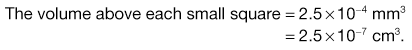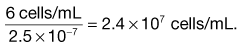[level-membership-for-basic-science-category]
Fundamentals of microbiology
Geoffrey W. Hanlon
Chapter contents
Microscopy and staining of bacteria
Growth and reproduction of bacteria
Handling and storage of microorganisms
Isolation of pure bacterial cultures
Key points
Introduction
Microorganisms are ubiquitous in nature and are vital components in the cycle of life. The majority are free-living organisms growing on dead or decaying matter whose prime function is the turnover of organic materials in the environment. Pharmaceutical microbiology, however, is concerned with the relatively small group of biological agents that cause human disease, spoil prepared medicines or can be used to produce compounds of medical interest.
In order to understand microorganisms more fully, living organisms of similar characteristics have been grouped together into taxonomic units. The most fundamental division is between prokaryotic and eukaryotic cells, which differ in a number of respects (Table 13.1) but particularly in the arrangement of their nuclear material. Eukaryotic cells contain chromosomes, which are separate from the cytoplasm and contained within a limiting nuclear membrane, i.e. they possess a true nucleus. Prokaryotic cells do not possess a true nucleus and their nuclear material is free within the cytoplasm, although it may be aggregated into discrete areas called nuclear bodies. Prokaryotic organisms make up the lower forms of life and include Eubacteria and Archaeobacteria. Eukaryotic cell types embrace all the higher forms of life, of which only the fungi will be dealt with in this chapter.
Table 13.1
| Structure | Prokaryotes | Eukaryotes |
| Cell wall structure | Usually contains peptidoglycan | Peptidoglycan absent |
| Nuclear membrane | Absent | Present. Possess a true nucleus |
| Nucleolus | Absent | Present |
| Number of chromosomes | One | More than one |
| Mitochondria | Absent | Present |
| Mesosomes | Present | Absent |
| Ribosomes | 70S | 80S |
One characteristic shared by all microorganisms is the fact that they are small; however, it is a philosophical argument whether all infectious agents can be regarded as living. Some are little more than simple chemical entities incapable of any free-living existence. Viroids, for example, are small circular, single-stranded RNA molecules not complexed with protein. One particularly well-studied viroid has only 359 nucleotides (one-10th the size of the smallest known virus) and yet causes a disease in potatoes. Prions are small, self-replicating proteins devoid of any nucleic acid. The prion associated with Creutzfeld–Jakob disease in humans, scrapie in sheep and bovine spongiform encephalitis in cattle has only 250 amino acids and is highly resistant to inactivation by normal sterilization procedures.
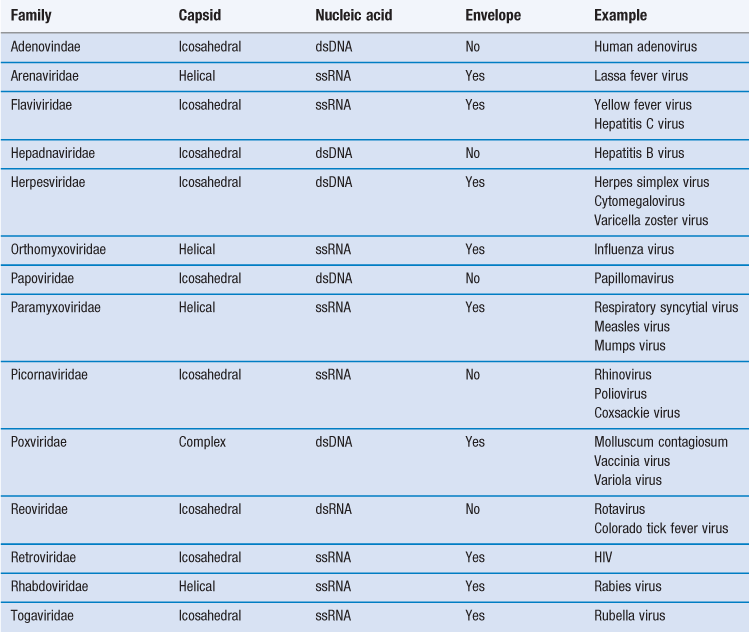
Viruses are more complex than viroids or prions, possessing both protein and nucleic acid. Despite being among the most dangerous infectious agents known, they are still not regarded as living. Table 13.2 shows the major groups of viruses infecting humans.
Viruses
Viruses are obligate intracellular parasites with no intrinsic metabolic activity, being devoid of ribosomes and energy-producing enzyme systems. They are thus incapable of leading an independent existence and cannot be cultivated on cell-free media, no matter how nutritious. The size of human viruses ranges from the largest poxviruses, measuring about 300 nm, to the picornaviruses, such as the poliovirus which is approximately 20 nm. When one considers that a bacterial coccus measures 1000 nm in diameter, it can be appreciated that only the very largest virus particles may be seen under the light microscope, and electron microscopy is required for visualizing the majority. It will also be apparent that few of these viruses are large enough to be retained on the 200 nm (0.2 µm) membrane filters used to sterilize thermolabile liquids.
Viruses consist of a core of nucleic acid (either DNA as in vaccinia virus or RNA as in poliovirus) surrounded by a protein shell or capsid. Most DNA viruses have linear, double-stranded DNA but in the case of the parvovirus it is single stranded. The majority of RNA-containing viruses contain one molecule of single-stranded RNA, although in reoviruses it is double stranded. The protein capsid comprises 50–90% of the weight of the virus and, as nucleic acid can only synthesize approximately 10% its own weight of protein, the capsid must be made up of a number of identical protein molecules. These individual protein units are called capsomeres and are not in themselves symmetrical but are arranged around the nucleic acid core in characteristic symmetrical patterns. Additionally, many of the larger viruses possess a lipoprotein envelope surrounding the capsid arising from the membranes within the host cell. In many instances the membranes are virus modified to produce projections outwards from the envelope, such as haemagglutinins or neuraminidase. The enveloped viruses are often called ether sensitive, as ether and other organic solvents may dissolve the membrane.
The arrangement of the capsomeres can be of a number of types.
Reproduction of viruses
Because viruses have no intrinsic metabolic capability, they require the functioning of the host cell machinery in order to manufacture and assemble new virus particles. It is this intimate association between the virus and its host that makes the treatment of viral infections so complex. Any chemotherapeutic approach which damages the virus will almost inevitably cause injury to the host cells and hence lead to side-effects. An understanding of the life cycle of the virus is vital in determining suitable target sites for antiviral chemotherapy. The replication of viruses within host cells can be broken down into a number of stages.
Adsorption to host cell
The first step in the infection process involves virus adsorption on to the host cell. This usually occurs via an interaction between protein or glycoprotein moieties on the virus surface with specific receptors on the host cell outer membrane. Different cells possess receptors for different viruses. For example, the human immunodeficiency virus (HIV) possesses two proteins involved in adsorption to T lymphocytes and these are known as gp41 and gp120. There are receptors on the lymphocyte surface to which HIV will bind. The main receptor is CD4 to which the gp120 protein attaches. Other receptors are CXCR4 or CCR5 to which the gp41 protein binds. Both attachments are necessary for infection and lead to conformational changes in the HIV envelope proteins, resulting in membrane fusion.
Penetration
Enveloped viruses fuse the viral membrane with the host cell membrane and release the nucleocapsid directly into the cytoplasm. Naked virions generally penetrate the cell by phagocytosis. Bacteriophages are viruses which specifically attack bacteria and these inject their DNA into the host cell while the rest of the virus remains on the outside.
Uncoating
In this stage the capsid is removed as a result of attack by cellular proteases and this releases the nucleic acid into the cytoplasm. These first three stages are similar for both DNA and RNA viruses.
Nucleic acid and protein synthesis
The detailed mechanisms by which DNA- and RNA-containing viruses replicate inside the cell are outside the scope of this chapter and the reader is referred to the bibliography for further information. After nucleic acid replication, early viral proteins are produced, the function of which is to switch off host cell metabolic activity and direct the activities of the cell towards the synthesis of proteins necessary for the assembly of new virus particles.
Assembly of new virions
Again, there are differences in the detail of how the viruses are assembled within the host cell, but construction of new virions occurs at this stage and up to 100 new virus particles may be produced per cell.
Release of virus progeny
The newly formed virus particles may be liberated from the cell as a burst, in which case the host cell ruptures and dies. Infection with influenza virus results in a lytic response. Alternatively, the virions may be released gradually from the cell by budding of the host cell plasma membrane. These are often called ‘persistent’ infection; an example being hepatitis B.
Latent infections
In some instances a virus may enter a cell but not go through the replicative cycle outlined above and the host cell may be unharmed. The genome of the virus is conserved and may become integrated into the host cell genome where it may be replicated along with the host DNA during cell division. At some later stage the latent virus may become reactivated and progress through a lytic phase, causing cell damage/death and the release of new virions. Examples of this type of infection are those which occur with the herpes simplex viruses associated with cold sores, genital herpes and also chickenpox where the dormant virus may reactivate to give shingles later in life.
Oncogenic viruses
Oncogenic viruses have the capacity to transform the host cell into a cancer cell. In some cases this may lead to relatively harmless, benign growths, such as warts caused by papovavirus, but in other cases more severe, malignant tumours may arise. Cellular transformation may result from viral activation or mutation of normal host genes, called protooncogenes, or the insertion of viral oncogenes.
Bacteriophages
Bacteriophages (phages) are viruses that attack bacteria but not animal cells. It is generally accepted that the interaction between phage and bacterium is highly specific, and there is probably at least one phage for each species of bacterium. In many cases the infection of a bacterial cell by a phage results in lysis of the bacterium; such phages are termed virulent. Some phages, however, can infect a bacterium without causing lysis. In this case the phage DNA becomes incorporated within the bacterial genome. The phage DNA can then be replicated along with the bacterial cell DNA; this is then termed a prophage. Bacterial cells carrying a prophage are called lysogenic and phages capable of inducing lysogeny are called temperate. Occasionally some of the prophage genes may be expressed and this will confer on the bacterial cell the ability to produce new proteins. The ability to produce additional proteins as a result of prophage DNA is termed lysogenic conversion.
The discovery of bacteriophages in the early 20th century is attributed to two workers, Frederick Twort and Felix d’Herelle. In 1896 Ernest Hankin had made an observation that the waters of the Ganges River possessed antibacterial properties which may have led to a reduction in cases of dysentery and cholera in the areas surrounding the river. Twort and d’Herelle independently came to the conclusion that this effect must be due to a virus. Twort did not carry on with his research but d’Herelle quickly established the potential of bacteriophages in antibacterial therapy 10 years before the advent of antibiotics. It was the discovery of penicillin by Alexander Fleming in 1928 that led to the demise of bacteriophage therapy but interest is now increasing again due to the emergence of antibiotic-resistant strains of bacteria.
Archaeobacteria
Archaeobacteria are a fascinating group of prokaryotic microorganisms that are frequently found living in hostile environments. They differ in a number of respects from Eubacteria, particularly in the composition of their cell walls. They comprise methane producers, sulfate reducers, halophiles and extreme thermophiles. However, they are of little significance from a pharmaceutical or clinical standpoint and so will not be considered further.
Eubacteria
Eubacteria constitute the major group of prokaryotic cells that have pharmaceutical and clinical significance. They include a diverse range of microorganisms, from the primitive parasitic rickettsias that share some of the characteristics of viruses, through the more typical free-living bacteria to the branching, filamentous actinomycetes, which at first sight resemble fungi rather than bacteria.
Atypical bacteria
Rickettsiae
The family Rickettsiaceae includes three clinically important genera, Rickettsia, Coxiella and Bartonella. Although these are prokaryotic cells, they differ from most other bacteria both in their structure and in the fact that the majority of species lead an obligate intracellular existence. This means that, with a few exceptions, they cannot be grown on cell-free media, although unlike many viruses they do possess some independent enzymes. They have a pleomorphic appearance, ranging from coccoid through to rod-shaped cells; multiplication is by binary fission. Their cell wall composition bears similarities to that of Gram-negative bacteria (see later in this chapter) and in general they stain this way. The genus Rickettsia has a number of species that give rise to human diseases, in particular epidemic typhus (R. prowazekii), murine typhus (R. typhi) and spotted fevers (various species). These are characterized by transmission via insect vectors, particularly mites, ticks, fleas and lice.
The mode of transmission by these vectors varies depending upon the insect concerned. In the case of lice and fleas, the microorganisms multiply within the insect and get into the faeces. These insects then colonize humans and transmit the microorganism when the faeces or the insect itself is crushed onto the skin. No bite is necessary and the faeces may also be inhaled. Mites and ticks pick up the microorganism when they take a blood meal from an infected animal. They then pass on the infection to humans when they accidentally bite us.
