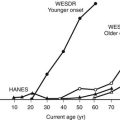
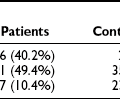Development of the Endocrine Pancreas
Pancreas Morphogenesis
During gastrulation, mesoendodermal cells that migrate through the node region start to segregate to form distinct mesodermal and endodermal germ layers. Endodermal cells aggregate to form a contiguous epithelium that spans the whole length of the forming gastrointestinal tract of the developing embryo. Digestive organs develop along the anteroposterior and dorsoventral axes of this sheet of cells in a precise and predetermined pattern.1–3
The pancreas is formed from clumps of cells that bud from the dorsal and ventral aspects of the gut endoderm near the foregut/midgut junction4,5 (Fig. 5-1). Well before any morphologic evidence of the pancreas becomes apparent, interactions between the mesodermally derived notochord and the endodermal epithelium initiate dorsal pancreas formation.6 Similarly, interactions between the lateral plate mesoderm and the lateral endoderm ensure proper induction of the ventral pancreas.7 The first morphologic sign of pancreas formation is a thickening of the dorsal endodermal sheet caudal to the stomach anlage. This bud structure continues to grow and initiates contact with the dorsal aorta that has fused in the midline and thus separated the notochord and the endoderm.8 Subsequently, dorsal mesenchyme replaces the aorta and provides essential signals that stimulate pancreatic epithelial branching and cell differentiation.
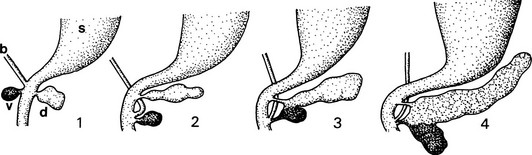
FIGURE 5-1 Schematic outline of the development of the mammalian pancreas. 1, Dorsal and ventral buds arise from the duodenal part of the primitive gut endoderm (blackened). 2, The dominant ventral bud, following the bile duct, turns and approaches the dorsal bud. 3, The two buds fuse together. 4, The ventral bud becomes part of the head of the pancreas (the PP-rich duodenal lobe), whereas the dorsal bud gives rise to the body and tail (the glucagon-rich splenic lobe); the pancreatic juice is drained off by the main and accessory pancreatic ducts. b, bile duct; d, dorsal bud; s, stomach; v, ventral bud.
Similar tissue interactions guide the development of the two ventral buds. In contrast to the dorsal bud that forms in the midline of the epithelium, the ventral buds are derived from the ventrolateral edges of the endodermal sheet. Instructive signals provided by the lateral plate mesenchyme initiate ventral pancreas formation, and interaction with smaller blood vessels further supports organ growth.8 One of these ventral buds degenerates, while the other one fuses with the dorsal bud when stomach and gut rotate to form the mature organ. The regulation of this rotation is not well understood, but perturbations lead to malformations in humans, including annular pancreas, wherein a part of the ventral pancreas is mislocalized and can constrict the adjacent duodenum.9
Mesenchymal-epithelial interactions are required for proper development of mature tissue10,11; however, the epithelial cells start to outgrow the mesenchyme during later stages of development, and few remnants of this tissue remain in the adult pancreas. Epithelial cells branch into the surrounding mesenchyme to form an elaborate duct system that allows transport of exocrine enzymes into the duodenum. Distal cells of the branching epithelium differentiate into acinar cells that are connected to the duct system via centroacinar cells. Acinar cells are organized as small glands that produce a variety of digestive enzymes, including amylases, peptidases, nucleases, and lipases.
In contrast to the continuous acinar duct system, endocrine precursors delaminate from immature ducts. These distinct islet cells produce a variety of hormones, including insulin, glucagon, pancreatic polypeptide, and somatostatin, which are secreted directly into the bloodstream and regulate gastrointestinal function and nutrient storage and utilization. Because of intimate interactions between endocrine cells and blood vessels, islets are highly vascularized. Similarly, hormone secretion is controlled at least in part by the nervous system; the sympathetic, parasympathetic, and sensory neurons that innervate the islets are derived from cells that migrate from the neural crest into the pancreas just as the two buds fuse.12
Early Organ Specification and Bud Formation
Tissue Interaction and Signaling Pathways
With the exception of supporting mesenchyme, blood vessels, and innervating neurons, all mature pancreatic cell types are derived from epithelial cells of endodermal origin.13–15 Several studies have addressed the question of how the pancreas anlage becomes specified within the epithelial sheet that forms the endoderm, and increasing evidence points to a series of tissue interactions as important steps during early stages of organogenesis. During gastrulation, the mesoectodermal portion of the embryo signals to the endodermal sheet to establish a prepattern along the anterior-posterior axis.3
Although the exact signals required for patterning of the fore-midgut area are not completely understood, studies in mice have implicated the fibroblast growth factor (FGF) signaling pathway in this process.16 Before any morphologic signs of pancreas formation are apparent, the midline of the endodermal epithelium comes in close contact with the overlying notochord, a mesodermal structure known to regulate organogenesis and cell differentiation in adjacent structures. The notochord provides numerous secreted signaling proteins that are required for initiation of dorsal pancreas organogenesis. Members of the transforming growth factor-β (TGF-β) and FGF signaling pathways have been implicated in the notochord-mediated induction of pancreas development. A critical aspect of their activity involves repression of the expression of Sonic Hedgehog (Shh), a member of the Hedgehog signaling pathway within the pancreas anlage.17 Ectopic elevation of Hedgehog signaling at the onset of pancreas formation results in pancreas agenesis, indicating that tight regulation of the activity of this pathway is essential for proper organ formation. However, the notochord provides only permissive signals because it cannot induce expression of pancreatic markers in nonpancreatic endoderm.6
In contrast to the singular dorsal pancreas, two distinct pancreatic buds form within the ventrolateral endoderm, a region that is not contacted by the notochord. Tissue recombination experiments in chick and mice demonstrate that similar but distinct signals from the lateral plate mesoderm (LPM) ensure the correct temporal-spatial induction of the ventral pancreas.7,16 Before contact is made with the LPM, no pancreatic markers are detectable in the presumptive ventral pancreas, but LPM or the TGF-β superfamily members activin and morphogenetic proteins (BMPs), as well as retinoic acid (RA), have been shown to induce expression of pancreatic genes in underlying endoderm (see the following section).7 It is important to note that mesenchyme isolated from the pancreatic region carries the potential to induce pancreatic gene expression in anterior endoderm normally fated to develop into stomach and esophagus. Thus, in contrast to the notochord-endoderm interaction that induces dorsal pancreas development in a permissive fashion, the LPM actively instructs uncommitted endoderm to differentiate into pancreatic tissue. Activin, BMP, and RA signals that mimic the pancreas instructive activity of the LPM are potentially useful in designing a cocktail of growth factors designed to regulate differentiation of uncommitted progenitor cells toward a pancreatic fate.
