency to power the cell. Peroxisomes contain no genes and depend entirely on nuclear genes to encode their proteins. Their evolutionary origins are obscure. Peroxisomes contain enzymes that catalyze oxidation reactions that are essential for normal human physiology. Patients who lack peroxisomes have severe neural defects.
Mitochondria
Evolution and Structure of Mitochondria
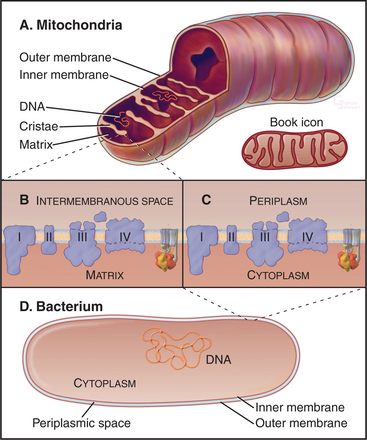
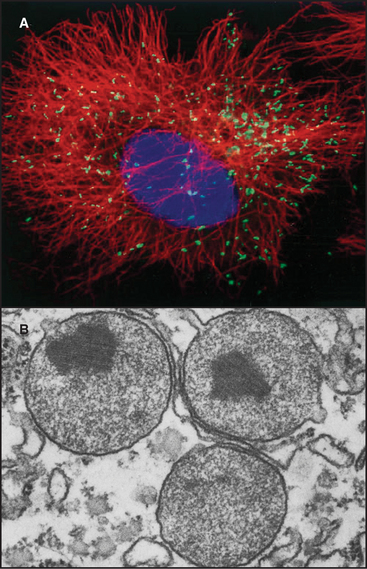
Mitochondria (Fig. 19-1) arose about 2 billion years ago when a Bacterium fused with an archaeal cell or established a symbiotic relationship with a primitive eukaryotic cell (see Fig. 2-5 and associated text). The details are not preserved in the fossil record, but the bacterial origins of mitochondria are apparent in their many common features (Fig. 19-2). The closest extant relatives of the Bacterium that gave rise to mitochondria are Rickettsia, aerobic α-proteobacteria with a genome of 1.1 megabase pairs. These intracellular pathogens cause typhus and Rocky Mountain spotted fever. However, some evidence argues that the actual progenitor bacterium had the genes required for both aerobic and anaerobic metabolism.
As primitive eukaryotes diverged from each other, most of the bacterial genes were lost or moved to the nuclei of the host eukaryotes. The pace of the gene transfer to the nucleus varied considerably depending on the species, but all known mitochondria retain some bacterial genes. A very few eukaryotes, such as Entamoeba, that branched well after their ancestors acquired mitochondria lost the organelle, leaving behind a few mitochondrial genes in the nucleus.
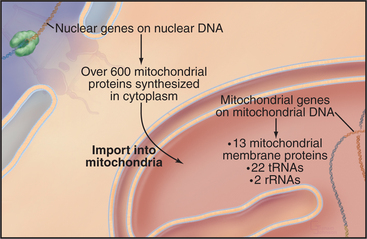
Chromosomes of contemporary mitochondria vary in size from 366,924 base pairs (bp) in the plant Arabidopsis to only 5966bp in Plasmodium. These small, usually circular genomes encode RNAs and proteins that are essential for mitochondrial function, including some subunits of proteins responsible for adenosine triphosphate (ATP) synthesis. The highly pared-down human mitochondrial genome with 16,569bp encodes only 13 mitochondrial membrane proteins, two ribosomal RNAs, and just enough tRNAs (22) to translate these genes. The number of proteins encoded by other mitochondrial genomes ranges from just 3 in Plasmodium to 97 in a protozoan. Nuclear genes encode the other 600 to 1000 mitochondria proteins, including those required to synthesize proteins in the matrix. All mitochondrial proteins that are encoded by nuclear genes are synthesized in the cytoplasm and subsequently imported into mitochondria (see Figs. 18-2 and 18-3).
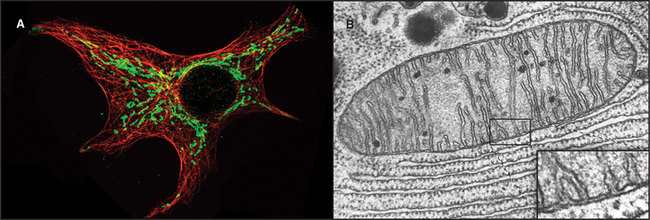
Mitochondria consist of two membrane-bounded compartments, one inside the other (Fig. 19-2). The outer membrane surrounds the intermembranous space. The inner membrane surrounds the matrix. Each membrane and compartment has a distinct protein composition and functions. Porins in the outer membrane provide channels for passage of molecules of less than 5000D, including most metabolites required for ATP synthesis. The highly impermeable inner membrane is specialized for converting energy provided by breakdown of nutrients in the matrix into ATP. Four complexes (I to IV) of integral membrane proteins use the transport of energetic electrons to create a gradient of protons across the inner membrane. The F1F0 ATP synthase (see Fig. 8-5) utilizes the proton gradient to synthesize ATP. The area of inner membrane available for these reactions is increased by folds called cristae that vary in number and shape depending on the species, tissue, and metabolic state. Cristae may be tubular or flattened sacs. Contacts between the inner and outer membranes are sites of protein import (see Fig. 18-4). Proteins in the intermembranous space participate in ATP synthesis but, when released into the cytoplasm, trigger programmed cell death (see Fig. 46-15).
Biogenesis of Mitochondria
Mitochondria grow by importing most of their proteins from the cytoplasm and by internal synthesis of some proteins and replication of the genome (Fig. 19-3). Targeting and sorting signals built into the mitochondrial proteins that are synthesized in the cytoplasm direct them to their destinations (see Fig. 18-4).
Similar to cells, mitochondria divide, but unlike most cells, they also fuse with other mitochondria. These fusion and division reactions were first observed nearly one hundred years ago. Now it is appreciated that a balance between ongoing fusion and division determines the number of mitochondria within a cell. Both fusion and division depend on proteins with guanosine triphosphatase (GTPase) domains related to dynamin (see Fig. 22-11). In fact, eukaryotes might have acquired their dynamin genes from the bacterium that became mitochondria.