Coxiella burnetii is the only species in the genus Coxiella and this gives rise to a disease called Q fever. Although the source of the disease is infected animals, usually no insect vector is involved and the most common route of transmission is by inhalation of infected dust. Bartonella quintana is the causative agent of trench fever which, as the name suggests, occurs typically under conditions of war and deprivation. Each of the infections described here can be treated with doxycycline, although the duration of therapy may vary depending upon the nature of the disease and its severity.
Chlamydiae
These are obligate intracellular parasitic bacteria that possess some independent enzymes but lack the ability to generate ATP. Two cellular forms are identified: a small (0.3 µm) highly infectious elementary body which, after infection, enlarges to give rise to the replicative form called the initial or reticulate body (0.8–1.2 µm). These divide by binary fission within membrane-bound vesicles in the cytoplasm of infected cells. Insect vectors are not required for the transmission of infection. Chlamydiae lack peptidoglycan in their cell walls and have weak Gram-negative characteristics.
Chlamydia trachomatis is the most important member of the group, being responsible for the disease trachoma, characterized by inflammation of the eyelids, which can lead to scarring of the cornea. This is the most common cause of infectious blindness worldwide. It is estimated that 400 million people are infected, with at least 6 million totally blind. The same species is also recognized as one of the major causes of sexually transmitted disease. C. psittaci and C. pneumoniae are responsible for respiratory tract infections. Chlamydial infections are responsive to treatment by tetracyclines, either topical or systemic as appropriate.
Mycoplasmas
The mycoplasmas are a group of very small (0.3–0.8 µm) prokaryotic microorganisms that are capable of growing on cell-free media but which lack cell walls. The cells are surrounded by a double-layered plasma membrane that contains substantial amounts of phospholipids and sterols. This structure has no rigidity owing to the absence of peptidoglycan, and so the cells are susceptible to osmotic lysis. The lack of peptidoglycan is also the reason for these bacteria being resistant to the effects of cell wall-acting antibiotics such as the penicillins, and also the enzyme lysozyme. Members of this group are pleomorphic, varying in shape from coccoid to filamentous. Most are facultative anaerobes capable of growth at 35 °C, and on solid media produce colonies with a characteristic ‘fried egg’ appearance. They contain a number of genera, of which the most important from a clinical point of view are Mycoplasma and Ureaplasma. M. pneumoniae is a major cause of respiratory tract infections in children and young adults, whereas U. urealyticum has been implicated in non-specific genital infections. Despite being resistant to the β-lactam antibiotics, these infections can be effectively treated using either tetracyclines or erythromycin.
Actinomycetes
Many of the macroscopic features of the actinomycetes are those that are more commonly found among the filamentous fungi but they are indeed prokaryotic cells. They are a diverse group of Gram-positive bacteria morphologically distinguishable from other bacteria because they have a tendency to produce branching filaments and reproductive spores. Nocardia contain a number of species that have been shown to be pathogenic to humans, but they occur principally in tropical climates. Reproduction in this genus is by fragmentation of the hyphal strands into individual cells, each of which can form a new mycelium. The genus Streptomyces contains no human pathogens but most species are saprophytic bacteria found in the soil. They are aerobic microorganisms producing a non-fragmenting, branching mycelium that may bear spores. The reason for their pharmaceutical importance is their ability to produce a wide range of therapeutically useful antibiotics, including streptomycin, chloramphenicol, oxytetracycline, erythromycin and neomycin.
Typical bacteria
Shape, size and aggregation
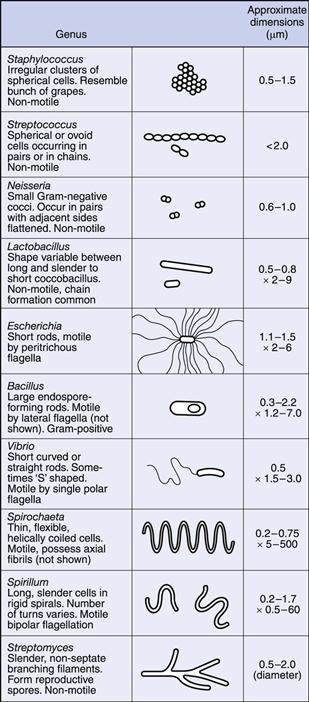
Bacteria occur in a variety of shapes and sizes, determined not only by the nature of the organisms themselves but also by the way in which they are grown (Fig. 13.1). In general, bacterial dimensions lie in the range 0.75–5 µm. The most common shapes are the sphere (coccus) and the rod (bacillus).
Some bacteria grow in the form of rods with a distinct curvature, e.g. vibrios are rod-shaped cells with a single curve resembling a comma, whereas a spirillum possesses a partial rigid spiral; spirochaetes are longer and thinner, exhibit a number of turns and are also more flexible. Rod-shaped cells occasionally grow in the form of chains but this is dependent upon growth conditions rather than being a characteristic of the species.
Cocci, however, show considerable variation in aggregation, which is characteristic of the species. The plane of cell division and the strength of adhesion of the cells determine the extent to which they aggregate after division. Cocci growing in pairs are called diplococci, those in four are tetrads and groups of eight are sarcina. If a chain of cells is produced resembling a string of beads this is termed a streptococcus, whereas an irregular cluster similar in appearance to a bunch of grapes is called a staphylococcus. In many cases this is sufficiently characteristic to give rise to the name of the bacterial genus, e.g. Staphylococcus aureus, Streptococcus pneumoniae.
Anatomy
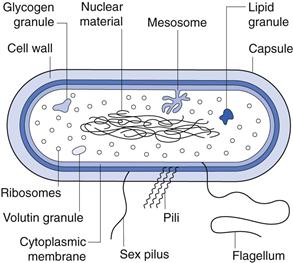
Figure 13.2 is a diagrammatic representation of a typical bacterial cell. The various components are described below.
Capsule.
Many bacteria produce extracellular polysaccharides, which may take the form of either a discrete capsule firmly adhered to the cell or a more diffuse layer of slime. Not all bacteria produce a capsule and even those that can will only do so under certain circumstances. Many encapsulated pathogens, when first isolated, give rise to colonies on agar which are smooth (S) but subculturing leads to the formation of rough colonies (R). This S to R transition is due to loss in capsule production. Reinoculation of the R cells into an animal results in the resumption of capsule formation, indicating that the capacity has not been lost irrevocably.
The function of the capsule is generally regarded as protective, as encapsulated cells are more resistant to disinfectants, desiccation and phagocytic attack. In some organisms, however, it serves as an adhesive mechanism; for example, Streptococcus mutans is an inhabitant of the mouth that metabolizes sucrose to produce a polysaccharide capsule enabling the cell to adhere firmly to the teeth. This is the initial step in the formation of dental plaque, which is a complex array of microorganisms and organic matrix that adheres to the teeth and ultimately leads to decay. The substitution of sucrose by glucose prevents capsule formation and hence eliminates plaque.
A similar picture emerges with Staph. epidermidis. This bacterium forms part of the normal microflora of the skin and was originally thought of as non-pathogenic. With the increased usage of indwelling medical devices coagulase-negative staphylococci, in particular Staph. epidermidis, have emerged as the major cause of device-related infections. The normal microbial flora has developed the ability to produce extracellular polysaccharide, which enables the cells to form resistant biofilms attached to the devices. These biofilms are very difficult to eradicate and have profound resistance to antibiotics and disinfectants. It is now apparent that the dominant mode of growth for aquatic bacteria is not planktonic (free swimming) but sessile, i.e. attached to surfaces and covered with protective extracellular polysaccharide or glycocalyx.
Cell wall.
Bacteria can be divided into two broad groups by the use of the Gram-staining procedure (see later in this chapter for details), which reflects differences in cell wall structure. The classification is based upon the ability of the cells to retain the dye methyl violet after washing with a decolourizing agent such as absolute alcohol. Gram-positive cells retain the stain whereas Gram-negative cells do not. As a very rough guide, the majority of small rod-shaped cells are Gram negative. Most large rods, such as the Bacillaceae, lactobacilli and actinomycetes, are Gram positive. Similarly, most cocci are Gram positive, although there are notable exceptions, such as the Neisseriaceae.
Bacteria are unique in that they possess peptidoglycan in their cell walls. This is a complex molecule with repeating units of N-acetylmuramic acid and N-acetylglucosamine (Fig. 13.3). This extremely long molecule is wound around the cell and crosslinked by polypeptide bridges to form a structure of great rigidity. The degree and nature of crosslinking vary between bacterial species. Crosslinking imparts to the cell its characteristic shape and has principally a protective function. Peptidoglycan (also called murein or mucopeptide) is the site of action of a number of antibiotics, such as penicillin, bacitracin, vancomycin and cycloserine. The enzyme lysozyme is also capable of hydrolysing the β–1–4 linkages between N-acetylmuramic acid and N-acelylglucosamine.
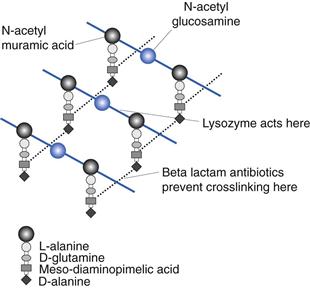
Fig. 13.3 Peptidoglycan.
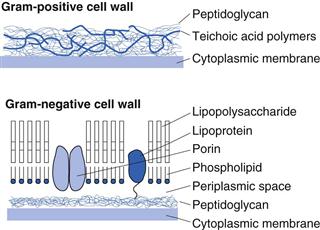
Figure 13.4 shows simplified diagrams of a Gram-positive and a Gram-negative cell wall. The Gram-positive cell wall is much simpler in layout, containing peptidoglycan interspersed with teichoic acid polymers. These latter are highly antigenic but do not provide structural support. Functions attributed to teichoic acids include the regulation of enzyme activity in cell wall synthesis, sequestration of essential cations, cellular adhesion and mediation of the inflammatory response in disease. In general, proteins are not found in Gram-positive cell walls. Gram-negative cell walls are more complex, comprising a much thinner layer of peptidoglycan surrounded by an outer bilayered membrane. This outer membrane acts as a diffusional barrier and is the main reason why many Gram-negative cells are much less susceptible to antimicrobial agents than are Gram-positive cells.
The lipopolysaccharide component of the outer membrane can be shed from the wall upon cell death. It is a highly heat-resistant molecule known as endotoxin, which has a number of toxic effects on the human body, including fever, shock and even death. For this reason it is important that solutions for injection or infusion are not just sterile but are also free from endotoxins.
Cytoplasmic membrane.
The cytoplasmic membranes of most bacteria are very similar and are composed of protein, lipids, phospholipids and a small amount of carbohydrate. The components are arranged into a bilayer structure with a hydrophobic interior and a hydrophilic exterior. The cytoplasmic membrane has a variety of functions.
• It serves as an osmotic barrier.
• It is selectively permeable and is the site of carrier-mediated transport.
• It is the site of ATP generation and cytochrome activity.
The cytoplasmic membrane has very little tensile strength and the internal hydrostatic pressure of up to 20 bar forces it firmly against the inside of the cell wall. Treatment of bacterial cells with lysozyme may remove the cell wall and, as long as the conditions are isotonic, the resulting cell will survive. These cells are called protoplasts and, as the cytoplasmic membrane is now the limiting structure, the cell assumes a spherical shape. Protoplasts of Gram-negative bacteria are difficult to obtain because the layer of lipopolysaccharide protects the peptidoglycan from attack. In these cases mixtures of EDTA and lysozyme are used and the resulting cells, which still retain fragments of cell envelope, are termed spheroplasts.
Nuclear material.
The genetic information necessary for the functioning of the cell is contained within a single circular molecule of double-stranded DNA. When unfolded, this would be about 1000 times as long as the cell itself and so exists within the cytoplasm in a considerably compacted state. It is condensed into discrete areas called chromatin bodies that are not surrounded by a nuclear membrane. Rapidly dividing cells may contain more than one area of nuclear material but these are copies of the same chromosome, not different chromosomes, and arise because DNA replication proceeds ahead of cell division.
In addition to the main chromosome, cells may contain extra pieces of circular double-stranded DNA which are called plasmids. These can encode a variety of products which are not necessary for the normal functioning of the cell but confer some sort of selective advantage. For example, the plasmids may contain genes conferring antibiotic resistance or the ability to synthesize toxins or virulence factors. Plasmids replicate autonomously (i.e. independent of the main chromosome) and in some cases are able to be transferred from one cell to another (maybe of a different species).