Other studies have shown that interactions between heart mesenchyme and ventral foregut endoderm regulate differentiation of ventral pancreas and liver progenitor cells.18 In an analogy to the notochord-dorsal pancreas bud interactions, the septum transversum and heart mesenchyme produce BMP and FGF signals, respectively, that regulate organ differentiation of the ventral endodermal organs, including liver and pancreas. Because of its close proximity, the concentration of FGF ligands produced by the heart mesenchyme is higher at the area fated to develop into liver and lower at the more distal region that develops into ventral pancreas.18 This is also the case for dorsal pancreatic epithelium, in which low levels of FGF signals have been shown to initiate pancreatic gene expression and block Shh expression, while higher levels of FGF signaling block pancreas induction via increased Hedgehog signaling. Thus, formation of dorsal and ventral pancreatic buds depends on low FGF signals and inhibition of Hedgehog signaling. It is important to note that Hedgehog signaling is active in areas immediately adjacent to the dorsal and ventral buds, thereby establishing a molecular boundary that prevents ectopic expansion of pancreatic tissue beyond its normal borders.
During subsequent stages, the flattened endodermal sheet expands and folds itself to form a tube-like structure. The first morphologic sign of pancreas formation is an epithelial thickening at the dorsal side of the forming gut tube caudal to the stomach anlage, shortly followed by the appearance of two ventral thickenings next to the liver diverticulum. Outgrowth and tissue-specific gene expression continue to depend on cues derived from adjacent tissues. In contrast to earlier stages, mesenchymal signals are now produced from forming blood vessels, the dorsal aorta, and smaller vitelline veins that contact the dorsal and ventral buds, respectively.19 Depletion of blood vessels via explantation in Xenopus or via homozygous recombination in transgenic mice that lack the Flk-1 gene, a receptor for vascular epithelial growth factor (VEGF), impairs the differentiation of the dorsal pancreatic epithelium.19,20 Vice versa, ectopic expression of VEGF leads to hypervascularization and islet hyperplasia in pancreatic tissues, as well as ectopic insulin expression in nonpancreatic stomach epithelium, suggesting that VEGF or molecules secreted by endothelial cells support endocrine cell differentiation. Detailed analysis of the interaction between blood vessels and pancreas buds has revealed physical contact between the aorta and the dorsal bud epithelium, while vitelline veins and ventral bud epithelium are separated by a fine band of mesenchymal cells, although endothelial cells are less critical for ventral bud formation.20
Recent results have provided further evidence for the role of signaling molecules in setting up organ boundaries in the pancreas anlage. Studies in zebrafish suggest that Fgf10 expression is critical for separating pancreatic and liver cells in the area of the hepatopancreatic ductal system. Fgf10 is expressed in the mesenchyme surrounding the developing hepatopancreatic duct and functions to restrain inappropriate differentiation of adjacent pancreatic into liver cells, as well as liver into pancreatic cells.21 Thus, secreted signaling molecules are essential in defining pancreas organ boundaries.
Upon specification of the dorsal and ventral pancreatic anlagen, additional mesenchymal-epithelial interactions are required for later steps of pancreas organogenesis.10,11 Splanchnic mesoderm expands medially and surrounds the pancreatic epithelial buds. Signals from the mesenchyme promote epithelial expansion that results in the formation of a branching structure composed of undifferentiated ductal cells. Based on morphologic criteria, Rutter and colleagues argued that early endodermal cells are “protodifferentiated,” and that mature exocrine and endocrine cells appear only after the secondary transition, a process during which the numbers of differentiated acinar and β cells increase significantly.4,22 More recent studies have shown that early pancreatic mesenchyme generally induces growth and proliferation of epithelial cells, while at later stages, mesenchymal signals promote exocrine and prohibit endocrine cell differentiation.23,24
Induction of the Pancreatic Gene Expression Program
Transcription factors expressed broadly in the endoderm provide competence to respond to the endoderm patterning signals and include several transcription factors originally described in the adult liver, such as hepatic nuclear factors 1b (a POU-homeodomain factor),25 3a and b (forkhead factors now known as Foxa1 and Foxa2),26 4a (an orphan nuclear receptor),27,28 and 6 (a cut-homeodomain factor now known as Onecut1),29,30 and the zinc-finger transcription factors Gata4 and Gata6.31,32 These genes function in a transcriptional hierarchy that not only controls endoderm patterning but also persists in endoderm-derived organs and plays a role in the mature pancreas.33–38 Several of these endoderm factors have been directly implicated in the control of mature pancreatic gene expression (see Fig. 5-5 and Table 5-1).
The earliest genes selectively expressed in prepancreatic endoderm are two transcription factors, the parahox homeodomain factor Pdx139–41 and the basic-helix-loop-helix (bHLH) transcription factor Ptf1a.42,43 Pdx1 expression first appears in prepancreatic endoderm more than a day before the initial formation of the dorsal pancreatic bud, and is preceded immediately by the appearance of another parahox factor, Mnx1 (also known as HB9),44,45 which is expressed more broadly than Pdx1 in anterior endoderm. Expression of both Mnx1 and Pdx1 persists in the initial pancreatic buds, although Mnx1 expression is extinguished quickly, and Pdx1 expression lasts a few more days. Expression of both factors is reactivated in mature β cells.
Dorsal, but not ventral, expression of Pdx1 depends on Mnx1.44,45 Because Mnx1 is also expressed in the notochord during the same period, it is formally possible that Mnx1 could control dorsal Pdx1 expression via signals from the notochord. Pdx1 expression in the dorsal prepancreatic endoderm does not require signals from the notochord, however, suggesting that Mnx1 functions cell intrinsically in inducing Pdx1. Extrinsic signals that induce Pdx1 expression in the dorsal prepancreatic endoderm have not been identified.6 On the other hand, as described in the preceding section, signals from the LPM, including activin, BMPs, and retinoic acid, can induce Pdx1 expression in the ventral prepancreatic endoderm.7
Studies of the Pdx1 promoter have identified several additional endodermal transcription factors as potential intrinsic regulators of Pdx1 expression.46–53 These include members of the Hnf1 and Foxa families of transcription factors, Onecut1, the paired homeodomain factor Pax6, and Pdx1 itself46,47,52,54,55 (Mnx1 also may act through these same Pdx1 binding sites, given its similarity to Pdx1 in the DNA-binding homeodomain). Hnf1a and Pax6 are not expressed early enough or broadly enough to initiate the early expression of Pdx1 in the embryonic gut and pancreatic buds, but the expression patterns of Onecut1, Hnf1β, Foxa1, and Foxa2 suggest that they could play this role (Fig. 5-2). Mice lacking Onecut1 have reduced expression of Pdx1,55 and embryoid bodies lacking Foxa2 fail to activate the pdx1 gene,52 suggesting that these two factors may be bona fide activators of Pdx1 expression in the prepancreatic endoderm and pancreatic buds and therefore lie directly upstream of Pdx1 in the hierarchy of factors involved in initiating pancreas development (see Fig. 5-5).
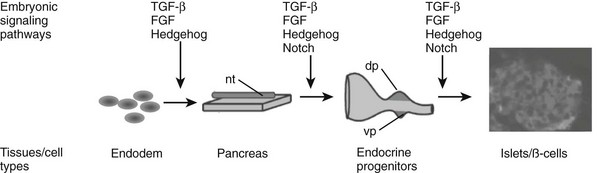
FIGURE 5-2 Embryonic signaling pathways regulate different steps of pancreas and endocrine islet formation. dp, Dorsal pancreas; nt, notochord; vp, ventral pancreas.