One dynamin-related GTPase is required for division of mitochondria. This GTPase self-assembles into spirals that appear to pinch mitochondria in two. During apoptosis (see Chapter 46), this GTPase also participates in the fragmentation of mitochondria.
Synthesis of ATP by Oxidative Phosphorylation
Mitochondria use energy extracted from the chemical bonds of nutrients to generate a proton gradient across the inner membrane. This proton gradient drives the F1F0 ATP synthase to synthesize ATP from ADP and inorganic phosphate. Enzymes in the inner membrane and matrix cooperate with pumps, carriers, and electron transport proteins in the inner membrane to move electrons, protons, and other energetic intermediates across the impermeable inner membrane. This is a classic chemiosmotic process (see Fig. 11-1).
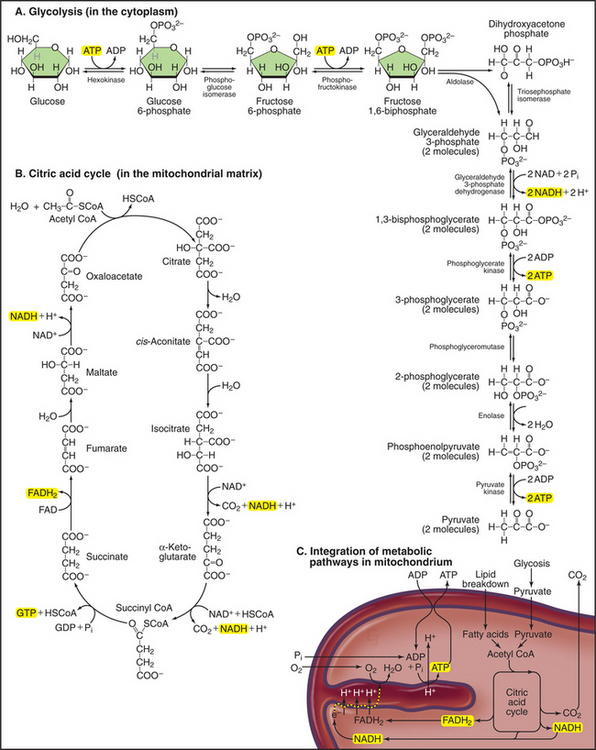
Mitochondria receive energy-yielding chemical intermediates from two ancient metabolic pathways, glycolysis and fatty acid oxidation (Fig. 19-4), that evolved in the common ancestor of living things. Both pathways feed into the equally ancient citric acid cycle of energy-yielding reactions in the mitochondrial matrix:
Breakdown of acetyl-CoA during one turn of the citric acid cycle produces three molecules of NADH, one molecule of FADH2, and two molecules of carbon dioxide. Energetic electrons donated by NADH and FADH2 drive an electron transport pathway in the inner mitochondrial membrane that powers a chemiosmotic cycle to produce ATP (Fig. 19-5). Electrons use two routes to pass through three protein complexes in the inner mitochondrial membrane. Starting with NADH, electrons pass through complex I to complex III to complex IV. Electrons from FADH2 pass through complex II to complex III to complex IV. Along both routes, energy is partitioned off to transfer multiple protons (at least 10 electrons per NADH oxidized) across the inner mitochondrial membrane from the matrix to the inner membrane space. The resulting electrochemical gradient of protons drives ATP synthesis (see Fig. 8-5).
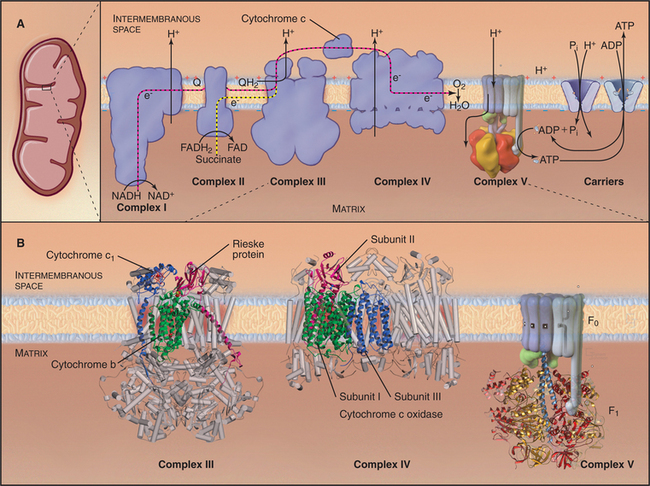
Figure 19-5 chemiosmotic cycle of the respiratory electron transport chain and atp synthase. a, left panel, A mitochondrion for orientation. Right panel, The electron transport system of the inner mitochondrial membrane. Note the pathway of electrons through the four complexes (red and yellow arrows) and the sites of proton translocation between the matrix to the intermembranous space (black arrows). The stoichiometry is not specified, but at the last step, four electrons are required to reduce oxygen to water. ATP synthase uses the electrochemical proton gradient produced by the electron transport reactions to drive ATP synthesis. B, The available atomic structures of the electron transport chain are shown. In the cytochrome bc1 complex III, the 3 of 11 mitochondrial subunits used by bacteria are shown as ribbon models. The supporting subunits found in mitochondria are shown as cylinders. The four subunits of complex IV encoded by the mitochondrial genome are shown as ribbon models. They form the functional core of the complex, which is supported by additional subunits shown as cylinders. See Figure 8-4 for further details of ATP synthase (complex V).
(B, Images of complex III and complex IV courtesy of M. Saraste, European Molecular Biology Laboratory, Heidelberg, Germany. Reference: Zhang Z, Huang L, Schulmeister VM, et al: Electron transfer by domain movement in cytochrome bc1. Nature 392:677–684, 1998. PDB file: 1BCC. Reference: Yoshikawa S, Shinzawα-Itoh K, Nakashima R, et al: Redox-coupled crystal structural changes in bovine heart cytochrome c oxidase. Science 280:1723–1729, 1998. PDB file: 2OCC.)