Mesosomes.
These are irregular invaginations of the cytoplasmic membrane which are quite prominent in Gram-positive bacteria but less so in Gram-negative bacteria. It has been proposed that they have a variety of functions, including cross-wall synthesis during cell division and furnishing an attachment site for nuclear material, facilitating the separation of segregating chromosomes during cell division. They have also been implicated in enzyme secretions and may act as a site for cell respiration. However, it has also been suggested that they are simply artefacts which arise as a result of preparation for electron microscopy.
Ribosomes.
The cytoplasm of bacteria is densely populated with ribosomes, which are complexes of RNA and protein in discrete particles 20 nm in diameter. They are the sites of protein synthesis within the cell and the numbers present reflect the degree of metabolic activity of the cell. They are frequently found organized in clusters called polyribosomes or polysomes. Prokaryotic ribosomes have a sedimentation coefficient of 70S, compared to 80S ribosomes of eukaryotic cells. This distinction aids the selective toxicity of a number of antibiotics. The 70S ribosome is made up of RNA and protein and can dissociate into one 30S and one 50S subunit.
Inclusion granules.
Certain bacteria tend to accumulate reserves of materials after active growth has ceased, and these become incorporated within the cytoplasm in the form of granules. The most common are glycogen granules, volutin granules (containing polymetaphosphate) and lipid granules (containing poly β-hydroxybutyric acid). Other granules, such as sulphur and iron, may also be found in the more primitive bacteria.
Flagella.
A flagellum is made up of protein called flagellin and it operates by forming a rigid helix that turns rapidly like a propeller. This can propel a motile cell up to 200 times its own length in 1 second. Under the microscope bacteria can be seen to exhibit two kinds of motion: swimming and tumbling. When tumbling, the cell stays in one position and spins on its own axis but when swimming, it moves in a straight line. Movement towards or away from a chemical stimulus is referred to as chemotaxis. The flagellum arises from the cytoplasmic membrane and is composed of a basal body, hook and filament. The number and arrangement of flagella depend upon the organism and vary from a single flagellum (monotrichous) to a complete covering (peritrichous).
Pili and fimbriae.
These terms are often used interchangeably but in reality these structures are functionally distinct from each other. Fimbriae are smaller than flagella and are not involved in motility. They are found all over the surface of certain bacteria (mainly Gram-negative cells) and are believed to be associated with adhesiveness and pathogenicity. They are also antigenic. Pili (of which there are different types) are larger and of a different structure to fimbriae and are involved in the transfer of genetic information from one cell to another. This is of major importance in the transfer of drug resistance between cell populations.
Endospores.
Under conditions of specific nutrient deprivation some genera of bacteria, in particular Bacillus and Clostridium, undergo a differentiation process at the end of logarithmic growth and change from an actively metabolizing vegetative form to a resting spore form. The process of sporulation is not a reproductive mechanism, as found in certain actinomycetes and filamentous fungi, but serves to enable the organism to survive periods of hardship. A single vegetative cell differentiates into a single spore. Subsequent encounter with favourable conditions results in germination of the spore and the resumption of vegetative activities.
Endospores are very much more resistant to heat, disinfectants, desiccation and radiation than are vegetative cells, making them difficult to eradicate from foods and pharmaceutical products. Heating at 80 °C for 10 minutes would kill most vegetative bacteria, whereas some spores will resist boiling for several hours. The sterilization procedures now routinely used for pharmaceutical products are thus designed specifically with reference to the destruction of the bacterial spore.
The mechanism of this extreme heat resistance was a perplexing issue for many years. At one time it was thought to be due to the presence of a unique spore component, dipicolinic acid (DPA). This compound is found only in bacterial spores where it is associated in a complex with calcium ions. The isolation of heat-resistant DPA-less mutants, however, led to the demise of this theory. Spores do not have a water content appreciably different from that of vegetative cells, but the distribution within the different compartments is unequal and this is thought to generate the heat resistance. The central core of the spore houses the genetic information necessary for growth after germination and this becomes dehydrated by expansion of the cortex against the rigid outer protein coats. Water is thus squeezed out of the central core. Osmotic pressure differences also help to maintain this water imbalance. Endospores are also highly unusual because of their ability to remain dormant and ametabolic for prolonged periods of time. Bacterial spores have been isolated from lake sediments where they were deposited 1000 years previously and there have even been claims of spores revived from geological specimens up to 40 million years old.
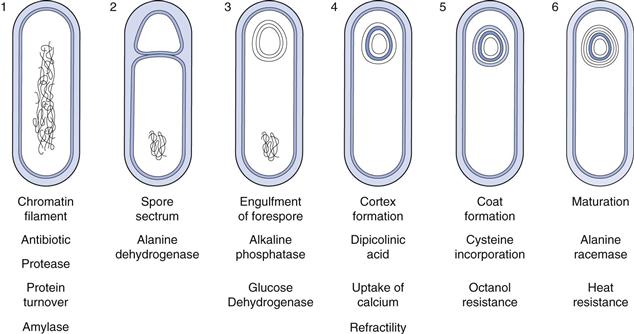
The sequence of events involved in sporulation is illustrated in Figure 13.5. It is a continuous process, although for convenience it may be divided into six stages. The complete process takes about 8 hours, although this may vary depending on the species and the conditions used. Occurring simultaneously with the morphological changes is a number of biochemical events that have been shown to be associated with specific stages and occur in an exact sequence. One important biochemical event is the production of antibiotics. Peptides possessing antimicrobial activity have been isolated from the majority of Bacillus species and many of these have found pharmaceutical applications. Examples of antibiotics include bacitracin, polymyxin and gramicidin. Similarly, the proteases produced by Bacillus species during sporulation are used extensively in a wide variety of industries.
Microscopy and staining of bacteria
Bacterial cells contain about 80% water by weight and this accounts for their very low refractility, i.e. they are transparent when viewed under ordinary transmitted light. Consequently, in order to visualize bacteria under the microscope, the cells must be killed and stained with some compound that scatters the light or, if live preparations are required, special adaptations must be made to the microscope. Such adaptations are found in phase-contrast, dark-ground and differential-interference contrast microscopy.
The microscopic examination of fixed and stained preparations is a routine procedure in most laboratories, but it must be appreciated that not only are the cells dead, but they may also have been altered morphologically by the often quite drastic staining process. The majority of stains used routinely are basic dyes, i.e. the chromophore has a positive charge and this readily combines with the abundant negative charges present both in the cytoplasm in the form of nucleic acids and on the cell surface. These dyes remain firmly adhered even after washing with water. This type of staining is called simple staining and all bacteria and other biological material are stained the same colour. Differential staining is a much more useful process as different organisms or even different parts of the same cell can be stained distinctive colours.
To prepare a film ready for staining, the glass microscope slide must be carefully cleaned to remove all traces of grease and dust. If the culture of bacteria is in liquid form then a loopful of suspension is transferred directly to the slide. Bacteria from solid surfaces require suspension with a small drop of water on the slide to give a faintly turbid film. A common fault with inexperienced workers is to make the film too thick. The films must then be allowed to dry in air. When thoroughly dry the film is fixed by passing the back of the slide through a small Bunsen flame until the area is just too hot to touch on the palm of the hand. The bacteria are killed by this procedure and are also stuck on to the slide. Fixing also makes the bacteria more permeable to the stain and inhibits lysis. Chemical fixation is commonly carried out using formalin or methyl alcohol; this causes less damage to the specimen but tends to be used principally for blood films and tissue sections.
Differential stains
A large number of differential stains have been developed and the reader is referred to the bibliography for more details. Only a few of those available will be discussed here.
Gram’s stain.
By far the most important in terms of use and application is the Gram stain, developed by Christian Gram in 1884 and subsequently modified. The fixed film of bacteria is flooded initially with a solution of methyl violet. This is followed by a solution of Gram’s iodine, which is an iodine–potassium iodide complex acting as a mordant, fixing the dye firmly in certain bacteria and allowing easy removal in others. Decolourization is effected with either alcohol or acetone or mixtures of the two. After treatment some bacteria retain the stain and appear a dark purple colour and these are called Gram positive. Others do not retain the stain and appear colourless (Gram negative). The colourless cells may be stained with a counterstain of contrasting colour, such as 0.5% safranin, which is red.
This method, although extremely useful, must be used with caution as the Gram reaction may vary with the age of the cells and the technique of the operator. For this reason, known Gram-positive and Gram-negative controls should be stained alongside the specimen of interest.
Ziehl–Neelsen’s acid-fast stain.
The bacterium responsible for the disease tuberculosis (Mycobacterium tuberculosis) contains within its cell wall a high proportion of lipids, fatty acids and alcohols, which render it resistant to normal staining procedures. The inclusion of phenol in the dye solution, together with the application of heat, enables the dye (basic fuchsin) to penetrate the cell and, once attached, to resist vigorous decolourization by strong acids, e.g. 20% sulphuric acid. These organisms are therefore called acid fast. Any unstained material can be counterstained with a contrasting colour, e.g. methylene blue.
Fluorescence microscopy
Certain materials, when irradiated by short-wave illuminations, e.g. UV light, become excited and emit visible light of a longer wavelength. This phenomenon is termed fluorescence and will persist only for as long as the material is irradiated. A number of dyes have been shown to fluoresce and are useful in that they tend to be specific to various tissues, which can then be demonstrated by UV irradiation and subsequent fluorescence of the attached fluorochrome. Coupling antibodies to the fluorochromes can enhance specificity, and this technique has found wide application in microbiology. As with the staining procedures described above, this technique can only be applied to dead cells. The three following techniques have been developed for the examination of living organisms.
Dark-ground microscopy
The usual function of the microscope condenser is to concentrate as much light as possible through the specimen and into the objective lens. The dark-ground condenser performs the opposite task, producing a hollow cone of light that comes to a focus on the specimen. The rays of light in the cone are at an oblique angle, such that after passing across the specimen they continue without meeting the front lens of the objective, resulting in a dark background. Any objects present at the point of focus scatter the light, which then enters the objective to show up as a bright image against the dark background.
Specimen preparation is critical, as very dilute bacterial suspensions are required, preferably with all the objects in the same plane of focus. Air bubbles must be absent from both the film and the immersion oil, if used. Dust and grease also scatter light and destroy the uniformly black background required for this technique. With this technique it is not possible to see any real detail but it is useful to study motility.
Phase-contrast microscopy
This technique allows us to see transparent objects well contrasted from the background in clear detail and is the most widely used image enhancement method in microbiology. In essence, an annulus of light is produced by the condenser of the microscope and focused on the back focal plane of the objective where a phase plate, comprising a glass disc containing an annular depression, is situated. The direct rays of the light source annulus pass through the annular groove and any diffracted rays pass through the remainder of the disc. Passage of the diffracted light through this thicker glass layer results in retardation of the light. This alters its phase relationship to the direct rays and increases contrast.
Differential-interference contrast microscopy
This method uses polarized light and has other applications outside the scope of this chapter, such as detecting surface irregularities in opaque specimens. It offers some advantages over phase-contrast microscopy, notably the elimination of haloes around the object edges, and enables extremely detailed observation of specimens. It does, however, tend to be more difficult to set up.
Electron microscopy
The highest magnification available using a light microscope is about 1500 times. This limitation is imposed not by the design of the microscope itself, as much higher magnifications are possible, but by the wavelength of light. An object can only be seen if it causes a ray of light to deflect. If a particle is very small indeed then no deflection is produced and the object is not seen. Visible light has a wavelength between 0.3 and 0.8 µm and objects less than 0.3 µm will not be clearly resolved, i.e. even if the magnification were increased no more detail would be seen. In order to increase the resolution it is necessary to use light of a shorter wavelength, such as UV light. This has been done and resulted in some useful applications but generally, for the purposes of increased definition, electrons are now used and they can be thought of as behaving like very short wavelength light. Transmission electron microscopy requires the preparation of ultrathin (50–60 nm) sections of material mounted on grids for support. Because of the severe conditions applied to the specimen during preparation, and the likelihood of artefacts, care must be taken in the interpretation of information from electron micrographs.