Pdx1 is required for outgrowth of the pancreatic buds. In mice with targeted disruptions of the pdx1 gene, the pancreatic buds fail to branch and expand after initial formation, yielding an animal that lacks a pancreas at birth.40,41,56 Confirming its role upstream of Pdx1 in the endoderm, lack of Onecut1 results in reduced pancreatic size,55 and removal of Mnx1 specifically arrests the growth of the dorsal pancreatic bud.44,45 A homozygous null mutation in the IPF1 gene, which encodes human Pdx1, has been identified in a human patient with pancreatic agenesis,57 and mutations in the zebrafish gene encoding Pdx1 also affect pancreas development,58 reflecting an evolutionary conservation of Pdx1 function.
Expression of Ptf1a follows shortly after Pdx1 and is restricted within the endoderm to the dorsal and ventral pancreatic buds.59,60 The aorta induces expression of Ptf1a in the dorsal pancreatic bud, but vascular-derived signals do not appear to play as critical a role in its expression in the ventral buds.20 Although Ptf1a was initially described as an exocrine-specific transcription factor,43,60 in fact it plays an essential role in determining pancreatic cell fate. In mouse embryos lacking Ptf1a, cells normally fated to contribute to both exocrine and endocrine lineages in the pancreas instead revert to duodenal epithelium.42
Cell Type Differentiation
FGF7 and FGF10 are expressed transiently in pancreatic mesenchyme and have been shown to promote proliferation of pancreatic epithelial cells.61,62 Transgenic mice that carry a homozygous deletion in the FGF10 gene display severe defects in pancreas morphogenesis that result from loss of Pdx1-positive, pancreatic epithelial progenitor cells.62 FGF signaling also has been shown to regulate proliferation of endocrine and exocrine cells,63,64 and expression of a dominant active form of the FGF receptor 2 (FGFR2) inhibits endocrine and exocrine cells.63–65
TGF-β/activin signaling has been shown to affect differentially exocrine and endocrine cell differentiation. Treatment of isolated pancreatic buds with soluble TGF-β1 stimulates differentiation of endocrine cells, particularly β and PP cells.66 Signaling via activins, a subgroup of TGF-β signaling members, also has been shown to affect preferentially endocrine cell differentiation. Treatment of pancreatic buds with follistatin, an antagonist that physically binds to activins, inhibits endocrine cell differentiation while promoting exocrine cell formation.67 Transgenic mice ectopically expressing dominant-negative activin type II receptor (dnActRII) develop islet hypoplasia, a phenotype also observed in mice carrying targeted mutations in both ActRIIA and ActRIIB.68–70 More recent work has focused on the downstream signaling cascade of TGF-β/activin signaling. Ligand binding to their respective receptors on receiving cells results in the phosphorylation of receptor-associated Sma- and Mad-related proteins 1, 2, 3, 5, and 8 (R-Smads). Another Smad, Smad4, acts as a universal partner for the R-Smads, and the R-Smad/Smad4 complex interacts with distinct DNA-binding factors to regulate target gene transcription. It is surprising that loss of Smad4 has no appreciable effects on pancreas formation,71,72 suggesting Smad4-independent TGF-β/activin functions. Additional support for the TGF-β/activin requirement during pancreas development comes from recent studies in which Smad7, a negative factor that interferes with Smad-receptor and Smad-Smad interactions, is ectopically expressed in pancreatic tissue. Embryonic expression in pancreatic epithelium results in pancreas and β cell hypoplasia.73 Of note, ectopic expression specifically in adult pancreatic Pdx1+ cells resulted in hypoinsulinemia followed by diabetes.
The role of Hedgehog signaling during endocrine cell differentiation is less clear. At early stages, elevation of Hedgehog signaling in the pancreas anlage results in transformation of the pancreatic mesenchyme into duodenal mesoderm.17,74 Increased Hedgehog signaling impairs pancreatic organogenesis, at least in part, because signals normally provided by the pancreatic mesenchyme are missing. Also, as has been shown for other organs, ectopic activation of Hedgehog signaling reduces FGF10 expression known to promote pancreatic epithelial and endocrine cell expansion.75,76 By contrast, a low level of Hedgehog signaling is detected throughout pancreas organogenesis and in mature islets.77 In addition, cell culture experiments have revealed that Hedgehog signaling activates Pdx1 and insulin promoters in cultured insulinoma cells,78,79 suggesting a different and potentially important role for the pathway in maintaining mature β cell function. As they result in early embryonic lethality before pancreas formation is initiated, conventional knockout mice lacking all Hedgehog signaling have not proved useful in determining whether pancreatic Hedgehog signaling is essential for some aspects of pancreas organogenesis.80 These questions await tissue-specific inactivation of the pathway in the pancreas.
Hedgehog signaling shares many features with another embryonic pathway, the canonical Wnt cascade. Several Wnt ligands and their respective frizzled receptors have been identified in embryonic and adult pancreas.81,82 Functional studies in which dominant-negative receptors were expressed ectopically, or β-catenin, an essential component of the canonical Wnt signaling cascade, was eliminated, have pointed to an important role of this pathway during embryonic pancreas formation; however, the exact function of each specific pancreas cell type is still under debate. Although several studies found a requirement for canonical Wnt signaling during exocrine development,83–85 others have noted a role for β-catenin in endocrine function.86 Ectopic activation of β-catenin signaling results in different phenotypes, depending on the time of expression. Activation at the onset of pancreas formation blocks progenitor cell expansion, and later expression results in a profound increase in acinar cell numbers.87 A recent study analyzing the requirement for the mouse homolog of pygopus 2 (mPygo-2), a necessary nuclear component of the Wnt cascade, revealed a novel role for Wnt signaling in early pancreatic mesenchyme.88 According to this study, mesenchymal mPygo-2 is essential in regulating epithelial progenitor proliferation, including endocrine progenitors. Thus, Wnt signaling regulates pancreas formation at various stages in both mesenchyme and epithelium. Furthermore, studies in adult animals indicate a role for canonical Wnt signaling in adult β cell proliferation.89
In addition to the TGF-β, FGF, Hedgehog, and Wnt signaling pathways, Notch signaling contributes to cell fate choices and differentiation in the pancreas, as it does in many other organs during embryogenesis. Notch signaling commonly regulates cell fate decision via a process known as lateral inhibition, in which a given cell within a homogenous field of cells becomes less susceptible to ligand-activated Notch signaling. As a consequence, this cell initiates a specific differentiation program while continuing to provide Notch ligands to adjacent cells, thereby blocking the differentiation of its neighbors (Fig. 5-3).90 A similar mechanism regulates the differentiation of endocrine cells during pancreas development. Reduction of Notch pathway activity in transgenic mice lacking essential Notch signaling components results in upregulation of expression of Neurogenin3 (Neurog3), a bHLH transcription factor required for endocrine formation91 (see following section), and precocious endocrine differentiation.92,93
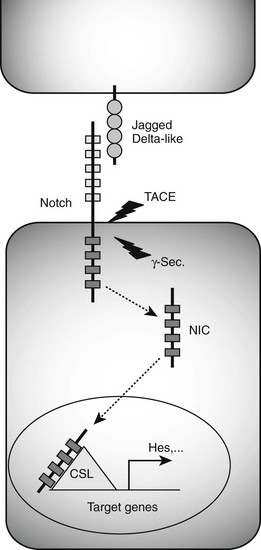
FIGURE 5-3 Notch signaling pathway. Binding of Jagged/Delta-like ligands to Notch receptors activates the Notch signaling cascade. Two proteolytic events mediated by the tumor necrosis factor-α–converting enzyme (TACE) and γ-secretase (γ-Sec.) enzymes result in the generation of an intracellular Notch fragment (NIC) that translocates from the membrane to the nucleus. Interaction with transcription factors from the CSL (CBF1/suppressor of hairless/Lag-1) family activates transcription of Notch target genes, including Hes genes known to block the transcription of Neurog3.