This process is called oxidative phosphorylation, since molecular oxygen is the sink for energy-bearing electrons at the end of the pathway and since the reactions add phosphate to ADP. Eukaryotes that live in environments with little or no oxygen use other acceptors for these electrons and produce nitrite, nitric oxide, or other reduced products rather than water. Oxidative phosphorylation is understood in remarkable detail, thanks to atomic structures of ATP synthase and three of the four electron transfer complexes. Nuclear genes encode most of the protein subunits of these complexes, but mitochondrial genes are responsible for a few key subunits.
Bacteria and mitochondria share homologous proteins for the key steps in oxidative phosphorylation (Fig. 19-2), but the machinery in mitochondria is usually more complex. Thus, bacteria are useful model systems with which to study the common mechanisms. Plasma membranes of bacteria and inner membranes of mitochondria have equivalent components, and the bacterial cytoplasm corresponds to the mitochondrial matrix (Fig. 19-2).
Energy enters this pathway in the form of electrons that are produced when NADH is oxidized to NAD+, releasing one H+ and two electrons (Fig. 19-5). If the proton and electrons were to combine immediately with oxygen, their energy would be lost as heat. Instead, these high-energy electrons are separated from the protons and then passed along the electron transport pathway before finally recombining to reduce molecular oxygen to form water. Along the pathway, electrons associate transiently with a series of oxidation/reduction acceptors, generally metal ions associated with organic cofactors, such as hemes in cytochromes and iron-sulfur centers (2Fe2S) and copper centers in complex IV. Electrons move along the transport pathway at rates of up to 1000s−1. To travel at this rate through a transmembrane protein complex spanning a 35-nm lipid bilayer, at least three redox cofactors are required in each complex, because the efficiency of quantum mechanical tunneling of electrons between redox cofactors falls off rapidly with distance. Two cofactors, even with optimal orientation, would be too slow.
The electrochemical proton gradient produced by the electron transport chain provides energy to synthesize ATP. Chapter 8 explained how the rotary ATP synthase (complex V) can either use ATP hydrolysis to pump protons or use the transit of protons down an electrochemical gradient to synthesize ATP (see Figs. 8-5 and 8-6). The proton gradient across the inner mitochondrial membrane drives rotation of the γ-subunit. The rotating mg-subunit physically changes the conformations of the α- and β-subunits, bringing together ADP and inorganic phosphate to make ATP. An antiporter in the inner membrane exchanges cytoplasmic ADP for ATP synthesized in the matrix (see Fig. 9-2A).
Mitochondria and Disease
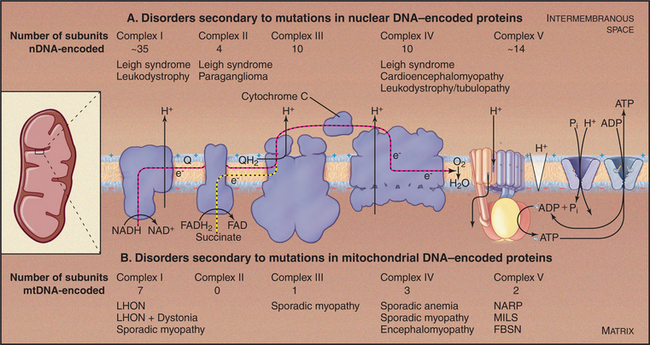
As expected from the central role of mitochondria in energy metabolism, mitochondrial dysfunction contributes to a remarkable diversity of human diseases (Fig. 19-6) including seizures, strokes, optic atrophy, neuropathy, myopathy, cardiomyopathy, hearing loss, and Type 2 diabetes mellitus. These disorders arise from mutations in genes for mitochondrial proteins encoded by both mitochondrial DNA (mtDNA) and nuclear DNA. More than half of the known disease-causing mutations are in genes for mitochondrial transfer RNAs.
Mutations in nuclear genes for mitochondrial proteins cause similar diseases (Fig. 19-6A). A mutation in one subunit of the protein import machinery (see Fig. 18-5), Tim8, causes a type of deafness.
Chloroplasts
Structure and Evolution of Photosynthesis Systems
Photosynthetic Bacteria and chloroplasts of algae and plants (Fig. 19-7) use chlorophyll to capture the remarkable amount of energy carried by single photons to boost electrons to an excited state. These high-energy electrons drive a chemiosmotic cycle to make NADPH and ATP, energy currency that is used by all cells. Photosynthetic organisms use ATP and the reducing power of NADPH to synthesize three-carbon sugar phosphates from carbon dioxide. Glycolytic reactions (Fig. 19-4) running backward use this three-carbon sugar phosphate to make six-carbon sugars and more complex carbohydrates for use as metabolic energy sources and structural components. Some Archaea, such as Halobacteria halobium, and some recently discovered Bacteria use a completely different light-driven pump lacking chlorophyll to generate a proton gradient to synthesize ATP. Retinol associated with bacteriorhodopsin absorbs light to drive proton transport (see Fig. 8-3).
Photosynthesis originated approximately 3.5 billion years ago in a Bacterium, most likely a gram-negative purple bacterium (see Fig. 2-4). These bacteria evolved components to assemble a transmembrane complex of proteins, pigments, and oxidation/reduction cofactors called a reaction center (Fig. 19-8). Reaction centers absorb light and initiate an electron transport pathway that pumps protons out of the cell. Such photosystems turn sunlight into electrical and chemical energy with 40% efficiency, better than any human-made photovoltaic cell. Given their alarming complexity and physical perfection, it is remarkable that photosystems emerged only a few hundred million years after the origin of life itself.