Growth and reproduction of bacteria
The growth and multiplication of bacteria can be examined in terms of individual cells or populations of cells. During the cell division cycle a bacterium assimilates nutrients from the surrounding medium and increases in size. When a predetermined size has been reached the DNA duplicates itself and a cross-wall will be produced, dividing the large cell into two daughter cells, each containing a copy of the parent chromosome. The daughter cells part and the process is known as binary fission. In a closed environment, such as a culture in a test tube, the rate at which cell division occurs varies according to the conditions, and this manifests itself in characteristic changes in the population concentration. When fresh medium is inoculated with a small number of bacterial cells, the number remains static for a short time while the cells undergo a period of metabolic adjustment. This period is called the lag phase (Fig. 13.6) and its length depends on the degree of readjustment necessary. Once the cells are adapted to the environment, they begin to divide in the manner described above, and this division occurs at regular intervals. The numbers of bacteria during this period increase in an exponential fashion, i.e. 2, 4, 8, 16, 32, 64, 128, etc., and this is therefore termed the exponential or logarithmic phase. When cell numbers are plotted on a log scale against time a straight line results for this phase.
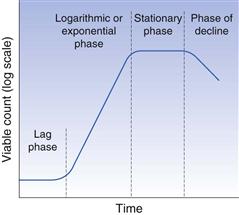
Fig. 13.6 Phases of bacterial growth.
During exponential growth (Fig. 13.6) the medium undergoes continuous change, as nutrients are consumed and metabolic waste products excreted. The fact that the cells continue to divide exponentially during this period is a tribute to their physiological adaptability. Eventually, the medium becomes so changed, due to either substrate exhaustion or excessive concentrations of toxic products, that it is unable to support further growth. At this stage cell division slows and eventually stops, leading to the stationary phase. During this period some cells lyse and die whereas others sporadically divide, but the cell numbers remain more or less constant. Gradually all the cells lyse and the culture enters the phase of decline.
It should be appreciated that this sequence of events is not a characteristic of the cell but a consequence of the interaction of the organisms with the nutrients in a closed environment. It does not necessarily reflect the way in which the organism would behave in vivo.
Genetic exchange
In addition to mutations, bacteria can alter their genetic make-up by transferring information from one cell to another, either as fragments of DNA or in the form of small extrachromosomal elements (plasmids). Transfer can be achieved in three ways: by transformation, transduction or conjugation.
Transformation.
When bacteria die they lyse and release cell fragments, including DNA, into the environment. Several bacterial genera (Bacillus, Haemophilus, Streptococcus, etc.) are able to take up these DNA fragments and incorporate them into their own chromosome, thereby inheriting the characteristics carried on that fragment. Cells able to participate in transformation are called competent. The development of competence has been shown in some cases to occur synchronously in a culture under the action of specific inducing proteins.
Transduction.
Some bacteriophages infect a bacterial cell and incorporate their nucleic acid into the host cell chromosome, with the result that the viral genes are replicated along with the bacterial DNA. In many instances this is a dormant lysogenic state for the phage but sometimes it is triggered into action and lysis of the cell occurs with liberation of phage particles. These new phage particles may have bacterial DNA incorporated into the viral genome and this will infect any new host cell. On entering a new lysogenic state, the new host cell will replicate the viral nucleic acid in addition to that portion received from the previous host. Bacteria in which this has been shown to occur include Mycobacterium, Salmonella, Shigella and Staphylococcus.
Conjugation.
Gram-negative bacteria such as Salmonella, Shigella and Escherichia coli have been shown to transfer genetic material conferring antibiotic resistance by cellular contact. This process is called conjugation and is controlled by an R-factor plasmid, which is a small circular strand of duplex DNA replicating independently from the bacterial chromosome. R factor comprises a region containing resistance transfer genes that control the formation of sex pili, together with a variety of genes that code for the resistance to drugs. Conjugation is initiated when the resistance transfer genes stimulate the production of a sex pilus and random motion brings about contact with a recipient cell. One strand of the replicating R factor is nicked and passes through the sex pilus into the recipient cell. Upon receipt of this single strand of plasmid DNA, the complementary strand is produced and the free ends are joined. For a short time afterwards this cell has the ability to form a sex pilus itself and so transfer the R factor further.
This is by no means an exhaustive discussion of genetic exchange in bacteria and the reader is referred to the bibliography for further information.
Bacterial nutrition
Bacteria require certain elements in fairly large quantities for growth and metabolism, including carbon, hydrogen, oxygen and nitrogen. Sulphur and phosphorus are also required but not in such large amounts. Only low concentrations of iron, calcium, potassium, sodium, magnesium and manganese are needed. Some elements, such as cobalt, zinc and copper, are required only in trace amounts and an actual requirement may be difficult to demonstrate.
The metabolic capabilities of bacteria differ considerably and this is reflected in the form in which nutrients may be assimilated. Bacteria can be classified according to their requirements for carbon and energy.
Lithotrophs (synonym: autotrophs).
These utilize carbon dioxide as their main source of carbon. Energy is derived from different sources within this group:
Oxygen requirements
As mentioned above, all bacteria require elemental oxygen in order to build up the complex materials necessary for growth and metabolism, but many organisms also require free oxygen as the final electron acceptor in the breakdown of carbon and energy sources. These organisms are called aerobes. If the organism will only grow in the presence of air it is called a strict aerobe, but most organisms can either grow in its presence or its absence and are called facultative anaerobes. A strict anaerobe cannot grow and may even be killed in the presence of oxygen, because some other compound replaces oxygen as the final electron acceptor in these organisms. A fourth group of microaerophilic organisms has also been recognized which grow best in only trace amounts of free oxygen and usually prefer an increased carbon dioxide concentration.
Influence of environmental factors on the growth of bacteria
The rate of growth and metabolic activity of bacteria is the sum of a multitude of enzyme reactions. It follows that those environmental factors that influence enzyme activity will also affect growth rate. Such factors include temperature, pH and osmolarity.
Temperature.
Bacteria can survive wide limits of temperature but each organism will exhibit minimum, optimum and maximum growth temperatures and on this basis fall into three broad groups:
Organisms kept below their minimum growth temperature will not divide but can remain viable. As a result, very low temperatures (–70 °C) are used to preserve cultures of organisms for many years. Temperatures in excess of the maximum growth temperature have a much more injurious effect and this is considered in more detail in Chapter 16.
pH.
Most bacteria grow best at around neutrality, in the pH range 6.8–7.6. There are, however, exceptions, such as the acidophilic organism lactobacillus, a contaminant of milk products, which grows best at pHs between 5.4 and 6.6. Yeasts and moulds prefer acid conditions with an optimum pH range of 4–6. The difference in pH optima between fungi and bacteria is used as a basis for the design of media permitting the growth of one group of organisms at the expense of others. Sabouraud medium, for example, has a pH of 5.6 and is a fungal medium, whereas nutrient broth, which is used routinely to cultivate bacteria, has a pH of 7.4. The adverse effect of extremes of pH has for many years been used as a means of preserving foods against microbial attack, for example by pickling in acidic vinegar.
Osmotic pressure.
Bacteria tend to be more resistant to extremes of osmotic pressure than other cells owing to the presence of a very rigid cell wall. The concentration of intracellular solutes gives rise to an osmotic pressure equivalent to between 5 and 20 bar, and most bacteria will thrive in a medium containing around 0.75% w/v sodium chloride. Staphylococci have the ability to survive higher than normal salt concentrations. This has enabled the formulation of selective media, such as mannitol salt agar containing 7.5% w/v sodium chloride, which will support the growth of staphylococci but restrict other bacteria. Halophilic organisms can grow at much higher osmotic pressures but these are all saprophytic and are non-pathogenic to humans. High osmotic pressures generated by either sodium chloride or sucrose have for a long time been used as preservatives. Syrup BP contains 66.7% w/w sucrose and is of sufficient osmotic pressure to resist microbial attack. This is used as a basis for many oral pharmaceutical preparations.
Handling and storage of microorganisms
Because microorganisms have such a diversity of nutritional requirements there has arisen a bewildering array of media for the cultivation of bacteria, yeasts and moulds. Media are produced either as liquids or solidified with agar. Agar is an extract of seaweed, which at concentrations of between 1% and 2% sets to form a firm gel below 45 °C. Unlike gelatin, bacteria cannot use agar as a nutrient and so even after growth the gel remains firm. Liquid media are stored routinely in test tubes or flasks, depending upon the volume, both secured with either loose-fitting caps or plugs of sterile cotton wool. Small amounts of solid media are stored in Petri dishes or slopes (also known as slants), whereas larger volumes may be incorporated in Roux bottles or Carrell flasks.
Bacteria may only be maintained on agar in Petri dishes for a short time (days) before the medium dries out. For longer storage periods the surface of an agar slope is inoculated and after growth the culture may be stored at 4 °C for several weeks. If even longer storage periods are required then the cultures may be stored at low temperatures (–70 °C), usually in the presence of a cryoprotectant such as glycerol. Alternatively they may be freeze-dried (lyophilized) before being stored at 4 °C. Some vegetative cells can survive lyophilization and may retain their viability for many years.
When a single cell is placed on the surface of an over-dried agar plate, it becomes immobilized but can still draw nutrients from the substrate, and consequently grows and divides. Eventually the numbers of bacterial cells are high enough to become visible and a colony is formed. Each of the cells in that colony is a descendant from the initial single cell or group of cells, and so the colony is assumed to be a pure culture with each cell having identical characteristics. The formation of single colonies is one of the primary aims of surface inoculation of solid media and allows the isolation of pure cultures from specimens containing mixed flora.
Inoculation of agar surfaces by streaking
The agar surface must be smooth. The surface should also be without moisture as this could cause the bacteria to become motile and the colonies to merge together. To dry the surface of the agar, the plates are placed in an incubator or drying cabinet for a short while. Inoculating loops can be made of either platinum or nichrome wire twisted along its length to form a loop 2–3 mm in diameter at the end. Nichrome wire is cheaper than platinum but has similar thermal properties. The wire is held in a handle with an insulated grip and the entire length of the wire is heated in a Bunsen flame to red heat to sterilize it. The first few centimetres of the holder are also flamed before the loop is set aside in a rack to cool. Alternatively, disposable pre-sterilized plastic loops are now frequently used.
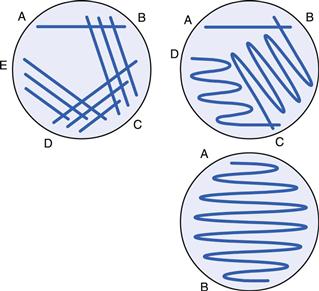
The loop is used to remove a small portion of liquid from a bacterial suspension and this is then drawn across the agar surface from A to B, as indicated in Figure 13.7. The loop is then resterilized (or replaced if plastic) and without reinoculating is streaked over the surface again, ensuring a small area of overlap with the previous streak line. The procedure is repeated as necessary. The pattern of streaking (other examples are shown in Fig. 13.7) is dictated largely by the concentration of the original bacterial suspension. The object of the exercise is to dilute the culture such that, after incubation, single colonies will arise in the later streak lines where the cells were sufficiently separated. All plates are incubated in an inverted position to prevent condensation from the lid falling on the surface of the medium and spreading the colonies.
Inoculation of slopes
A wire needle may be used to transfer single colonies from agar surfaces to the surface of slopes for maintenance purposes. The needle is similar to the loop except that the wire is single and straight, not terminating in a closed end. This is flamed and cooled as before and a portion of a single colony picked off the agar surface. The needle is then drawn upwards along the surface of the slant. Before incubation, the screw cap of the bottle should be loosened slightly to prevent oxygen starvation during growth. Some slopes are prepared with a shallower slope and a deeper butt to allow the needle to be stabbed into the agar when testing for gas production.
Transference of liquids
Graduated pipettes and Pasteur pipettes may be used for this purpose, the latter being short glass tubes one end of which is drawn into a fine capillary. Both types should be plugged with sterile cotton wool and filled via pipette fillers of appropriate capacity. Mouth pipetting should never be permitted. Automatic pipettes have generally replaced glass graduated pipettes in most areas of science for the measurement of small volumes of liquid. Provided they are properly maintained and calibrated, they have the advantage of being easy to use and reliable in performance.
Release of infectious aerosols
During all of these manipulations two considerations must be borne in mind. First, the culture must be transferred with the minimum risk of contamination from outside sources. To this end all pipettes, tubes, media, etc., are sterilized and the manipulations carried out under aseptic conditions. Second, the safety of the operator is paramount. During operations with microorganisms, it must be assumed that all organisms are capable of causing disease and that any route of infection is possible.
Most infections acquired in laboratories cannot be traced to a specific incident but arise from the inadvertent release of infectious aerosols. Two types of aerosols may be produced. The first kind produces large droplets (>5 µm), containing many organisms, which settle locally and contaminate surfaces in the vicinity of the operator. These may initiate infections if personnel touch the surfaces and subsequently transfer the organisms to eyes, nose or mouth. The second type of aerosol contains droplets less than 5 µm in size, which dry instantly to form droplet nuclei that remain suspended in the air for considerable periods. This allows them to be carried on air currents to situations far removed from the site of initiation. These particles are so small that they are not trapped by the usual filter mechanisms in the nasal passages and may be inhaled, giving rise to infections of the lungs.