Recent studies have revealed roles for Notch signaling in other steps of pancreatic formation, both before and after its well-recognized role in regulating endocrine specification. Even before pancreas is specified, Notch signaling in the foregut endoderm helps to define the area of the pancreas domain.94 Ectopic expression of a constitutively active, truncated form of the Notch1 receptor at the onset of pancreas formation blocks both exocrine and endocrine cell differentiation, suggesting that Notch signaling normally may allow the expansion of undifferentiated precursor cells.95–97 After endocrine specification, constitutive activation of Notch signaling also blocks endocrine cell differentiation in endocrine precursor cells and can change the differentiation potential of some α/β cells away from the endocrine phenotype toward the ductal fate.98 By contrast, Notch activation in fully matured endocrine cells is not sufficient to change their differentiation status.
Although the exact mechanism of Notch regulation during pancreas formation remains unresolved, recent evidence suggests that FGF signaling moderates Notch activity during pancreas formation. Ectopic expression in the pancreas epithelium of FGF10, an FGF ligand normally found in pancreatic mesenchyme, activates expression of Notch ligands and receptors and the Notch target gene Hes1.99,100 Given the recent evidence of interactions between embryonic signaling pathways during the formation of other organs, it will be critical to determine how these exchanges regulate pancreas development and pancreas cell differentiation.
Transcription Factors
The target of notch inhibition in the embryonic pancreas, bHLH factor Neurog3, functions as a proendocrine factor: It is sufficient by itself to drive precursor cells to an endocrine fate.92,101 When expressed broadly in the developing pancreatic bud, Neurog3 can force all cells in the developing pancreas to differentiate prematurely into islet cells. It is interesting to note that Neurog3 is expressed transiently, predominantly during development in the pancreas in scattered ductal cells and occasional periductal cells, but it is never found in mature, hormone-producing cells. Together, these data support a model in which Neurog3 acts upstream of other islet differentiation factors, initiating the differentiation of endocrine cells, but switching off before the time of final differentiation. This conclusion is supported by the observation that mice homozygous for a targeted disruption of the Neurog3 gene fail to form any endocrine cells in the pancreas and do not express the other islet differentiation factors,91 as well as by lineage tracing experiments.102
Although Notch signaling restricts Neurog3 to scattered cells within the pancreatic epithelium, positive signals must initiate Neurog3 expression in the absence of Notch signaling. Studies of the Neurog3 promoter implicate several transcription factors in this role, including Hnf1, Foxa, Onecut1, and Sox9,103–106 which function together as an interacting network105 (Fig. 5-4). Roles for Onecut1 and Sox9 have been confirmed in vivo.104,106
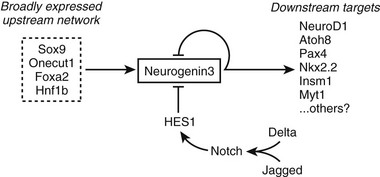
FIGURE 5-4 Control of Neurog3 expression. A model is shown for the positive and negative regulation of Neurog3 expression in pancreatic endocrine progenitor cells.
NeuroD1 expression initiates slightly later than Neurog3 during pancreatic development, but, unlike Neurog3, it persists in the mature islet cells, where it plays a role in the expression of a number of differentiated endocrine cell products, including insulin.107–110 Neurog3 can bind to and activate the neuroD1 gene promoter, and it can induce neuroD1 expression in Xenopus embryos111 and pancreatic ductal cells.112,113 Conversely, Neurog3 null embryos completely lack neuroD1 expression in the pancreas,91 and Neurog3 expression is unchanged in neuroD1 null embryos.101 Expression of the distally related bHLH factor Atoh8 and the zinc-finger factors Myt1 and IA1 also is activated by Neurog3, and Atoh8 and IA1 appear to play a role in feedback regulation of Neurog3 expression.114–116 In addition, Myt1 expression persists in the differentiated endocrine cells and may play a role in their maturation.117
Just as it can with the neuroD1 gene promoter, Neurog3 binds to and activates the pax4 gene promoter118; it can activate the intact gene in cultured cells,113,119 and in its absence, pax4 expression is lost in the pancreas.91 In addition, studies of the pax4 gene promoter implicate the more broadly expressed endoderm factors HNF1 and HNF4, along with Neurog3, in cooperative activation of pax4 expression.118,119 In the case of the gene encoding Nkx2.2, Neurog3 binds to and activates one of three alternate promoters in the gene, and it cooperates with Foxa transcription factors in activating the gene.120
Unlike NeuroD1, Atoh8, and Myt1, however, Pax4 and Nkx2.2 expression is not induced in all islet lineages. Pax4, therefore, falls into a class of factors that play specific roles in the differentiation of the different islet cell subtypes. These factors can be grouped into the early group—factors like Pax4 and the NK homeodomain factors Nkx2.2121 and Nkx6.1,122 which are coexpressed with Neurog3 in endocrine progenitor cells,101 and the late group—genes like the basic leucine zipper transcription factor MafA,123 pdx1 (in a second phase of expression), pax6,124,125 the paired-related homeodomain transcription factor Arx,126 and the pou-homeodomain factor Brn4/Pou3f4,101,127 which are expressed at the final stage of differentiation and are largely restricted to differentiated islet cells (Table 5-1). It is not clear whether any of these genes can actually determine the cell-type fate of the cell in which they are expressed, or if they simply complete the differentiation process initiated by other genes.
Table 5-1
Pancreas Transcription Factors
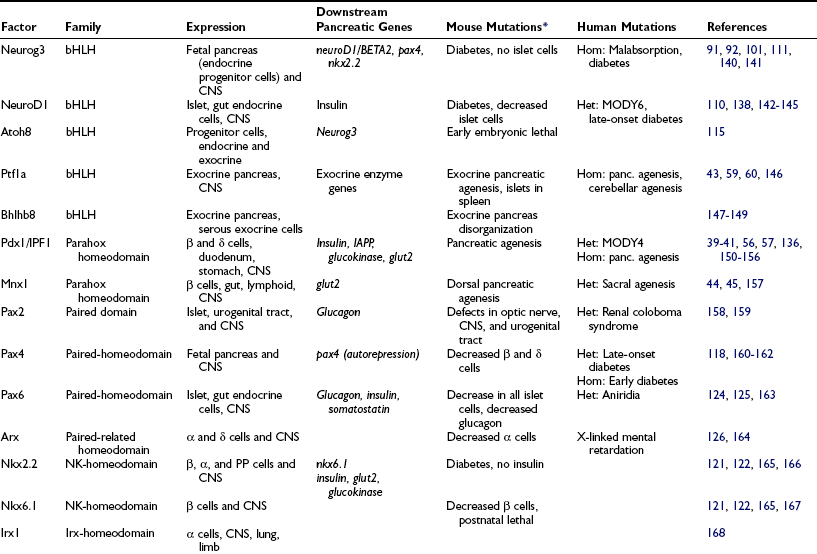


*All mouse phenotypes are for homozygous mutant animals unless stated otherwise.