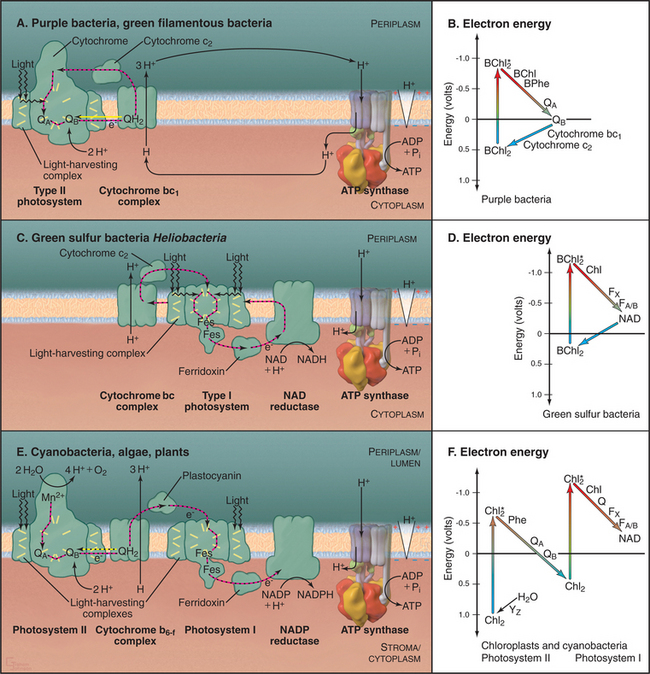
Broadly speaking, photosynthetic reaction centers of contemporary organisms can be divided into two different groups (Fig. 19-8). The reaction centers of purple bacteria and green filamentous bacteria utilize the pigment pheophytin and a quinone as the electron acceptor, similar to photosystem II of cyanobacteria and chloroplasts. The reaction centers of green sulfur bacteria and heliobacteria have iron-sulfur centers as electron acceptors, similar to photosystem I of cyanobacteria and chloroplasts.
Cyanobacteria are unique among Bacteria in that they have both types of photosystems as well as a manganese enzyme that splits water, releasing from two water molecules four electrons, four protons, and oxygen (Fig. 19-7E). Coupling this enzyme to photosynthesis was a pivotal event in the history of the earth, as this reaction is the source of most of the oxygen in the earth’s atmosphere.
Chloroplasts of eukaryotic cells arose from a symbiotic cyanobacterium (see Fig. 2-8). Much evidence indicates that this event occurred just once, giving all chloroplasts a common origin. However, to account for chloroplasts in organisms that diverged prior to the acquisition of chloroplasts, one must also postulate lateral transfer of chloroplasts from, for example, a green alga to Euglena. Less likely, but not ruled out conclusively, cyanobacteria may have colonized eukaryotic cells on up to three different occasions, giving rise to organelles that evolved into chloroplasts.
Chloroplasts have retained up to 250 original bacterial genes on circular genomes, whereas many bacte-rial genes were lost or moved to the nucleus of host eukaryotes. Chloroplast genomes encode subunits of many proteins responsible for photosynthesis and chloroplast division, ribosomal RNAs and proteins, and a complete set of tRNAs. Chloroplast proteins encoded by nuclear genes are transported posttranslationally into chloroplasts (Fig. 18-6) after their synthesis in cytoplasm.
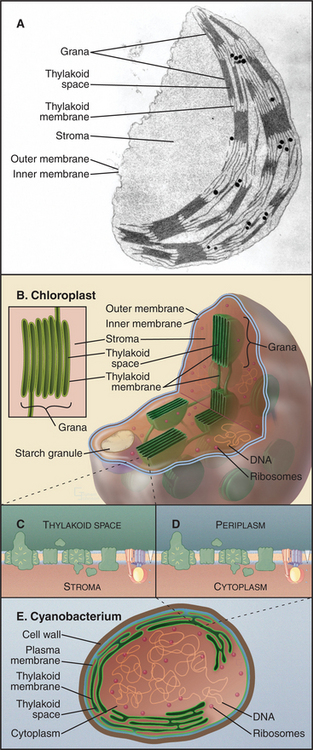
The organization of cyanobacterial membranes ex-plains the architecture of chloroplasts (Fig. 19-7C-E). In cyanobacteria, light-absorbing pigments, as well as protein complexes involved with electron transport and ATP synthesis, are concentrated in invaginations of the plasma membrane. The F1 domain of ATP synthase faces the cytoplasm, and the lumen of this membrane system is periplasmic. This internal membrane system remains in chloroplasts but is separated from the inner membrane (the former plasma membrane). These thylakoid membranes contain photosynthetic hardware and enclose the thylakoid membrane space. Like the bacterial plasma membrane, the chloroplast “inner membrane” is a permeability barrier, containing carriers for metabolites. The inner membrane surrounds the stroma, the cytoplasm of the original symbiotic bacterium, a protein-rich compartment devoted to synthesis of three-carbon sugar phosphates, chloroplast proteins, and all plant fatty acids. The stroma also houses the genomes and stores starch. The outer membrane, like the comparable bacterial and mitochondrial membranes, has large pore channels that allow free passage of metabolites.
Light and Dark Reactions
Photosynthetic mechanisms capture energy from photons to drive two types of reactions:
All photosynthetic systems use similar mechanisms to capture energy from photons (Fig. 19-8). Pigments associated with transmembrane proteins in photosynthetic reaction centers absorb photons and use the energy to boost electrons to a high-energy, excited state. Subsequent electron transfer reactions partition this energy in several steps to generate a proton gradient across the membrane. Generation of this proton electrochemical gradient and chemiosmotic produc-tion of ATP are similar to oxidative phosphorylation (Fig. 19-5).
Specific photosynthetic systems differ in the complexity of the hardware, the source of electrons, and the products (Fig. 19-8). Most photosynthetic bacteria use either a type I photosystem or a type II photosystem to create a proton gradient to synthesize ATP. Cyanobacteria and green plants use both types of reaction cen-ters in series to raise electrons to an energy sufficient to make NADPH in addition to ATP. These advanced systems also use water as the electron donor and produce molecular oxygen as a by-product.