The aerosols described above may be produced by a variety of means, such as heating wire loops, placing hot loops into liquid cultures, splashing during pipetting, rattling loops and pipettes inside test tubes, opening screw-capped tubes and ampoules, etc. All microbiologists should have an awareness of the dangers of aerosol production and learn the correct techniques to minimize them.
Cultivation of anaerobes
Anaerobic microbiology is a much-neglected subject owing principally to the practical difficulties involved in growing organisms in the absence of air. However, with the increasing implication of anaerobes in certain disease states and improved cultivation systems, the number of workers in this field is growing.
The most common liquid medium for cultivation of anaerobes is thioglycollate medium. In addition to sodium thioglycollate, the medium contains methylene blue as a redox indicator, and it permits the growth of aerobes, anaerobes and microaerophilic organisms. When in test tubes, the medium may be used after sterilization until not more than one-third of the liquid is oxidized, as indicated by the colour of the methylene blue indicator. Boiling and cooling of the medium just prior to inoculation are recommended for maximum performance. In some cases the presence of methylene blue poses toxicity problems and under these circumstances the indicator may be removed.
Anaerobic jars have improved considerably in recent years, making the cultivation of even strict anaerobes now relatively simple. The most common consist of a clear polycarbonate jar and are designed to be used with disposable oxygen absorbants and CO2 generators such as the AnaeroGen sachet. Once opened, the sachet will rapidly absorb atmospheric oxygen from the jar and simultaneously generate carbon dioxide. It is important therefore to open the sachet, place within the jar and seal the lid of the jar within one minute. The oxygen level will be reduced to below 1% within 30 minutes and the final carbon dioxide level will be between 9 and 11%. Carbon dioxide is produced to allow the growth of many fastidious anaerobes, which fail to grow in its absence. The absence of oxygen can be demonstrated by the action of a redox indicator, which in the case of methylene blue will be colourless.
Counting bacteria
Estimates of bacterial numbers in a suspension can be evaluated from a number of standpoints, each equally valid, depending upon the circumstances and the information required. In some cases it may be necessary to know the total amount of biomass produced within a culture, irrespective of whether the cells are actively metabolizing. In other instances only an assessment of living bacteria may be required. Bacterial counts can be divided into total counts and viable counts.
Total counts
These counts estimate the total number of bacteria present within a culture, both dead and living cells. A variety of methods is available for the determination of total counts and the one chosen will depend largely upon the characteristics of the cells being studied, i.e. whether they aggregate together.
Microscope methods.
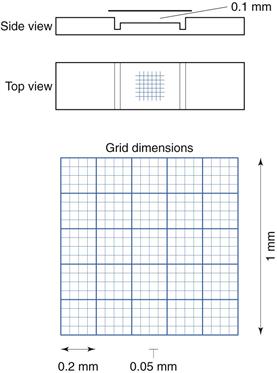
Microscope methods employ a haemocytometer counting chamber (Fig. 13.8), which has a platform engraved with a grid of small squares each 0.0025 mm2 in area. The platform is depressed 0.1 mm and a glass coverslip is placed over the platform, enclosing a space of known dimensions. The volume above each square is 0.00025 mm3. For motile bacteria the culture is fixed by adding two to three drops of 40% formaldehyde solution per 10 mL of culture to prevent the bacteria from moving across the field of view. A drop of the suspension is then applied to the platform at the edge of the coverslip. The liquid is drawn into the space by capillary action. It is important to ensure that liquid does not enter a trench that surrounds the platform; the liquid must fill the whole space between the coverslip and the platform. This slide is examined using phase-contrast or dark-ground microscopy and, if necessary, the culture is diluted to give 2–10 bacteria per small square. A minimum of 300 bacterial cells should be counted to give statistically significant results.
Spectroscopic methods.
These methods are simple to use and very rapid but require careful calibration if meaningful results are to be obtained. Either opacity or light scattering may be used but both methods may only be used for dilute, homogeneous suspensions as at higher concentrations the cells obscure each other in the light path and the relationship between optical density and concentration is not linear. Simple colorimeters and nephelometers can be used but more accurate results are obtained using a spectrophotometer.
Electronic methods.
A variety of automated methods is available for bacterial cell counting, including electronic particle counting, microcalorimetry, changes in impedance or conductivity, and radiometric and infrared systems for monitoring CO2 production.
Other methods.
If an organism is prone to excessive clumping, or if a measure of biomass is needed rather than numbers, then estimates may be made by performing dry weight or total nitrogen determinations. For dry weight determinations, a sample of suspension is centrifuged and the pellet washed free of culture medium by further centrifugation in water. The pellet is collected and dried to a constant weight in a desiccator. Total nitrogen measures the total quantity of nitrogenous material within a cell population. A known volume of suspension is centrifuged and washed as before and the pellet digested using sulphuric acid in the presence of a CuSO4-K2SO4-selenium catalyst. This produces ammonia, which is removed using boric acid and estimated either by titration or colorimetrically.
Viable counts
These are counts to determine the number of bacteria in a suspension that are capable of division. In all these methods, the assumption is made that a colony arises from a single cell, although clearly this is often not the case, as cells frequently clump or grow as aggregates, e.g. Staphylococcus aureus. For this reason viable counts are usually expressed as colony-forming units (cfu) per mL rather than cells per mL.
Spread plates.
A known volume, usually no more than 0.2 mL, of a suitably diluted culture is pipetted on to an over-dried agar plate and distributed evenly over the surface using a sterile spreader made of glass or plastic. All the liquid must be allowed to soak in before the plates are inverted. A series of 10-fold dilutions should be made in a suitable sterile diluent and replicates plated out at each dilution in order to ensure that countable numbers of colonies (30–300) are obtained per plate.
The viable count is calculated from the average colony count per plate, knowing the dilution and the volume pipetted onto the agar.
Pour plates.
A series of dilutions of original culture is prepared as before, ensuring that at least one is in the range 30–300 organisms/mL. One-millilitre quantities are placed into empty sterile Petri dishes. Molten agar, cooled to 45°C, is poured on to the suspension and mixed by gentle swirling. After setting, the plates are inverted and incubated. Because the colonies are embedded within the agar they do not exhibit the characteristic morphology seen with surface colonies. In general, they assume a lens shape and are usually smaller. Because the oxygen tension below the surface is reduced this method is not suitable for strict aerobes. Calculations are similar to that given above, except that no correction is necessary for volume placed upon the plate.
Membrane filtration.
This method is particularly useful when the level of contamination is very low, such as in water supplies. A known volume of sample is passed through a membrane filter, typically made of cellulose acetate/nitrate, of sufficient pore size to retain bacteria (0.2–0.45 µm). The filtrate is discarded and the membrane placed bacteria uppermost on the surface of an over-dried agar plate, avoiding trapped air between membrane and surface. Upon incubation the bacteria draw nutrients through the membrane and form countable colonies.
ATP determination.
There are sometimes instances when viable counts are required for clumped cultures or for bacteria adhered to surfaces, for example in biofilms. Conventional plate count techniques are not appropriate here and ATP determinations can be used. The method assumes that viable bacteria contain a relatively constant level of ATP, but this falls to zero when the cells die. ATP is extracted from the cells using a strong acid such as trichloroacetic acid, and the extract is then neutralized by dilution with buffer. The ATP assay is based upon the quantitative measurement of a stable level of light produced as a result of an enzyme reaction catalysed by firefly luciferase.

The amount of ATP is calculated by reference to light output from known ATP concentrations and the number of bacterial cells is calculated by reference to a previously constructed calibration plot.
Isolation of pure bacterial cultures
Mixed bacterial cultures from pathological specimens or other biological materials are isolated first on solid media to give single colonies. The resultant pure cultures can then be subjected to identification procedures. The techniques used for isolation depend upon the proportion of the species of interest compared to the background contamination. Direct inoculation can only be used when an organism is found as a pure culture in nature. Examples include bacterial infections of normally sterile fluids such as blood or cerebrospinal fluid.
Streaking is the most common method employed. If the proportions of bacteria in the mixed culture are roughly equal then streaking on an ordinary nutrient medium should yield single colonies of all microbial types. More usually, the organism of interest is present only as a very small fraction of the total microbial population, necessitating the use of selective media.
A selective enrichment broth is initially inoculated with the mixed population of cells and this inhibits the growth of the majority of the background population. At the same time the growth of the organism of interest is encouraged. After incubation in these media the cultures are streaked out on to solid selective media, which frequently contain indicators to further differentiate species on the basis of fermentation of specific sugars.
Classification and identification
Taxonomy is the ordering of living organisms into groups on the basis of their similarities. In this way we can construct a hierarchy of interrelationships such that species with similar characteristics are grouped within the same genus, genera which have similarities are grouped within the same family, families grouped into orders, orders into classes and classes into divisions. The classification of bacteria does pose a problem because a species is defined as a group of closely related organisms that reproduce sexually to produce fertile offspring. Of course, bacteria do not reproduce sexually and so a bacterial species is simply defined as a population of cells with similar characteristics.
Nomenclature
The total number of different bacterial species on the planet can only be speculated and probably runs into tens of millions, however, the number of known, named species is just over 6,000. It is therefore extremely important to be sure there is no confusion when describing any one particular bacterial species. Although we are familiar with the use of trivial names in ornithology and botany (we understand what we mean when we describe a sparrow or a daffodil), such an approach could have disastrous consequences in clinical microbiology. For this reason, we use the binomial system of nomenclature developed by Carolus Linnaeus in the 18th century. In this system every bacterium is given two names, the first being the genus name and the second the species name. By convention, the name is italicized or underlined, and the genus name always begins with a capital letter whereas the species name begins in lower case.
Identification
The organization of bacteria into groups of related microorganisms is based upon the similarity of their chromosomal DNA. Although this provides a very accurate indicator of genetic relatedness it is far too cumbersome a tool to use for the identification of an unknown bacterium isolated from a sample. In this instance, a series of rapid and simple tests is required that probe the phenotypic characteristics of the microorganism. The tests are conducted in a logical series of steps, the results from each test providing information for the next stage of the investigation. An example of such a procedure is given below:
| Morphology: | microscope investigations using a wet mount to determine cell size, shape, formation of spores, aggregation, motility, etc. |
| Staining reactions: | Gram stain, acid-fast stain, spore stain |
| Cultural reactions: | appearance on solid media (colony formation, shape, size, colour, texture, smell, pigments, etc.); aerobic/anaerobic growth, temperature requirements, pH requirements |
| Biochemical reactions: | enzymatic activities are probed to distinguish between closely related bacteria. This can be performed in traditional mode or using kits. |
Biochemical tests.
These are designed to examine the enzymatic capabilities of the organism. As there is a large number of biochemical tests that can be performed, the preliminary steps help to narrow down the range to those that will be most discriminatory. Given below are a few examples of commonly used biochemical tests.
Sugar fermentation is very frequently used and examines the ability of the organism to ferment a range of sugars. A number of tubes of peptone water are prepared, each containing a different sugar. An acid–base indicator is incorporated into the medium that also contains a Durham tube (a small inverted tube filled with medium) capable of collecting any gas produced during fermentation. After inoculation and incubation, the tubes are examined for acid production (as indicated by a change in the colour of the indicator) and gas production (as seen by a bubble of gas collected in the inverted Durham tube).
Proteases are produced by a number of bacteria, e.g. Bacillus species and Pseudomonas, and they are responsible for the breakdown of protein into smaller units. Gelatin is a protein that can be added to liquid media to produce a stiff gel similar to agar. Unlike agar, which cannot be utilized by bacteria, those organisms producing proteases will destroy the gel structure and liquefy the medium. A medium made of nutrient broth solidified with gelatin is normally incorporated in boiling tubes or small bottles and inoculated by means of a stab wire. After incubation, it is important to refrigerate the gelatin prior to examination; otherwise false positives may be produced. Proteases can also be detected using milk agar, which is opaque. Protease producers form colonies with clear haloes around them where the enzyme has diffused into the medium and digested the casein.
Oxidase is produced by Neisseria and Pseudomonas and can be detected using 1% tetramethylparaphenylene diamine. The enzyme catalyses the transport of electrons between electron donors in the bacteria and the redox dye. A positive reaction is indicated by a deep purple colour in the reduced dye. The test is carried out by placing the reagent directly on to an isolated colony on an agar surface. Alternatively, a filter paper strip impregnated with the dye is moistened with water and, using a platinum loop, a bacterial colony spread across the surface. If positive, a purple colour will appear within 10 seconds. Note that the use of iron loops may give false-positive reactions.