Once Neurog3 is expressed in an appropriate progenitor, that cell is destined to become an islet cell. But the decision as to which of the four islet cell types it will become apparently is controlled by other factors. Ectopic broad expression of Neurog3 in the developing pancreatic bud causes uniform and precocious endocrine differentiation, but the type of endocrine cells depends on when Neurog3 is induced: early expression of Neurog3 produces α cells,92,101 but later expression of Neurog3 produces other islet cell types.127a It can be hypothesized, therefore, that the cell intrinsic competence of the progenitor cells to generate different islet cell types changes over time, or that extracellular signals that change with time guide the differentiation of the Neurog3-expressing cells into the different islet cell types. The identity of the signals and factors that determine individual islet subtype fates remains to be determined.
Islet Formation
Cell culture assays have implicated matrix metalloproteinases (MMPs) in this process, although experimental evidence from in vivo studies supporting these results is currently missing.128 Cell attachment to and migration via the ECM are mediated by integrins, heterodimeric transmembrane proteins that serve as extracellular matrix receptors.129 Functional evidence for an essential role of integrins during migration of islet precursors comes from in vivo studies in which islet formation in human fetal pancreas transplanted under the kidney capsule of recipient mice was inhibited by injection of integrin-blocking peptides.130 The integrin family comprises a large number of different members with distinct substrate specificities. A recent study reveals that fetal and mature human β cells express different repertoires of integrins in fetal versus mature.131 Changes in integrin composition might explain why fetal β cells are more motile than their adult counterparts—a difference that also may provide insight into the molecular mechanisms that prevent dispersion of endocrine cells once mature islet structures have formed.
Nonetheless, a number of open questions remain with regard to the regulation of islet progenitor migration and islet formation. For example, studies using transgenic animals have shown that a given islet originates from several independent endocrine precursors.132 Thus, endocrine cells delaminating from the ductal structures during embryogenesis must follow a specific guiding mechanism that coordinates their migration path toward a specific location. Although cell migration has been well studied in neural cells that respond to attractive and repulsive guidance cues, a putative, localized source of chemoattractant molecules that stimulate islet precursor migration has not been identified. Furthermore, evidence from studies in human diabetic patients suggests neogenesis of endocrine cells throughout life.133 Although still controversial, some endocrine cells appear to form away from already existing islets, raising the question of whether these cells migrate toward already formed islets, or cluster with other newly formed endocrine cells to initiate aggregation of a new islet. In either case, it is likely that specific, as yet unidentified, guidance mechanisms control these processes.
The Role of Pancreatic Development in Human Diabetes
As a consequence, it is not surprising that mutations in a number of human genes involved in pancreatic islet development, especially the transcription factor genes, can lead to diabetes (see Table 5-1). The transcription factors shown in Fig. 5-5 represent five of the six genes known to cause maturity-onset diabetes of the young (MODY).38,134–138 In addition, mutations in the coding sequence of isl1 and pax4 have been implicated in families with later-onset diabetes.138,139 Analysis of these mutations has provided insight into the process by which β cell dysfunction may lead to the development of diabetes.
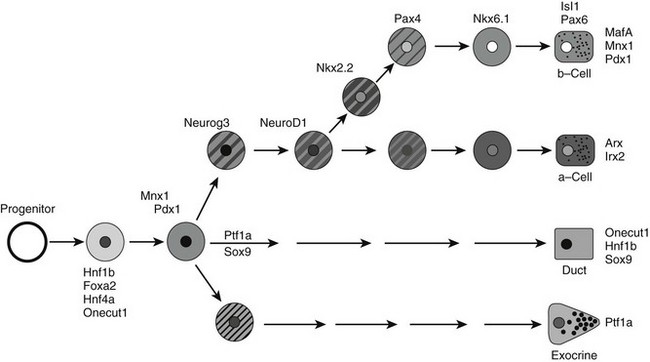
FIGURE 5-5 A simplified model for the role of islet transcription factors in endocrine differentiation in the developing pancreas. The proposed position for each transcription factor is based on its timing of expression, timing of predominant functional role, or both. Clearly, some factors function at multiple steps, but only key steps are shown for simplicity.
References
1. Tam, PP, Kanai-Azuma, M, Kanai, Y. Early endoderm development in vertebrates: lineage differentiation and morphogenetic function. Curr Opin Genet Dev. 2003;13:393–400.
2. Ober, EA, Field, HA, Stainier, DY. From endoderm formation to liver and pancreas development in zebrafish. Mech Dev. 2003;120:5–18.
3. Wells, JM, Melton, DA. Vertebrate endoderm development. Annu Rev Cell Dev Biol. 1999;15:393–410.
4. Pictet, R, Rutter, WJ. Development of the embryonic endocrine pancreas. In: Society AP, Steiner DF, Frenkel N, eds. Handbook of Physiology. Washington DC: Williams and Wilkins; 1972:25–66.
5. Slack, JM. Developmental biology of the pancreas. Development. 1995;121:1569–1580.
6. Kim, SK, Hebrok, M, Melton, DA. Notochord to endoderm signaling is required for pancreas development. Development. 1997;124:4243–4252.
7. Kumar, M, Jordan, N, Melton, D, et al. Signals from lateral plate mesoderm instruct endoderm toward a pancreatic fate. Dev Biol. 2003;259:109–122.
8. Lammert, E, Cleaver, O, Melton, D. Role of endothelial cells in early pancreas and liver development. Mech Dev. 2003;120:59–64.
9. Hill, D, Lebenthal, E. Congenital abnormalities of the exocrine pancreas. In: Go VLW, Dimagno EP, Gardner JD, Lebenthal E, Reber HA, Scheele GA, eds. The Pancreas: Biology, Pathobiology, and Disease. 2nd ed. New York: Raven Press Ltd; 1993:1029–1040.
10. Golosow, N, Grobstein, C. Epitheliomesenchymal interaction in pancreatic morphogenesis. Dev Biol. 1962;4:242–255.
11. Wessells, NK, Cohen, JH. Early pancreas organogenesis: morphogenesis, tissue interactions and mass effects. Dev Biol. 1967;15:237–270.
12. Nekrep, N, Wang, J, Miyatsuka, T, et al. Signals from the neural crest regulate beta-cell mass in the pancreas. Development. 2008;135:2151–2160.
13. Pictet, RL, Rall, LB, Phelps, P, et al. The neural crest and the origin of the insulin-producing and other gastrointestinal hormone-producing cells. Science. 1976;191:191–192.
14. Fontaine, J, Le Douarin, NM. Analysis of endoderm formation in the avian blastoderm by the use of quail-chick chimaeras. The problem of the neurectodermal origin of the cells of the APUD series. J Embryol Exp Morphol. 1977;41:209–222.
15. Le Douarin, NM. On the origin of pancreatic endocrine cells. Cell. 1988;53:169–171.
16. Wells, JM, Melton, DA. Early mouse endoderm is patterned by soluble factors from adjacent germ layers. Development. 2000;127:1563–1572.