Energy Capture and Transduction by Type II Photosystems and Photosystem II
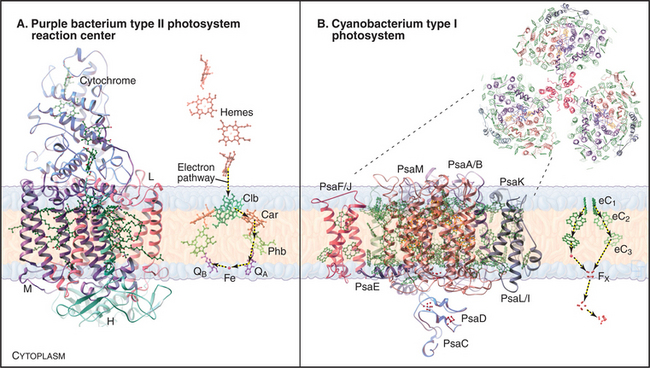
The reaction center from the purple bacterium Rhodopseudomonas viridis (Fig. 19-9A) is a model for the more complex photosystem II of cyanobacteria and chloroplasts. This bacterial reaction center consists of just four subunits. A cytochrome subunit on the periplasmic side of the membrane donates electrons. Two core subunits form a rigid transmembrane framework to bind 10 cofactors in orientations that favor transfer of high-energy electrons from two “special” bacteriochlorophylls through chlorophyll b and bacteriopheophytin b.
Photosynthesis begins with absorption of a photon by the special pair bacteriochlorophylls. Photons in the visible part of the spectrum are quite energetic, 40 to 80kcalmol−1, enough to make several ATPs. The purple bacterium reaction center absorbs relatively low-energy, 870-nm red light. The energy elevates an electron in the special pair bacteriochlorophylls to an excited state (Fig. 19-8B). In an organic solvent, the excited state would decay rapidly (109s−1), and the energy would dissipate as heat or emission of a less energetic photon by fluorescence or phosphorescence. However, reaction centers are optimized to transfer excited-state electrons rapidly and efficiently from the special pair bacteriochlorophylls to bacteriopheophytin (3 × 10−12s) and then to tightly bound quinone A (200 × 10−12s). Transfer is by quantum mechanical tunneling right through the protein molecule. Because the tunneling rate falls off quickly with distance, four redox centers must be spaced close together to allow an energetic electron to transfer across the lipid bilayer faster than spontaneous decay of the excited state.
On the cytoplasmic side of the membrane, two electrons transfer from quinone A to loosely bound quinone B (100 × 10−9s), where they combine with two protons to make a high-energy reduced quinone, QH2 (Fig. 19-8A). In purple bacteria, these cytoplasmic protons are taken up through water-filled channels in the reaction center, contributing to the proton gradient.
QH2 has a low affinity for the reaction center and diffuses in the hydrophobic core of the bilayer to the next component in the pathway, the chloroplast equivalent of the mitochondrial cytochrome bc1 complex III (Fig. 19-8A). As in mitochondria, passage of energetic electrons through this complex releases protons from QH2 on the periplasmic side of the membrane, adding to the electrochemical gradient. The electron circuit is completed by transfer of low-energy electrons from complex bc1 to a soluble periplasmic protein, cytochrome c2. Electrons then move to the cytochrome subunit of the reaction center, which supplies special pair chlorophylls with electrons for the photosynthetic reaction cycle.
The net result of this cycle is the conversion of the energy of two photons into transport of three protons to the periplasm. A diagram of the energy levels of the various intermediates in the cycle (Fig. 19-8B) shows how energy is partitioned after an electron is excited by a photon and then moves, step by step, through protein-associated redox centers back to the ground state.
The proton electrochemical gradient established by photosynthetic electron transfer reactions is used to drive an ATP synthase (see Fig. 8-5) similar to those of nonphotosynthetic prokaryotes and mitochondria.
Light Harvesting
Reaction center chlorophylls absorb light themselves, but both chloroplasts and bacteria increase the efficiency of light collection with proteins that absorb light and transfer the energy to a reaction center. Most of these light-harvesting complexes are small, transmembrane proteins that cluster around a reaction center, although some bacteria and algae also have soluble light-harvesting proteins. Transmembrane, light-harvesting proteins consist of a few α-helices associated with multiple chlorophyll and carotenoid pigments (Figs. 19-8A and C and 19-9B). The use of different pigments broadens the range of wavelengths absorbed. Multiple pigments increase the efficiency of photon capture. Leaves are green because chlorophylls and carotenoids absorb purple and blue wavelengths (<530nm) as well as red wavelengths (>620nm), reflecting only yellow-green wavelengths in between.
Energy Capture and Transduction by Photosystem I
The reaction centers of green sulfur bacteria and heliobacteria are similar to photosystem I of cyanobacteria and chloroplasts. Generation of a proton gradient by photosystem I has many parallels with photosystem II. Direct absorption of light or resonance energy transfer from surrounding light-harvesting complexes excites special pair chlorophylls in photosystem I (Fig. 19-8C-D). Excited-state electrons move rapidly within the reaction center from these chlorophylls through two accessory chlorophylls and to an iron-sulfur center. The pathway includes a quinone in cyanobacteria and chloroplasts. Electrons then move to the iron-sulfur center of a subunit on the cytoplasmic side of the membrane. The subsequent events in green sulfur bacteria and heliobacteria are still under investigation but are thought to include electron transfer by the soluble protein ferridoxin to an NAD reductase, followed by transfer by a lipid intermediate to cytochrome bc complex, and then back to the reaction center via a cytochrome c.
Oxygen-Producing Synthesis of NADPH and ATP by Dual Photosystems
Chloroplasts and cyanobacteria combine photosystem II and photosystem I in the same membrane to form a system capable of accepting low-energy electrons from the oxidation of water and producing both a proton gradient to drive ATP synthesis and reducing equivalents in the form of NADPH (Fig. 19-8E-F). Both photosystems are more elaborate in dual systems than in single systems. Although plant photosystem II, with more than 25 protein subunits, is much more complicated than is the homologous reaction center of purple bacteria, the arrangement of transmembrane helices and chlorophyll cofactors in the core of the plant reaction center is similar to the simple reaction center of purple bacteria.