The indole test distinguishes those bacteria capable of decomposing the amino acid tryptophan to indole. Any indole produced can be tested by a colorimetric reaction with p-dimethylaminobenzaldehyde. After incubation in peptone water, 0.5 mL Kovacs reagent is placed on the surface of the culture, shaken, and a positive reaction is indicated by a red colour. Organisms giving positive indole reactions include E. coli and Proteus vulgaris.
Catalase is responsible for the breakdown of hydrogen peroxide into oxygen and water. The test may be performed by adding 1 mL of 10-vol hydrogen peroxide directly to the surface of colonies growing on an agar slope. A vigorous frothing of the surface liquid indicates the presence of catalase. Staphylococcus and Micrococcus are catalase positive, whereas Streptococcus is catalase negative.
Urease production enables certain bacteria to break down urea to ammonia and carbon dioxide:
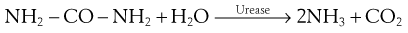
This test is readily carried out by growing the bacteria on a medium containing urea and an acid–base indicator. After incubation the production of ammonia will be shown by the alkaline reaction of the indicator. Examples of urease-negative bacteria include E. coli and Enterococcus faecalis.
Simmons citrate agar was developed to test for the presence of organisms that could utilize citrate as the sole source of carbon and energy and ammonia as the main source of nitrogen. It is used to differentiate members of the Enterobacteriaceae. The medium, containing bromothymol blue as indicator, is surface inoculated on slopes and citrate utilization demonstrated by an alkaline reaction and a change in the indicator colour from a dull green to a bright blue. E. coli, Shigella, Edwardsiella and Yersinia do not utilize citrate, whereas Serratia, Enterobacter, Klebsiella and Proteus do and so give a positive result.
The methyl red test is used to distinguish organisms that, during metabolism of glucose, produce and maintain a high level of acidity from those that initially produce acid but restore neutral conditions with further metabolism. The organism is grown on glucose phosphate medium and, after incubation, a few drops of methyl red are added and the colour immediately recorded. A red colour indicates acid production (positive), whereas a yellow colour indicates alkali (negative).
Some organisms can convert carbohydrates to acetyl methyl carbinol (CH3-CO-CHOH-CH3). This may be oxidized to diacetyl (CH3-CO-CO-CH3), which will react with guanidine residues in the medium under alkaline conditions to produce a colour. This is the basis of the Voges Proskauer test, which is usually carried out at the same time as the methyl red test. The organism is again grown in glucose phosphate medium and, after incubation, 40% KOH is added together with 5% α-naphthol in ethanol. After mixing, a positive reaction is indicated by a pink colour in 2–5 minutes, gradually becoming darker red up to 30 minutes. Organisms giving positive Voges Proskauer reactions usually give negative methyl red reactions, as the production of acetylmethyl carbinol is accompanied by low acid production. Klebsiella species typically give a positive Voges Proskauer reaction.
Rapid identification systems.
With the increasing demand for quick and accurate identification of bacteria, a number of micromethods have been developed combining a variety of biochemical tests selected for their rapidity of reading and high discrimination. The API bacterial identification system is an example of such a micromethod and comprises a plastic tray containing dehydrated substrates in a number of wells. Culture is added to the wells, dissolving the substrate and allowing the fermentation of carbohydrates or the presence of enzymes similar to those just described to be demonstrated. In some cases incubation times of 2 hours are sufficient for accurate identification. Kits are available with different reagents, permitting the identification of Enterobacteriaceae, Streptococcaceae, Staphylococci, anaerobes, yeasts and moulds. Accurate identification is made by reference to a table of results.
MALDI-TOF (matrix-assisted laser desorption/ionization – time of flight) mass spectroscopy is used increasingly. Here a bacterial sample is transferred to the MALDI target plate and overlaid with matrix solution. The sample is loaded into the mass spectrometer and a profile acquired. This profile is a unique fingerprint of the microorganism and is compared to the library of electronic MS spectra held within the software database. Although the equipment cost is high, this procedure is ideal for those laboratories that have a high throughput of microbial samples that require rapid processing.
The tests described so far will enable differentiation of an unknown bacterium to species level. However, it is apparent that not all isolates of the same species behave in an identical manner. For example, E. coli isolated from the intestines of a healthy person is relatively harmless compared to the well-publicized E. coli O157.H7, which causes intense food poisoning and haemolytic uraemic syndrome. On occasions it is therefore necessary to distinguish further between isolates from the same species. This can be performed using, among other things, serological tests and phage typing.
Serological tests.
Bacteria have antigens associated with their cell envelopes (O-antigens), with their flagella (H-antigens) and with their capsules (K-antigens). When injected into an animal, antibodies will be produced directed specifically towards those antigens and able to react with them. Specific antisera are prepared by immunizing an animal with a killed or attenuated bacterial suspension and taking blood samples. Serum containing the antibodies can then be separated. If a sample of bacterial suspension is placed on a glass slide and mixed with a small amount of specific antiserum, then the bacteria will be seen to clump when examined under the microscope. The test can be made more quantitative by using the tube dilution technique, where a given amount of antigen is mixed with a series of dilutions of specific antisera. The highest dilution at which agglutination occurs is called the agglutination titre.
Phage typing.
Many bacteria are susceptible to lytic bacteriophages whose action is very specific. Identification may be based on the susceptibility of a culture to a set of such type-specific lytic bacteriophages. This method enables very detailed identification of the organisms to be made, e.g. one serotype of Salmonella typhi has been further subdivided into 80 phage types using this technique.
Fungi
Fungus is a general term used to describe all yeasts and moulds, whereas a mould is a filamentous fungus exhibiting a mycelial form of growth. The study of fungi is called mycology. Yeasts and moulds are eukaryotic microorganisms possessing organized demonstrable nuclei enclosed within an outer membrane, a nucleolus and chromatin strands that become organized into chromosomes during cell division. Fungal cell walls are composed predominantly of polysaccharide. In most cases this is chitin mixed with cellulose, glucan and mannan. Proteins and glycoproteins are also present but peptidoglycan is absent. The polysaccharide polymers are crosslinked to provide a structure of considerable strength which gives the cell osmotic stability. The fungal membrane contains sterols such as ergosterol and zymosterol not found in mammalian cells, and this provides a useful target for antifungal antibiotics. The role of fungi in nature is predominantly a scavenging one and in this respect they are vital for the decomposition and recycling of organic materials. Of the more than 100 000 species of known fungi, fewer than 100 are human pathogens and most of these are facultative and not obligate parasites.
Fungal morphology
The fungi can be divided into five broad groups on the basis of their morphology.
Yeasts
These are spherical or ovoid unicellular bodies 2–4 µm in diameter which typically reproduce by budding. In liquid cultures and on agar they behave very much like bacteria. Examples include Saccharomyces cerevisiae, strains of which are used in baking and in the production of beers and wines. Cryptococcus neoformans is the only significant pathogen and this gives rise to a respiratory tract disease called cryptococcosis, which in most cases is relatively mild. However, the microorganism may disseminate, leading to multiorgan disease, including meningitis. Cryptococcosis is of particular significance in immune-compromised patients. If left untreated, 80% of patients with disseminated cryptococcosis will die within 1 year.
Yeast-like fungi
These organisms normally behave like typical budding yeasts but under certain circumstances the buds do not separate and become elongated. The resulting structure resembles a filament and is called a pseudomycelium. It differs from a true mycelium in that there are no interconnecting pores between the cellular compartments comprising the hyphae.
The most important member of this group is Candida albicans, which is usually resident in the mouth, intestines and vagina. Under normal conditions Candida does not cause problems but if the environmental balance is disturbed then problems can arise. These include vaginal thrush (vaginitis) and oral thrush. Overgrowth of Candida albicans within the gut can lead to symptoms of inexplicable fatigue and malaise that is difficult to diagnose. Predisposing factors may include poor diet, diabetes, alcoholism and long-term treatment with steroids.
Dimorphic fungi
These grow as yeasts or as filaments depending upon the cultural conditions. At 22 °C, either in the soil or in culture media, filamentous mycelial forms and reproductive spores are produced, whereas at 37 °C in the body, the microorganisms assume a yeast-like appearance. Histoplasma capsulatum is an important pathogen that gives rise to respiratory illness. The infectious form is the spore that is borne on the wind and is inhaled. It has been postulated that a single spore can elicit an infection. On entering the body, the spores germinate to give rise to the yeast form. Primary infections are often mild but progressive disseminated histoplasmosis is a very severe disease that can affect many organs of the body.
Filamentous fungi
This group comprises those multicellular moulds that grow in the form of long, slender filaments 2–10 µm in diameter called hyphae. The branching hyphae, which constitute the vegetative or somatic structure of the mould, intertwine and gradually spread over the entire surface of the available substrate, extracting nutrients and forming a dense mat or mycelium. The hyphae may be non-septate (coenocytic) or septate, but in each case the nutrients and cellular components are freely diffusible along the length of the filament. This is facilitated by the presence of pores within the septa.
Mushrooms and toadstools
This group is characterized by the production of large reproductive fruiting bodies of complex structure. They also possess elaborate propagation mechanisms. Some of these fungi are edible and are used in cooking but others, such as Amanita phalloides (death cap), produce potent mycotoxins that may result in death if eaten.
Reproduction of fungi
In the somatic portion of most fungi the nuclei are very small and the mechanism of nuclear division is uncertain. Under the correct environmental conditions the organisms will switch from the somatic or vegetative growth phase to a reproductive form, so that the fungus may propagate the species by producing new mycelia on fresh food substrates. Two types of reproduction are found: asexual and sexual.
Asexual reproduction
Asexual reproduction is in general more important for the propagation of the species. Mechanisms include binary fission, budding, hyphal fragmentation and spore formation. Each progeny is an exact replica of the parent and no species variation can occur. Some yeasts (e.g. Schizosaccharomyces rouxii) reproduce by binary fission in the same way as bacteria. The parent cell enlarges, its nucleus divides and, when a cross-wall is produced across the cell, two identical daughter cells form.
Budding occurs in the majority of yeasts and is the production of a small outgrowth or bud from the parent cell. As the bud increases in size, the nucleus divides and one of the pair migrates into the bud. The bud eventually breaks off from the parent to form a new individual. A scar is left behind on the parent cell and each parent can produce up to 24 buds.
Fungi growing in a filamentous form may employ hyphal fragmentation as a means of asexual propagation. The hyphal tips break up into component segments (called arthroconidia or arthrospores), each of which can disperse on the wind to other environments and fresh food substrates.
The formation of specialized spore-bearing structures containing reproductive spores is the most common method of asexual reproduction (Fig. 13.9). The spores can be borne in a sporangium, supported on a sporangio-phore. A limiting membrane surrounds the sporangium and the spores contained within it are called sporangiospores. The spores are released when the sporangium ruptures. This type of reproduction is found in the lower fungi possessing non-septate hyphae (e.g. Mucor and Rhizopus). Separate spores produced at the tips of specialized conidiophores are called conidiospores. A diverse range of structures is found in nature and Figure 13.9 illustrates some of the different types of asexual spores found in fungi.
Sexual reproduction
Sexual reproduction involves the union of two compatible nuclei and allows variation of the species. Mycology is made much more complex because individual fungi are given different names depending upon whether they are in the sexual or the asexual stage. Not all fungi have been observed to carry out sexual reproduction. Some species produce distinguishable male and female sex organs on the same mycelium and are therefore hermaphroditic, i.e. a single colony can reproduce sexually by itself. Others produce mycelia which are either male or female (called dioecious) and can therefore only reproduce when two dissimilar organisms come together.
Fungal classification
The pharmaceutically important fungi can be found within four main taxonomic classes.
Zygomycetes
These are terrestrial saprophytes possessing non-septate hyphae and are sometimes referred to as the lower fungi. Apart from their hyphae, they can be distinguished from other filamentous fungi by the presence of sporangia. Examples are Mucor and Rhizopus, which are important in the manufacture of organic acids and the biotransformation of steroids. They are also common spoilage organisms.
Ascomycetes
Ascomycetes possess septate hyphae and the sexual or perfect stage is characterized by the presence of a saclike reproductive structure called an ascus. This typically contains eight ascospores. The asexual or imperfect stage involves conidiospores. An example is Claviceps purpurea, which is a parasite of rye and is important as a source of ergot alkaloids used to control haemorrhage and in treating migraine. A subclass of the Ascomycetes is the Hemiascomycetes. This includes the yeasts such as Saccharomyces and Cryptococcus, together with Torulopsis and Candida.