17. Hebrok, M, Kim, SK, Melton, DA. Notochord repression of endodermal Sonic hedgehog permits pancreas development. Genes Dev. 1998;12:1705–1713.
18. Deutsch, G, Jung, J, Zheng, M, et al. A bipotential precursor population for pancreas and liver within the embryonic endoderm. Development. 2001;128:871–881.
19. Lammert, E, Cleaver, O, Melton, D. Induction of pancreatic differentiation by signals from blood vessels. Science. 2001;294:564–567.
20. Yoshitomi, H, Zaret, KS. Endothelial cell interactions initiate dorsal pancreas development by selectively inducing the transcription factor Ptf1a. Development. 2004;131:807–817.
21. Dong, PD, Munson, CA, Norton, W, et al. Fgf10 regulates hepatopancreatic ductal system patterning and differentiation. Nat Genet. 2007;39:397–402.
22. Pictet, RL, Clark, WR, Williams, RH, et al. An ultrastructural analysis of the developing embryonic pancreas. Dev Biol. 1972;29:436–467.
23. Li, Z, Manna, P, Kobayashi, H, et al. Multifaceted pancreatic mesenchymal control of epithelial lineage selection. Dev Biol. 2004;269:252–263.
24. Gittes, GK, Galante, PE, Hanahan, D, et al. Lineage-specific morphogenesis in the developing pancreas: role of mesenchymal factors. Development. 1996;122:439–447.
25. Ott, MO, Rey-Campos, J, Cereghini, S, et al. vHNF1 is expressed in epithelial cells of distinct embryonic origin during development and precedes HNF1 expression. Mech Dev. 1991;36:47–58.
26. Monaghan, AP, Kaestner, KH, Grau, E, et al. Postimplantation expression patterns indicate a role for the mouse forkhead/HNF-3 alpha, beta and gamma genes in determination of the definitive endoderm, chordamesoderm and neuroectoderm. Development. 1993;119:567–578.
27. Duncan, SA, Manova, K, Chen, WS, et al. Expression of transcription factor HNF-4 in the extraembryonic endoderm, gut, and nephrogenic tissue of the developing mouse embryo: HNF-4 is a marker for primary endoderm in the implanting blastocyst. Proc Natl Acad Sci U S A. 1994;91:7598–7602.
28. Taraviras, S, Monaghan, AP, Schutz, G, et al. Characterization of the mouse HNF-4 gene and its expression during mouse embryogenesis. Mech Dev. 1994;48:67–79.
29. Rausa, F, Samadani, U, Ye, H, et al. The cut-homeodomain transcriptional activator HNF-6 is coexpressed with its target gene HNF-3 beta in the developing murine liver and pancreas. Dev Biol. 1997;192:228–246.
30. Landry, C, Clotman, F, Hioki, T, et al. HNF-6 is expressed in endoderm derivatives and nervous system of the mouse embryo and participates to the cross-regulatory network of liver-enriched transcription factors. Dev Biol. 1997;192:247–257.
31. Decker, K, Goldman, DC, Grasch, CL, et al. Gata6 is an important regulator of mouse pancreas development. Dev Biol. 2006;298:415–429.
32. Ketola, I, Otonkoski, T, Pulkkinen, MA, et al. Transcription factor GATA-6 is expressed in the endocrine and GATA-4 in the exocrine pancreas. Mol Cell Endocrinol. 2004;226:51–57.
33. Zaret, KS. Molecular genetics of early liver development. Annu Rev Physiol. 1996;58:231–251.
34. Cereghini, S. Liver-enriched transcription factors and hepatocyte differentiation. Faseb J. 1996;10:267–282.
35. Zaret, K. Developmental competence of the gut endoderm: genetic potentiation by GATA and HNF3/fork head proteins. Dev Biol. 1999;209:1–10.
36. Boj, SF, Parrizas, M, Maestro, MA, et al. A transcription factor regulatory circuit in differentiated pancreatic cells. Proc Natl Acad Sci U S A. 2001;98:14481–14486.
37. Shih, DQ, Screenan, S, Munoz, KN, et al. Loss of HNF-1alpha function in mice leads to abnormal expression of genes involved in pancreatic islet development and metabolism. Diabetes. 2001;50:2472–2480.
38. Shih, DQ, Stoffel, M. Dissecting the transcriptional network of pancreatic islets during development and differentiation. Proc Natl Acad Sci U S A. 2001;98:14189–14191.
39. Guz, Y, Montminy, MR, Stein, R, et al. Expression of murine STF-1, a putative insulin gene transcription factor, in beta cells of pancreas, duodenal epithelium and pancreatic exocrine and endocrine progenitors during ontogeny. Development. 1995;121:11–18.
40. Ahlgren, U, Jonsson, J, Edlund, H. The morphogenesis of the pancreatic mesenchyme is uncoupled from that of the pancreatic epithelium in IPF1/PDX1-deficient mice. Development. 1996;122:1409–1416.
41. Offield, MF, Jetton, TL, Labosky, PA, et al. PDX-1 is required for pancreatic outgrowth and differentiation of the rostral duodenum. Development. 1996;122:983–995.
42. Kawaguchi, Y, Cooper, B, Gannon, M, et al. The role of the transcriptional regulator Ptf1a in converting intestinal to pancreatic progenitors. Nat Genet. 2002;32:128–134.
43. Krapp, A, Knofler, M, Frutiger, S, et al. The p48 DNA-binding subunit of transcription factor PTF1 is a new exocrine pancreas-specific basic helix-loop-helix protein. EMBO J. 1996;15:4317–4329.
44. Li, H, Arber, S, Jessell, TM, et al. Selective agenesis of the dorsal pancreas in mice lacking homeobox gene Hlxb9. Nat Genet. 1999;23:67–70.
45. Harrison, KA, Thaler, J, Pfaff, SL, et al. Pancreas dorsal lobe agenesis and abnormal islets of Langerhans in Hlxb9-deficient mice. Nat Genet. 1999;23:71–75.
46. Sharma, S, Jhala, US, Johnson, T, et al. Hormonal regulation of an islet-specific enhancer in the pancreatic homeobox gene STF-1. Mol Cell Biol. 1997;17:2598–2604.
47. Wu, KL, Gannon, M, Peshavaria, M, et al. Hepatocyte nuclear factor 3beta is involved in pancreatic beta-cell-specific transcription of the pdx-1 gene. Mol Cell Biol. 1997;17:6002–6013.
48. Stoffers, DA, Heller, RS, Miller, CP, et al. Developmental expression of the homeodomain protein IDX-1 in mice transgenic for an IDX-1 promoter/lacZ transcriptional reporter. Endocrinology. 1999;140:5374–5381.
49. Ben-Shushan, E, Marshak, S, Shoshkes, M, et al. A pancreatic beta -cell-specific enhancer in the human PDX-1 gene is regulated by hepatocyte nuclear factor 3beta (HNF-3beta ), HNF-1alpha, and SPs transcription factors. J Biol Chem. 2001;276:17533–17540.
50. Marshak, S, Ben-Shushan, E, Shoshkes, M, et al. Regulatory elements involved in human pdx-1 gene expression. Diabetes. 2001;50(Suppl 1)):S37–38.