Photosynthesis involves a tortuous electron transfer pathway powered at two way stations by absorption of photons. This process begins when the special pair chlorophylls of photosystem II are excited by direct aβ-sorption of light or by resonance energy transfer from surrounding light-harvesting complexes (Fig. 19-8E-F). Electrons come from splitting two waters into molecular oxygen and four protons. Excited-state electrons tunnel through the redox cofactors and combine with protons from the stroma (or cytoplasm in bacteria) to reduce quinone QB to QH2, a high-energy electron donor. QH2 diffuses to complex b6-f, the chloroplast equivalent of the mitochondrial bc1 complex. Passage of electrons through complex b6-f releases protons from QH2 into the thylakoid lumen (or bacterial periplasm), contributing to the proton gradient across the membrane.
Complex b6-f donates electrons from QH2 to photosystem I. Direct absorption of 680-nm light or resonance energy transfer from surrounding light-harvesting complexes boosts special pair chlorophyll electrons to a very high-energy, excited state (Fig. 19-8F). Excited-state electrons pass through chlorophyll and iron-sulfur centers of photosystem I to the iron-sulfur center of the redox protein, ferridoxin, on the cytoplasmic/stromal surface of the membrane. The enzyme NADP reductase combines electrons from ferridoxin with a proton to form NADPH, the final product of this tortuous electron transfer pathway powered at two way stations by absorption of photons. Uptake of stromal protons during NADPH formation contributes to the transmembrane proton gradient for the synthesis of ATP. Antiporters in the inner membrane exchange ATP for ADP, as in mitochondria.
Synthesis of Carbohydrates
ATP and NADPH produced by light reactions drive the unfavorable conversion of carbon dioxide into sugars. This is the first step in the earth’s annual production of about 1010 tons of carbohydrates by photosynthetic organisms. This process is very expensive, consuming three ATPs and two NADPHs for each carbon dioxide added to the five-carbon sugar ribulose 1,5-bisphosphate. The responsible enzyme, ribulose phosphate carboxylase (called RUBISCO), is the most abundant protein in the stroma and might be the most abundant protein on the earth. The products of combining the five-carbon sugar with carbon dioxide are two molecules of the three-carbon sugar 3-phosphoglycerate.
An antiporter in the inner chloroplast membrane exchanges 3-phosphoglycerate for inorganic phosphate, so 3-phosphoglycerate can join the glycolytic pathway in the cytoplasm (Fig. 19-4). Driven by this abundant supply of 3-phosphoglycerate, the glycolytic pathway runs backward to make six-carbon sugars, which are used to make disaccharides such as sucrose to nourish nonphotosynthetic parts of the plant, the glucose polymer starch to store carbohydrate, and cellulose for the extracellular matrix (see Figs. 3-25A and 32-12).
Peroxisomes
Peroxisomes are organelles bounded a single membrane (Fig. 19-10), named for their content of enzymes that produce and degrade hydrogen peroxide, H2O2. Oxidases produce H2O2 and peroxidases such as catalase break it down. Peroxisomes also contain diverse enzymes for the metabolism of lipids and other metabolites, including the β-oxidation of fatty acids and oxidation of bile acids and cholesterol. Peroxisomes lack nucleic acids, and there is no evidence that they arose from a bacterial ancestor. All peroxisomal proteins are encoded by nuclear genes, translated on cytoplasmic ribosomes, and then subsequently incorporated into peroxisomes (see Fig. 18-8).
Peroxisomes form in two different ways: de novo synthesis by budding from the endoplasmic reticulum and growth and division of preexisting peroxisomes (see Fig. 18-8). Cells that lack preexisting peroxisomes can form peroxisomes without a template by differentiation and budding of ER membranes. PEX3 and PEX16 target to the ER, where they recruit other peroxins to form a specialized domain that pinches off to form a nascent peroxisome. In addition to arising by outgrowth from the ER, new peroxisomes can form by fission of preexisting peroxisomes.
Defects in peroxisomal biogenesis cause a spectrum of lethal human diseases known as the peroxisomal biogenesis disorders (see Appendix 18-1). These diseases include Zellweger syndrome, neonatal adrenoleukodystrophy, infantile Refsum’s disease, and rhizomelic chondrodysplasia punctata. They are moderately rare, occurring in approximately 1 in 50,000 live births. Most patients with peroxisomal biogenesis disorders display no defect in peroxisome membrane synthesis or import of peroxisomal membrane proteins, but they do have mild-to-severe defects in matrix protein import. However, in rare cases, patients lack peroxisome membranes altogether. Studies of both yeast pex mutants and cells from patients with peroxisomal biogenesis disorders have provided clues regarding peroxisome biogenesis (see Fig. 18-7).
Blankenship RE, Hartman H. The origin and evolution of oxygenic photosynthesis. Trends Biochem Sci. 1998;23:94-97.
Cecchini G. Function and structure of complex II of the respiratory chain. Annu Rev Biochem. 2003;72:77-109.
Deisenhofer J, Michel H. The photosynthetic reaction center from the purple bacterium Rhodopseudomonas viridis. Science. 1989;245:1463-1473.
Frey TG, Mannella CA. The internal structure of mitochondria. Trends Biochem Sci. 2000;25:319-324.
Gray MW, Burger G, Lang BF. Mitochondrial evolution. Science. 1999;283:1476-1481.
Hosler JP, Ferguson-Miller S, Mills DA. Energy transduction: Proton transfer through the respiratory complexes. Annu Rev Biochem. 2006;75:165-187.
Iwata S, Barber J. Structure of photosystem II and molecular architecture of the oxygen-evolving centre. Curr Opin Struct Biol. 2004;14:447-453.
Lazarow PB. Peroxisome biogenesis: Advances and conundrums. Curr Opin Cell Biol. 2003;15:489-497.
Lowell BB, Shulman GI. Mitochondrial dysfunction and type 2 diabetes. Science. 2004;307:384-387.
Meeusen SL, Nunnari J. How mitochondria fuse. Curr Opin Cell Biol. 2005;17:389-394.
Moser CC, Keske JM, Warncke K, et al. Nature of biological electron transfer. Nature. 1992;355:796-802.
Osteryoung KW, Nunnari J. The division of endosymbiotic organelles. Science. 2003;302:1698-1704.