Deuteromycetes
Sometimes called the Fungi Imperfecti, this group includes those fungi in which the sexual stage of reproduction has not been observed. Penicillium and Aspergillus are Ascomycetes but classified among the Deuteromycetes as the perfect stage is apparently absent. Penicillium chrysogenum is important in the production of the antibiotic penicillin, whereas Aspergillus species have found widespread industrial usage owing to their extensive enzymic capabilities. Some Aspergillus species also produce mycotoxins and can cause serious infections in humans. The Deuteromycetes contain most of the human pathogens, such as Blastomyces and Coccidioides, and some of the dermatophyte fungi.
Basidiomycetes
This is the most advanced group, containing the mushrooms and toadstools. Sexual reproduction is by basidiospores. The group also includes the rusts (cereal parasites) and smuts.
Bibliography
1. Collins CH, Lyne PM, Grange JM, Falkinham J. Microbiological Methods. 8th edn London: Hodder Arnold; 2004.
2. Denyer SP, Hodges NA, Gorman SP. Hugo and Russell’s Pharmaceutical Microbiology. 7th edn Oxford: Blackwell Publishing; 2004.
3. Fraise A, Lambert PA, Maillard JY. Hugo, Russell and Ayliffe’s Principles and Practice of Disinfection, Preservation and Sterilization. 4th edn Oxford: Blackwell Science; 2004.
4. Gillespie SH, Bamford K. Medical Microbiology and Infection at a Glance. 3rd edn Oxford: Blackwell; 2007.
5. Russell AD, Chopra I. Understanding Antibacterial Action and Resistance. 2nd edn London: Ellis Horwood; 1996.
6. Stryer L, Berg J, Tymoczko J. Biochemistry. 7th edn New York: Freeman; 2006.
[/level-membership-for-basic-science-category][not-level-membership-for-basic-science-category]
Fundamentals of microbiology
Geoffrey W. Hanlon
Chapter contents
Microscopy and staining of bacteria
Growth and reproduction of bacteria
Handling and storage of microorganisms
Isolation of pure bacterial cultures
Key points
Introduction
Microorganisms are ubiquitous in nature and are vital components in the cycle of life. The majority are free-living organisms growing on dead or decaying matter whose prime function is the turnover of organic materials in the environment. Pharmaceutical microbiology, however, is concerned with the relatively small group of biological agents that cause human disease, spoil prepared medicines or can be used to produce compounds of medical interest.
In order to understand microorganisms more fully, living organisms of similar characteristics have been grouped together into taxonomic units. The most fundamental division is between prokaryotic and eukaryotic cells, which differ in a number of respects (Table 13.1) but particularly in the arrangement of their nuclear material. Eukaryotic cells contain chromosomes, which are separate from the cytoplasm and contained within a limiting nuclear membrane, i.e. they possess a true nucleus. Prokaryotic cells do not possess a true nucleus and their nuclear material is free within the cytoplasm, although it may be aggregated into discrete areas called nuclear bodies. Prokaryotic organisms make up the lower forms of life and include Eubacteria and Archaeobacteria. Eukaryotic cell types embrace all the higher forms of life, of which only the fungi will be dealt with in this chapter.
Table 13.1
| Structure | Prokaryotes | Eukaryotes |
| Cell wall structure | Usually contains peptidoglycan | Peptidoglycan absent |
| Nuclear membrane | Absent | Present. Possess a true nucleus |
| Nucleolus | Absent | Present |
| Number of chromosomes | One | More than one |
| Mitochondria | Absent | Present |
| Mesosomes | Present | Absent |
| Ribosomes | 70S | 80S |
One characteristic shared by all microorganisms is the fact that they are small; however, it is a philosophical argument whether all infectious agents can be regarded as living. Some are little more than simple chemical entities incapable of any free-living existence. Viroids, for example, are small circular, single-stranded RNA molecules not complexed with protein. One particularly well-studied viroid has only 359 nucleotides (one-10th the size of the smallest known virus) and yet causes a disease in potatoes. Prions are small, self-replicating proteins devoid of any nucleic acid. The prion associated with Creutzfeld–Jakob disease in humans, scrapie in sheep and bovine spongiform encephalitis in cattle has only 250 amino acids and is highly resistant to inactivation by normal sterilization procedures.
Viruses are more complex than viroids or prions, possessing both protein and nucleic acid. Despite being among the most dangerous infectious agents known, they are still not regarded as living. Table 13.2 shows the major groups of viruses infecting humans.
Viruses
Viruses are obligate intracellular parasites with no intrinsic metabolic activity, being devoid of ribosomes and energy-producing enzyme systems. They are thus incapable of leading an independent existence and cannot be cultivated on cell-free media, no matter how nutritious. The size of human viruses ranges from the largest poxviruses, measuring about 300 nm, to the picornaviruses, such as the poliovirus which is approximately 20 nm. When one considers that a bacterial coccus measures 1000 nm in diameter, it can be appreciated that only the very largest virus particles may be seen under the light microscope, and electron microscopy is required for visualizing the majority. It will also be apparent that few of these viruses are large enough to be retained on the 200 nm (0.2 µm) membrane filters used to sterilize thermolabile liquids.
Viruses consist of a core of nucleic acid (either DNA as in vaccinia virus or RNA as in poliovirus) surrounded by a protein shell or capsid. Most DNA viruses have linear, double-stranded DNA but in the case of the parvovirus it is single stranded. The majority of RNA-containing viruses contain one molecule of single-stranded RNA, although in reoviruses it is double stranded. The protein capsid comprises 50–90% of the weight of the virus and, as nucleic acid can only synthesize approximately 10% its own weight of protein, the capsid must be made up of a number of identical protein molecules. These individual protein units are called capsomeres and are not in themselves symmetrical but are arranged around the nucleic acid core in characteristic symmetrical patterns. Additionally, many of the larger viruses possess a lipoprotein envelope surrounding the capsid arising from the membranes within the host cell. In many instances the membranes are virus modified to produce projections outwards from the envelope, such as haemagglutinins or neuraminidase. The enveloped viruses are often called ether sensitive, as ether and other organic solvents may dissolve the membrane.
The arrangement of the capsomeres can be of a number of types.
Reproduction of viruses
Because viruses have no intrinsic metabolic capability, they require the functioning of the host cell machinery in order to manufacture and assemble new virus particles. It is this intimate association between the virus and its host that makes the treatment of viral infections so complex. Any chemotherapeutic approach which damages the virus will almost inevitably cause injury to the host cells and hence lead to side-effects. An understanding of the life cycle of the virus is vital in determining suitable target sites for antiviral chemotherapy. The replication of viruses within host cells can be broken down into a number of stages.
Adsorption to host cell
The first step in the infection process involves virus adsorption on to the host cell. This usually occurs via an interaction between protein or glycoprotein moieties on the virus surface with specific receptors on the host cell outer membrane. Different cells possess receptors for different viruses. For example, the human immunodeficiency virus (HIV) possesses two proteins involved in adsorption to T lymphocytes and these are known as gp41 and gp120. There are receptors on the lymphocyte surface to which HIV will bind. The main receptor is CD4 to which the gp120 protein attaches. Other receptors are CXCR4 or CCR5 to which the gp41 protein binds. Both attachments are necessary for infection and lead to conformational changes in the HIV envelope proteins, resulting in membrane fusion.
Penetration
Enveloped viruses fuse the viral membrane with the host cell membrane and release the nucleocapsid directly into the cytoplasm. Naked virions generally penetrate the cell by phagocytosis. Bacteriophages are viruses which specifically attack bacteria and these inject their DNA into the host cell while the rest of the virus remains on the outside.
Uncoating
In this stage the capsid is removed as a result of attack by cellular proteases and this releases the nucleic acid into the cytoplasm. These first three stages are similar for both DNA and RNA viruses.
Nucleic acid and protein synthesis
The detailed mechanisms by which DNA- and RNA-containing viruses replicate inside the cell are outside the scope of this chapter and the reader is referred to the bibliography for further information. After nucleic acid replication, early viral proteins are produced, the function of which is to switch off host cell metabolic activity and direct the activities of the cell towards the synthesis of proteins necessary for the assembly of new virus particles.
Assembly of new virions
Again, there are differences in the detail of how the viruses are assembled within the host cell, but construction of new virions occurs at this stage and up to 100 new virus particles may be produced per cell.
Release of virus progeny
The newly formed virus particles may be liberated from the cell as a burst, in which case the host cell ruptures and dies. Infection with influenza virus results in a lytic response. Alternatively, the virions may be released gradually from the cell by budding of the host cell plasma membrane. These are often called ‘persistent’ infection; an example being hepatitis B.
Latent infections
In some instances a virus may enter a cell but not go through the replicative cycle outlined above and the host cell may be unharmed. The genome of the virus is conserved and may become integrated into the host cell genome where it may be replicated along with the host DNA during cell division. At some later stage the latent virus may become reactivated and progress through a lytic phase, causing cell damage/death and the release of new virions. Examples of this type of infection are those which occur with the herpes simplex viruses associated with cold sores, genital herpes and also chickenpox where the dormant virus may reactivate to give shingles later in life.
Oncogenic viruses
Oncogenic viruses have the capacity to transform the host cell into a cancer cell. In some cases this may lead to relatively harmless, benign growths, such as warts caused by papovavirus, but in other cases more severe, malignant tumours may arise. Cellular transformation may result from viral activation or mutation of normal host genes, called protooncogenes, or the insertion of viral oncogenes.
Bacteriophages
Bacteriophages (phages) are viruses that attack bacteria but not animal cells. It is generally accepted that the interaction between phage and bacterium is highly specific, and there is probably at least one phage for each species of bacterium. In many cases the infection of a bacterial cell by a phage results in lysis of the bacterium; such phages are termed virulent. Some phages, however, can infect a bacterium without causing lysis. In this case the phage DNA becomes incorporated within the bacterial genome. The phage DNA can then be replicated along with the bacterial cell DNA; this is then termed a prophage. Bacterial cells carrying a prophage are called lysogenic and phages capable of inducing lysogeny are called temperate. Occasionally some of the prophage genes may be expressed and this will confer on the bacterial cell the ability to produce new proteins. The ability to produce additional proteins as a result of prophage DNA is termed lysogenic conversion.
The discovery of bacteriophages in the early 20th century is attributed to two workers, Frederick Twort and Felix d’Herelle. In 1896 Ernest Hankin had made an observation that the waters of the Ganges River possessed antibacterial properties which may have led to a reduction in cases of dysentery and cholera in the areas surrounding the river. Twort and d’Herelle independently came to the conclusion that this effect must be due to a virus. Twort did not carry on with his research but d’Herelle quickly established the potential of bacteriophages in antibacterial therapy 10 years before the advent of antibiotics. It was the discovery of penicillin by Alexander Fleming in 1928 that led to the demise of bacteriophage therapy but interest is now increasing again due to the emergence of antibiotic-resistant strains of bacteria.
Archaeobacteria
Archaeobacteria are a fascinating group of prokaryotic microorganisms that are frequently found living in hostile environments. They differ in a number of respects from Eubacteria, particularly in the composition of their cell walls. They comprise methane producers, sulfate reducers, halophiles and extreme thermophiles. However, they are of little significance from a pharmaceutical or clinical standpoint and so will not be considered further.
Eubacteria
Eubacteria constitute the major group of prokaryotic cells that have pharmaceutical and clinical significance. They include a diverse range of microorganisms, from the primitive parasitic rickettsias that share some of the characteristics of viruses, through the more typical free-living bacteria to the branching, filamentous actinomycetes, which at first sight resemble fungi rather than bacteria.
Atypical bacteria
Rickettsiae
The family Rickettsiaceae includes three clinically important genera, Rickettsia, Coxiella and Bartonella. Although these are prokaryotic cells, they differ from most other bacteria both in their structure and in the fact that the majority of species lead an obligate intracellular existence. This means that, with a few exceptions, they cannot be grown on cell-free media, although unlike many viruses they do possess some independent enzymes. They have a pleomorphic appearance, ranging from coccoid through to rod-shaped cells; multiplication is by binary fission. Their cell wall composition bears similarities to that of Gram-negative bacteria (see later in this chapter) and in general they stain this way. The genus Rickettsia has a number of species that give rise to human diseases, in particular epidemic typhus (R. prowazekii), murine typhus (R. typhi) and spotted fevers (various species). These are characterized by transmission via insect vectors, particularly mites, ticks, fleas and lice.