51. Gerrish, K, Cissell, MA, Stein, R. The role of hepatic nuclear factor 1 alpha and PDX-1 in transcriptional regulation of the pdx-1 gene. J Biol Chem. 2001;276:47775–47784.
52. Gerrish, K, Gannon, M, Shih, D, et al. Pancreatic beta cell-specific transcription of the pdx-1 gene. The role of conserved upstream control regions and their hepatic nuclear factor 3beta sites. J Biol Chem. 2000;275:3485–3492.
53. Samaras, SE, Cissell, MA, Gerrish, K, et al. Conserved sequences in a tissue-specific regulatory region of the pdx-1 gene mediate transcription in pancreatic beta cells: role for hepatocyte nuclear factor 3beta and Pax6. Mol Cell Biol. 2002;22:4702–4713.
54. Marshak, S, Benshushan, E, Shoshkes, M, et al. Functional conservation of regulatory elements in the pdx-1 gene: PDX-1 and hepatocyte nuclear factor 3beta transcription factors mediate beta- cell-specific expression. Mol Cell Biol. 2000;20:7583–7590.
55. Jacquemin, P, Lemaigre, FP, Rousseau, GG. The Onecut transcription factor HNF-6 (OC-1) is required for timely specification of the pancreas and acts upstream of Pdx-1 in the specification cascade. Developmental biology. 2003;258:105–116.
56. Jonsson, J, Carlsson, L, Edlund, T, et al. Insulin-promoter-factor 1 is required for pancreas development in mice. Nature. 1994;371:606–609.
57. Stoffers, DA, Zinkin, NT, Stanojevic, V, et al. Pancreatic agenesis attributable to a single nucleotide deletion in the human IPF1 gene coding sequence. Nature genetics. 1997;15:106–110.
58. Yee, NS, Yusuff, S, Pack, M. Zebrafish pdx1 morphant displays defects in pancreas development and digestive organ chirality, and potentially identifies a multipotent pancreas progenitor cell. Genesis. 2001;30:137–140.
59. Obata, J, Yano, M, Mimura, H, et al. p48 subunit of mouse PTF1 binds to RBP-Jkappa/CBF-1, the intracellular mediator of Notch signalling, and is expressed in the neural tube of early stage embryos. Genes Cells. 2001;6:345–360.
60. Krapp, A, Knofler, M, Ledermann, B, et al. The bHLH protein PTF1-p48 is essential for the formation of the exocrine and the correct spatial organization of the endocrine pancreas. Genes Dev. 1998;12:3752–3763.
61. Ye, F, Duvillie, B, Scharfmann, R. Fibroblast growth factors 7 and 10 are expressed in the human embryonic pancreatic mesenchyme and promote the proliferation of embryonic pancreatic epithelial cells. Diabetologia. 2005;48:277–281.
62. Bhushan, A, Itoh, N, Kato, S, et al. Fgf10 is essential for maintaining the proliferative capacity of epithelial progenitor cells during early pancreatic organogenesis. Development. 2001;128:5109–5117.
63. Le Bras, S, Miralles, F, Basmaciogullari, A, et al. Fibroblast growth factor 2 promotes pancreatic epithelial cell proliferation via functional fibroblast growth factor receptors during embryonic life. Diabetes. 1998;47:1236–1242.
64. Miralles, F, Czernichow, P, Ozaki, K, et al. Signaling through fibroblast growth factor receptor 2b plays a key role in the development of the exocrine pancreas. Proc Natl Acad Sci U S A. 1999;96:6267–6272.
65. Celli, G, LaRochelle, WJ, Mackem, S, et al. Soluble dominant-negative receptor uncovers essential roles for fibroblast growth factors in multi-organ induction and patterning. EMBO J. 1998;17:1642–1655.
66. Sanvito, F, Herrera, PL, Huarte, J, et al. TGF-beta 1 influences the relative development of the exocrine and endocrine pancreas in vitro. Development. 1994;120:3451–3462.
67. Miralles, F, Czernichow, P, Scharfmann, R. Follistatin regulates the relative proportions of endocrine versus exocrine tissue during pancreatic development. Development. 1998;125:1017–1024.
68. Shiozaki, S, Tajima, T, Zhang, YQ, et al. Impaired differentiation of endocrine and exocrine cells of the pancreas in transgenic mouse expressing the truncated type II activin receptor. Biochim Biophys Acta. 1999;1450:1–11.
69. Yamaoka, T, Idehara, C, Yano, M, et al. Hypoplasia of pancreatic islets in transgenic mice expressing activin receptor mutants. J Clin Invest. 1998;102:294–301.
70. Kim, SK, Hebrok, M, Li, E, et al. Activin receptor patterning of foregut organogenesis. Genes Dev. 2000;14:1866–1871.
71. Izeradjene, K, Combs, C, Best, M, et al. Kras(G12D) and Smad4/Dpc4 haploinsufficiency cooperate to induce mucinous cystic neoplasms and invasive adenocarcinoma of the pancreas. Cancer Cell. 2007;11:229–243.
72. Bardeesy, N, Cheng, KH, Berger, JH, et al. Smad4 is dispensable for normal pancreas development yet critical in progression and tumor biology of pancreas cancer. Genes Dev. 2006;20:3130–3146.
73. Smart, NG, Apelqvist, AA, Gu, X, et al. Conditional expression of Smad7 in pancreatic beta cells disrupts TGF-beta signaling and induces reversible diabetes mellitus. PLoS Biol. 2006;4:e39.
74. Apelqvist, A, Ahlgren, U, Edlund, H. Sonic hedgehog directs specialised mesoderm differentiation in the intestine and pancreas. Curr Biol. 1997;7:801–804.
75. Kawahira, H, Ma, NH, Tzanakakis, ES, et al. Combined activities of Hedgehog signaling inhibitors regulate pancreas development. Development. 2003;130:4871–4879.
76. Chuang, PT, Kawcak, T, McMahon, AP. Feedback control of mammalian Hedgehog signaling by the Hedgehog-binding protein, Hip1, modulates Fgf signaling during branching morphogenesis of the lung. Genes Dev. 2003;17:342–347.
77. Hebrok, M, Kim, SK, St Jacques, B, et al. Regulation of pancreas development by hedgehog signaling. Development. 2000;127:4905–4913.
78. Thomas, MK, Lee, JH, Rastalsky, N, et al. Hedgehog signaling regulation of homeodomain protein islet duodenum homeobox-1 expression in pancreatic beta-cells. Endocrinology. 2001;142:1033–1040.
79. Thomas, MK, Rastalsky, N, Lee, JH, et al. Hedgehog signaling regulation of insulin production by pancreatic beta-cells. Diabetes. 2000;49:2039–2047.
80. Zhang, J, Rosenthal, A, de Sauvage, FJ, et al. Downregulation of Hedgehog signaling is required for organogenesis of the small intestine in Xenopus. Dev Biol. 2001;229:188–202.
81. Heller, RS, Dichmann, DS, Jensen, J, et al. Expression patterns of Wnts, Frizzleds, sFRPs, and misexpression in transgenic mice suggesting a role for Wnts in pancreas and foregut pattern formation. Dev Dyn. 2002;225:260–270.
82. Heller, RS, Klein, T, Ling, Z, et al. Expression of Wnt, Frizzled, sFRP, and DKK genes in adult human pancreas. Gene Expr. 2003;11:141–147.