Rhee K-H. Photosystem II: The solid structural era. Annu Rev Biophys Biomol Struct. 2001;30:307-328.
Rhee K-H, Morris EP, Barber J, Kuhlbrandt W. Three-dimensional structure of the plant photosystem II reaction centre at 8 Å resolution. Nature. 1998;396:283-286.
Rutherford AW, Boussac A. Water photolysis in biology. Science. 2004;303:1782-1784.
Scheffler IE. Mitochondria. New York: Wiley & Sons, 1999.
Schubert W-D, Klukas O, Krauss N, et al. Photosystem I of Synechococcus elongatus at 4 Å resolution: Comprehensive structure analysis. J Mol Biol. 1997;272:741-769.
Schultz BE, Chan SI. Structures and proton-pumping strategies of mitochondrial respiratory enzymes. Annu Rev Biophys Biomol Struct. 2001;30:23-65.
Smith JL, Zhang H, Yan J, et al. Cytochrome bc complexes: A common core of structure and function surrounded by diversity in the outlying provinces. Curr Opin Struct Biol. 2004;14:432-439.
Tielens AGM, Rotte C, van Hellemond JJ, Martin W. Mitochondria as we don’t know them. Trends Biochem Sci. 2002;27:564-572.
Wallace DC. Mitochondrial diseases in man and mouse. Science. 1999;283:1482-1488.
Wanders RJ, Waterham HR. Biochemistry of mammalian peroxisomes revisited. Annu Rev Biochem. 2006;75:295-332.
Wittenhagen LM, Kelley SO. Impact of disease-related mitochondrial mutations on tRNA structure and function. Trends Biochem Sci. 2003;28:605-611.
[/level-membership-for-basic-science-category][not-level-membership-for-basic-science-category]
CHAPTER 19 Mitochondria, Chloroplasts, Peroxisomes
This chapter considers three organelles formed by posttranslational import of proteins synthesized in the cytoplasm. Mitochondria and chloroplasts both arose from endosymbiotic bacteria, two singular events that occurred about one billion years apart. Both mitochondria and chloroplasts retain remnants of those prokaryotic genomes but depend largely on genes that were transferred to the nucleus of the host eukaryote. Both organelles brought biochemical mechanisms that allow their eukaryotic hosts to acquire and utilize energy more efficiently. In oxidative phosphorylation by mitochondria and photosynthesis by chloroplasts, energy from the breakdown of nutrients or from absorption of photons is used to energize electrons. As these electrons tunnel through transmembrane proteins, energy is partitioned off to create proton gradients. These proton gradients drive the rotary ATP synthase (see Fig. 8-5) to make adenosine triphosphate (ATP), which is used as energy currency to power the cell. Peroxisomes contain no genes and depend entirely on nuclear genes to encode their proteins. Their evolutionary origins are obscure. Peroxisomes contain enzymes that catalyze oxidation reactions that are essential for normal human physiology. Patients who lack peroxisomes have severe neural defects.
Mitochondria
Evolution and Structure of Mitochondria
Mitochondria (Fig. 19-1) arose about 2 billion years ago when a Bacterium fused with an archaeal cell or established a symbiotic relationship with a primitive eukaryotic cell (see Fig. 2-5 and associated text). The details are not preserved in the fossil record, but the bacterial origins of mitochondria are apparent in their many common features (Fig. 19-2). The closest extant relatives of the Bacterium that gave rise to mitochondria are Rickettsia, aerobic α-proteobacteria with a genome of 1.1 megabase pairs. These intracellular pathogens cause typhus and Rocky Mountain spotted fever. However, some evidence argues that the actual progenitor bacterium had the genes required for both aerobic and anaerobic metabolism.
As primitive eukaryotes diverged from each other, most of the bacterial genes were lost or moved to the nuclei of the host eukaryotes. The pace of the gene transfer to the nucleus varied considerably depending on the species, but all known mitochondria retain some bacterial genes. A very few eukaryotes, such as Entamoeba, that branched well after their ancestors acquired mitochondria lost the organelle, leaving behind a few mitochondrial genes in the nucleus.
Chromosomes of contemporary mitochondria vary in size from 366,924 base pairs (bp) in the plant Arabidopsis to only 5966bp in Plasmodium. These small, usually circular genomes encode RNAs and proteins that are essential for mitochondrial function, including some subunits of proteins responsible for adenosine triphosphate (ATP) synthesis. The highly pared-down human mitochondrial genome with 16,569bp encodes only 13 mitochondrial membrane proteins, two ribosomal RNAs, and just enough tRNAs (22) to translate these genes. The number of proteins encoded by other mitochondrial genomes ranges from just 3 in Plasmodium to 97 in a protozoan. Nuclear genes encode the other 600 to 1000 mitochondria proteins, including those required to synthesize proteins in the matrix. All mitochondrial proteins that are encoded by nuclear genes are synthesized in the cytoplasm and subsequently imported into mitochondria (see Figs. 18-2 and 18-3).
Mitochondria consist of two membrane-bounded compartments, one inside the other (Fig. 19-2). The outer membrane surrounds the intermembranous space. The inner membrane surrounds the matrix. Each membrane and compartment has a distinct protein composition and functions. Porins in the outer membrane provide channels for passage of molecules of less than 5000D, including most metabolites required for ATP synthesis. The highly impermeable inner membrane is specialized for converting energy provided by breakdown of nutrients in the matrix into ATP. Four complexes (I to IV) of integral membrane proteins use the transport of energetic electrons to create a gradient of protons across the inner membrane. The F1F0 ATP synthase (see Fig. 8-5) utilizes the proton gradient to synthesize ATP. The area of inner membrane available for these reactions is increased by folds called cristae that vary in number and shape depending on the species, tissue, and metabolic state. Cristae may be tubular or flattened sacs. Contacts between the inner and outer membranes are sites of protein import (see Fig. 18-4). Proteins in the intermembranous space participate in ATP synthesis but, when released into the cytoplasm, trigger programmed cell death (see Fig. 46-15).