The mode of transmission by these vectors varies depending upon the insect concerned. In the case of lice and fleas, the microorganisms multiply within the insect and get into the faeces. These insects then colonize humans and transmit the microorganism when the faeces or the insect itself is crushed onto the skin. No bite is necessary and the faeces may also be inhaled. Mites and ticks pick up the microorganism when they take a blood meal from an infected animal. They then pass on the infection to humans when they accidentally bite us.
Coxiella burnetii is the only species in the genus Coxiella and this gives rise to a disease called Q fever. Although the source of the disease is infected animals, usually no insect vector is involved and the most common route of transmission is by inhalation of infected dust. Bartonella quintana is the causative agent of trench fever which, as the name suggests, occurs typically under conditions of war and deprivation. Each of the infections described here can be treated with doxycycline, although the duration of therapy may vary depending upon the nature of the disease and its severity.
Chlamydiae
These are obligate intracellular parasitic bacteria that possess some independent enzymes but lack the ability to generate ATP. Two cellular forms are identified: a small (0.3 µm) highly infectious elementary body which, after infection, enlarges to give rise to the replicative form called the initial or reticulate body (0.8–1.2 µm). These divide by binary fission within membrane-bound vesicles in the cytoplasm of infected cells. Insect vectors are not required for the transmission of infection. Chlamydiae lack peptidoglycan in their cell walls and have weak Gram-negative characteristics.
Chlamydia trachomatis is the most important member of the group, being responsible for the disease trachoma, characterized by inflammation of the eyelids, which can lead to scarring of the cornea. This is the most common cause of infectious blindness worldwide. It is estimated that 400 million people are infected, with at least 6 million totally blind. The same species is also recognized as one of the major causes of sexually transmitted disease. C. psittaci and C. pneumoniae are responsible for respiratory tract infections. Chlamydial infections are responsive to treatment by tetracyclines, either topical or systemic as appropriate.
Mycoplasmas
The mycoplasmas are a group of very small (0.3–0.8 µm) prokaryotic microorganisms that are capable of growing on cell-free media but which lack cell walls. The cells are surrounded by a double-layered plasma membrane that contains substantial amounts of phospholipids and sterols. This structure has no rigidity owing to the absence of peptidoglycan, and so the cells are susceptible to osmotic lysis. The lack of peptidoglycan is also the reason for these bacteria being resistant to the effects of cell wall-acting antibiotics such as the penicillins, and also the enzyme lysozyme. Members of this group are pleomorphic, varying in shape from coccoid to filamentous. Most are facultative anaerobes capable of growth at 35 °C, and on solid media produce colonies with a characteristic ‘fried egg’ appearance. They contain a number of genera, of which the most important from a clinical point of view are Mycoplasma and Ureaplasma. M. pneumoniae is a major cause of respiratory tract infections in children and young adults, whereas U. urealyticum has been implicated in non-specific genital infections. Despite being resistant to the β-lactam antibiotics, these infections can be effectively treated using either tetracyclines or erythromycin.
Actinomycetes
Many of the macroscopic features of the actinomycetes are those that are more commonly found among the filamentous fungi but they are indeed prokaryotic cells. They are a diverse group of Gram-positive bacteria morphologically distinguishable from other bacteria because they have a tendency to produce branching filaments and reproductive spores. Nocardia contain a number of species that have been shown to be pathogenic to humans, but they occur principally in tropical climates. Reproduction in this genus is by fragmentation of the hyphal strands into individual cells, each of which can form a new mycelium. The genus Streptomyces contains no human pathogens but most species are saprophytic bacteria found in the soil. They are aerobic microorganisms producing a non-fragmenting, branching mycelium that may bear spores. The reason for their pharmaceutical importance is their ability to produce a wide range of therapeutically useful antibiotics, including streptomycin, chloramphenicol, oxytetracycline, erythromycin and neomycin.
Typical bacteria
Shape, size and aggregation
Bacteria occur in a variety of shapes and sizes, determined not only by the nature of the organisms themselves but also by the way in which they are grown (Fig. 13.1). In general, bacterial dimensions lie in the range 0.75–5 µm. The most common shapes are the sphere (coccus) and the rod (bacillus).
Some bacteria grow in the form of rods with a distinct curvature, e.g. vibrios are rod-shaped cells with a single curve resembling a comma, whereas a spirillum possesses a partial rigid spiral; spirochaetes are longer and thinner, exhibit a number of turns and are also more flexible. Rod-shaped cells occasionally grow in the form of chains but this is dependent upon growth conditions rather than being a characteristic of the species.
Cocci, however, show considerable variation in aggregation, which is characteristic of the species. The plane of cell division and the strength of adhesion of the cells determine the extent to which they aggregate after division. Cocci growing in pairs are called diplococci, those in four are tetrads and groups of eight are sarcina. If a chain of cells is produced resembling a string of beads this is termed a streptococcus, whereas an irregular cluster similar in appearance to a bunch of grapes is called a staphylococcus. In many cases this is sufficiently characteristic to give rise to the name of the bacterial genus, e.g. Staphylococcus aureus, Streptococcus pneumoniae.
Anatomy
Figure 13.2 is a diagrammatic representation of a typical bacterial cell. The various components are described below.
Capsule.
Many bacteria produce extracellular polysaccharides, which may take the form of either a discrete capsule firmly adhered to the cell or a more diffuse layer of slime. Not all bacteria produce a capsule and even those that can will only do so under certain circumstances. Many encapsulated pathogens, when first isolated, give rise to colonies on agar which are smooth (S) but subculturing leads to the formation of rough colonies (R). This S to R transition is due to loss in capsule production. Reinoculation of the R cells into an animal results in the resumption of capsule formation, indicating that the capacity has not been lost irrevocably.
The function of the capsule is generally regarded as protective, as encapsulated cells are more resistant to disinfectants, desiccation and phagocytic attack. In some organisms, however, it serves as an adhesive mechanism; for example, Streptococcus mutans is an inhabitant of the mouth that metabolizes sucrose to produce a polysaccharide capsule enabling the cell to adhere firmly to the teeth. This is the initial step in the formation of dental plaque, which is a complex array of microorganisms and organic matrix that adheres to the teeth and ultimately leads to decay. The substitution of sucrose by glucose prevents capsule formation and hence eliminates plaque.
A similar picture emerges with Staph. epidermidis. This bacterium forms part of the normal microflora of the skin and was originally thought of as non-pathogenic. With the increased usage of indwelling medical devices coagulase-negative staphylococci, in particular Staph. epidermidis, have emerged as the major cause of device-related infections. The normal microbial flora has developed the ability to produce extracellular polysaccharide, which enables the cells to form resistant biofilms attached to the devices. These biofilms are very difficult to eradicate and have profound resistance to antibiotics and disinfectants. It is now apparent that the dominant mode of growth for aquatic bacteria is not planktonic (free swimming) but sessile, i.e. attached to surfaces and covered with protective extracellular polysaccharide or glycocalyx.
Cell wall.
Bacteria can be divided into two broad groups by the use of the Gram-staining procedure (see later in this chapter for details), which reflects differences in cell wall structure. The classification is based upon the ability of the cells to retain the dye methyl violet after washing with a decolourizing agent such as absolute alcohol. Gram-positive cells retain the stain whereas Gram-negative cells do not. As a very rough guide, the majority of small rod-shaped cells are Gram negative. Most large rods, such as the Bacillaceae, lactobacilli and actinomycetes, are Gram positive. Similarly, most cocci are Gram positive, although there are notable exceptions, such as the Neisseriaceae.
Bacteria are unique in that they possess peptidoglycan in their cell walls. This is a complex molecule with repeating units of N-acetylmuramic acid and N-acetylglucosamine (Fig. 13.3). This extremely long molecule is wound around the cell and crosslinked by polypeptide bridges to form a structure of great rigidity. The degree and nature of crosslinking vary between bacterial species. Crosslinking imparts to the cell its characteristic shape and has principally a protective function. Peptidoglycan (also called murein or mucopeptide) is the site of action of a number of antibiotics, such as penicillin, bacitracin, vancomycin and cycloserine. The enzyme lysozyme is also capable of hydrolysing the β–1–4 linkages between N-acetylmuramic acid and N-acelylglucosamine.

Fig. 13.3 Peptidoglycan.
Figure 13.4 shows simplified diagrams of a Gram-positive and a Gram-negative cell wall. The Gram-positive cell wall is much simpler in layout, containing peptidoglycan interspersed with teichoic acid polymers. These latter are highly antigenic but do not provide structural support. Functions attributed to teichoic acids include the regulation of enzyme activity in cell wall synthesis, sequestration of essential cations, cellular adhesion and mediation of the inflammatory response in disease. In general, proteins are not found in Gram-positive cell walls. Gram-negative cell walls are more complex, comprising a much thinner layer of peptidoglycan surrounded by an outer bilayered membrane. This outer membrane acts as a diffusional barrier and is the main reason why many Gram-negative cells are much less susceptible to antimicrobial agents than are Gram-positive cells.
The lipopolysaccharide component of the outer membrane can be shed from the wall upon cell death. It is a highly heat-resistant molecule known as endotoxin, which has a number of toxic effects on the human body, including fever, shock and even death. For this reason it is important that solutions for injection or infusion are not just sterile but are also free from endotoxins.
Cytoplasmic membrane.
The cytoplasmic membranes of most bacteria are very similar and are composed of protein, lipids, phospholipids and a small amount of carbohydrate. The components are arranged into a bilayer structure with a hydrophobic interior and a hydrophilic exterior. The cytoplasmic membrane has a variety of functions.
• It serves as an osmotic barrier.
• It is selectively permeable and is the site of carrier-mediated transport.
• It is the site of ATP generation and cytochrome activity.
The cytoplasmic membrane has very little tensile strength and the internal hydrostatic pressure of up to 20 bar forces it firmly against the inside of the cell wall. Treatment of bacterial cells with lysozyme may remove the cell wall and, as long as the conditions are isotonic, the resulting cell will survive. These cells are called protoplasts and, as the cytoplasmic membrane is now the limiting structure, the cell assumes a spherical shape. Protoplasts of Gram-negative bacteria are difficult to obtain because the layer of lipopolysaccharide protects the peptidoglycan from attack. In these cases mixtures of EDTA and lysozyme are used and the resulting cells, which still retain fragments of cell envelope, are termed spheroplasts.
Nuclear material.
The genetic information necessary for the functioning of the cell is contained within a single circular molecule of double-stranded DNA. When unfolded, this would be about 1000 times as long as the cell itself and so exists within the cytoplasm in a considerably compacted state. It is condensed into discrete areas called chromatin bodies that are not surrounded by a nuclear membrane. Rapidly dividing cells may contain more than one area of nuclear material but these are copies of the same chromosome, not different chromosomes, and arise because DNA replication proceeds ahead of cell division.
In addition to the main chromosome, cells may contain extra pieces of circular double-stranded DNA which are called plasmids. These can encode a variety of products which are not necessary for the normal functioning of the cell but confer some sort of selective advantage. For example, the plasmids may contain genes conferring antibiotic resistance or the ability to synthesize toxins or virulence factors. Plasmids replicate autonomously (i.e. independent of the main chromosome) and in some cases are able to be transferred from one cell to another (maybe of a different species).
Mesosomes.
These are irregular invaginations of the cytoplasmic membrane which are quite prominent in Gram-positive bacteria but less so in Gram-negative bacteria. It has been proposed that they have a variety of functions, including cross-wall synthesis during cell division and furnishing an attachment site for nuclear material, facilitating the separation of segregating chromosomes during cell division. They have also been implicated in enzyme secretions and may act as a site for cell respiration. However, it has also been suggested that they are simply artefacts which arise as a result of preparation for electron microscopy.
Ribosomes.
The cytoplasm of bacteria is densely populated with ribosomes, which are complexes of RNA and protein in discrete particles 20 nm in diameter. They are the sites of protein synthesis within the cell and the numbers present reflect the degree of metabolic activity of the cell. They are frequently found organized in clusters called polyribosomes or polysomes. Prokaryotic ribosomes have a sedimentation coefficient of 70S, compared to 80S ribosomes of eukaryotic cells. This distinction aids the selective toxicity of a number of antibiotics. The 70S ribosome is made up of RNA and protein and can dissociate into one 30S and one 50S subunit.
Inclusion granules.
Certain bacteria tend to accumulate reserves of materials after active growth has ceased, and these become incorporated within the cytoplasm in the form of granules. The most common are glycogen granules, volutin granules (containing polymetaphosphate) and lipid granules (containing poly β-hydroxybutyric acid). Other granules, such as sulphur and iron, may also be found in the more primitive bacteria.
Flagella.
[/not-level-membership-for-basic-science-category]