Biogenesis of Mitochondria
Mitochondria grow by importing most of their proteins from the cytoplasm and by internal synthesis of some proteins and replication of the genome (Fig. 19-3). Targeting and sorting signals built into the mitochondrial proteins that are synthesized in the cytoplasm direct them to their destinations (see Fig. 18-4).
Similar to cells, mitochondria divide, but unlike most cells, they also fuse with other mitochondria. These fusion and division reactions were first observed nearly one hundred years ago. Now it is appreciated that a balance between ongoing fusion and division determines the number of mitochondria within a cell. Both fusion and division depend on proteins with guanosine triphosphatase (GTPase) domains related to dynamin (see Fig. 22-11). In fact, eukaryotes might have acquired their dynamin genes from the bacterium that became mitochondria.
One dynamin-related GTPase is required for division of mitochondria. This GTPase self-assembles into spirals that appear to pinch mitochondria in two. During apoptosis (see Chapter 46), this GTPase also participates in the fragmentation of mitochondria.
Synthesis of ATP by Oxidative Phosphorylation
Mitochondria use energy extracted from the chemical bonds of nutrients to generate a proton gradient across the inner membrane. This proton gradient drives the F1F0 ATP synthase to synthesize ATP from ADP and inorganic phosphate. Enzymes in the inner membrane and matrix cooperate with pumps, carriers, and electron transport proteins in the inner membrane to move electrons, protons, and other energetic intermediates across the impermeable inner membrane. This is a classic chemiosmotic process (see Fig. 11-1).
Mitochondria receive energy-yielding chemical intermediates from two ancient metabolic pathways, glycolysis and fatty acid oxidation (Fig. 19-4), that evolved in the common ancestor of living things. Both pathways feed into the equally ancient citric acid cycle of energy-yielding reactions in the mitochondrial matrix:
Breakdown of acetyl-CoA during one turn of the citric acid cycle produces three molecules of NADH, one molecule of FADH2, and two molecules of carbon dioxide. Energetic electrons donated by NADH and FADH2 drive an electron transport pathway in the inner mitochondrial membrane that powers a chemiosmotic cycle to produce ATP (Fig. 19-5). Electrons use two routes to pass through three protein complexes in the inner mitochondrial membrane. Starting with NADH, electrons pass through complex I to complex III to complex IV. Electrons from FADH2 pass through complex II to complex III to complex IV. Along both routes, energy is partitioned off to transfer multiple protons (at least 10 electrons per NADH oxidized) across the inner mitochondrial membrane from the matrix to the inner membrane space. The resulting electrochemical gradient of protons drives ATP synthesis (see Fig. 8-5).

Figure 19-5 chemiosmotic cycle of the respiratory electron transport chain and atp synthase. a, left panel, A mitochondrion for orientation. Right panel, The electron transport system of the inner mitochondrial membrane. Note the pathway of electrons through the four complexes (red and yellow arrows) and the sites of proton translocation between the matrix to the intermembranous space (black arrows). The stoichiometry is not specified, but at the last step, four electrons are required to reduce oxygen to water. ATP synthase uses the electrochemical proton gradient produced by the electron transport reactions to drive ATP synthesis. B, The available atomic structures of the electron transport chain are shown. In the cytochrome bc1 complex III, the 3 of 11 mitochondrial subunits used by bacteria are shown as ribbon models. The supporting subunits found in mitochondria are shown as cylinders. The four subunits of complex IV encoded by the mitochondrial genome are shown as ribbon models. They form the functional core of the complex, which is supported by additional subunits shown as cylinders. See Figure 8-4 for further details of ATP synthase (complex V).
(B, Images of complex III and complex IV courtesy of M. Saraste, European Molecular Biology Laboratory, Heidelberg, Germany. Reference: Zhang Z, Huang L, Schulmeister VM, et al: Electron transfer by domain movement in cytochrome bc1. Nature 392:677–684, 1998. PDB file: 1BCC. Reference: Yoshikawa S, Shinzawα-Itoh K, Nakashima R, et al: Redox-coupled crystal structural changes in bovine heart cytochrome c oxidase. Science 280:1723–1729, 1998. PDB file: 2OCC.)
This process is called oxidative phosphorylation, since molecular oxygen is the sink for energy-bearing electrons at the end of the pathway and since the reactions add phosphate to ADP. Eukaryotes that live in environments with little or no oxygen use other acceptors for these electrons and produce nitrite, nitric oxide, or other reduced products rather than water. Oxidative phosphorylation is understood in remarkable detail, thanks to atomic structures of ATP synthase and three of the four electron transfer complexes. Nuclear genes encode most of the protein subunits of these complexes, but mitochondrial genes are responsible for a few key subunits.
Bacteria and mitochondria share homologous proteins for the key steps in oxidative phosphorylation (Fig. 19-2), but the machinery in mitochondria is usually more complex. Thus, bacteria are useful model systems with which to study the common mechanisms. Plasma membranes of bacteria and inner membranes of mitochondria have equivalent components, and the bacterial cytoplasm corresponds to the mitochondrial matrix (Fig. 19-2).
Energy enters this pathway in the form of electrons that are produced when NADH is oxidized to NAD+, releasing one H+ and two electrons (Fig. 19-5). If the proton and electrons were to combine immediately with oxygen, their energy would be lost as heat. Instead, these high-energy electrons are separated from the protons and then passed along the electron transport pathway before finally recombining to reduce molecular oxygen to form water. Along the pathway, electrons associate transiently with a series of oxidation/reduction acceptors, generally metal ions associated with organic cofactors, such as hemes in cytochromes and iron-sulfur centers (2Fe2S) and copper centers in complex IV. Electrons move along the transport pathway at rates of up to 1000s−1. To travel at this rate through a transmembrane protein complex spanning a 35-nm lipid bilayer, at least three redox cofactors are required in each complex, because the efficiency of quantum mechanical tunneling of electrons between redox cofactors falls off rapidly with distance. Two cofactors, even with optimal orientation, would be too slow.
The first component of the electron transport pathway is called complex I
[/not-level-membership-for-basic-science-category]