83. Papadopoulou, S, Edlund, H. Attenuated wnt signaling perturbs pancreatic growth but not pancreatic function. Diabetes. 2005;54:2844–2851.
84. Murtaugh, LC, Law, AC, Dor, Y, et al. β-Catenin is essential for pancreatic acinar but not islet development. Development. 2005;132:4663–4674.
85. Wells, JM, Esni, F, Boivin, GP, et al. Wnt/beta-catenin signaling is required for development of the exocrine pancreas. BMC Dev Biol. 2007;7:4.
86. Dessimoz, J, Bonnard, C, Huelsken, J, et al. Pancreas-specific deletion of beta-catenin reveals Wnt-dependent and Wnt-independent functions during development. Curr Biol. 2005;15:1677–1683.
87. Heiser, PW, Lau, J, Taketo, MM, et al. Stabilization of beta-catenin impacts pancreas growth. Development. 2006;133:2023–2032.
88. Jonckheere, N, Mayes, E, Shih, HP, et al. Analysis of mPygo2 mutant mice suggests a requirement for mesenchymal Wnt signaling in pancreatic growth and differentiation. Dev Biol. 2008;318:224–235.
89. Rulifson, IC, Karnik, SK, Heiser, PW, et al. Wnt signaling regulates pancreatic beta cell proliferation. Proc Natl Acad Sci U S A. 2007;104:6247–6252.
90. Artavanis-Tsakonas, S, Rand, MD, Lake, RJ. Notch signaling: cell fate control and signal integration in development. Science. 1999;284:770–776.
91. Gradwohl, G, Dierich, A, LeMeur, M, et al. Neurogenin3 is required for the development of the four endocrine cell lineages of the pancreas. Proc Natl Acad Sci U S A. 2000;97:1607–1611.
92. Apelqvist, A, Li, H, Sommer, L, et al. Notch signaling controls pancreatic cell differentiation. Nature. 1999;400:877–881.
93. Jensen, J, Pedersen, EE, Galante, P, et al. Control of endodermal endocrine development by Hes-1. Nat Genet. 2000;24:36–44.
94. Fukuda, A, Kawaguchi, Y, Furuyama, K, et al. Ectopic pancreas formation in Hes1 -knockout mice reveals plasticity of endodermal progenitors of the gut, bile duct, and pancreas. J Clin Invest. 2006;116:1484–1493.
95. Hald, J, Hjorth, JP, German, MS, et al. Activated Notch1 prevents differentiation of pancreatic acinar cells and attenuate endocrine development. Developmental biology. 2003;260:426–437.
96. Murtaugh, LC, Stanger, BZ, Kwan, KM, et al. Notch signaling controls multiple steps of pancreatic differentiation. Proc Natl Acad Sci U S A. 2003;100:14920–14925.
97. Esni, F, Ghosh, B, Biankin, AV, et al. Notch inhibits Ptf1 function and acinar cell differentiation in developing mouse and zebrafish pancreas. Development. 2004;131:4213–4224.
98. Greenwood, AL, Li, S, Jones, K, et al. Notch signaling reveals developmental plasticity of Pax4(+) pancreatic endocrine progenitors and shunts them to a duct fate. Mech Dev. 2007;124:97–107.
99. Hart, A, Papadopoulou, S, Edlund, H. Fgf10 maintains notch activation, stimulates proliferation, and blocks differentiation of pancreatic epithelial cells. Dev Dyn. 2003;228:185–193.
100. Norgaard, GA, Jensen, JN, Jensen, J. FGF10 signaling maintains the pancreatic progenitor cell state revealing a novel role of Notch in organ development. Developmental biology. 2003;264:323–338.
101. Schwitzgebel, VM, Scheel, DW, Conners, JR, et al. Expression of neurogenin3 reveals an islet cell precursor population in the pancreas. Development. 2000;127:3533–3542.
102. Gu, G, Dubauskaite, J, Melton, DA. Direct evidence for the pancreatic lineage: NGN3+ cells are islet progenitors and are distinct from duct progenitors. Development. 2002;129:2447–2457.
103. Lee, JC, Smith, SB, Watada, H, et al. Regulation of the pancreatic pro-endocrine gene neurogenin3. Diabetes. 2001;50:928–936.
104. Jacquemin, P, Durviaux, SM, Jensen, J, et al. Transcription factor hepatocyte nuclear factor 6 regulates pancreatic endocrine cell differentiation and controls expression of the proendocrine gene ngn3. Mol Cell Biol. 2000;20:4445–4454.
105. Lynn, FC, Smith, SB, Wilson, ME, et al. Sox9 coordinates a transcriptional network in pancreatic progenitor cells. Proc Natl Acad Sci U S A. 2007;104:10500–10505.
106. Seymour, PA, Freude, KK, Dubois, CL, et al. A dosage-dependent requirement for Sox9 in pancreatic endocrine cell formation. Dev Biol. 2008;323:19–30.
107. Ohneda, K, Mirmira, RG, Wang, J, et al. The homeodomain of PDX-1 mediates multiple protein-protein interactions in the formation of a transcriptional activation complex on the insulin promoter. Mol Cell Biol. 2000;20:900–911.
108. Glick, E, Leshkowitz, D, Walker, MD. Transcription factor BETA2 acts cooperatively with E2A and PDX1 to activate the insulin gene promoter. J Biol Chem. 2000;275:2199–2204.
109. Qiu, Y, Guo, M, Huang, S, et al. Insulin gene transcription is mediated by interactions between the p300 coactivator and PDX-1, BETA2, and E47. Mol Cell Biol. 2002;22:412–420.
110. Naya, FJ, Stellrecht, CM, Tsai, MJ. Tissue-specific regulation of the insulin gene by a novel basic helix-loop-helix transcription factor. Genes Dev. 1995;9:1009–1019.
111. Huang, HP, Liu, M, El-Hodiri, HM, et al. Regulation of the pancreatic islet-specific gene BETA2 (neuroD) by neurogenin 3. Mol Cell Biol. 2000;20:3292–3307.
112. Heremans, Y, Van De Casteele, M, in’t Veld, P, et al. Recapitulation of embryonic neuroendocrine differentiation in adult human pancreatic duct cells expressing neurogenin 3. J Cell Biol. 2002;159:303–312.
113. Gasa, R, Mrejen, C, Leachman, N, et al. Proendocrine genes coordinate the pancreatic islet differentiation program in vitro. Proc Natl Acad Sci U S A. 2004;101:13245–13250.
114. Wang, S, Hecksher-Sorensen, J, Xu, Y, et al. Myt1 and Ngn3 form a feed-forward expression loop to promote endocrine islet cell differentiation. Dev Biol. 2008;317:531–540.
115. Lynn, FC, Sanchez, L, Gomis, R, et al. Identification of the bHLH factor Math6 as a novel component of the embryonic pancreas transcriptional network. PLoS ONE. 2008;3:e2430.
116. Mellitzer, G, Bonne, S, Luco, RF, et al. IA1 is NGN3-dependent and essential for differentiation of the endocrine pancreas. The EMBO journal. 2006;25:1344–1352.
117. Wang, S, Zhang, J, Zhao, A, et al. Loss of Myt1 function partially compromises endocrine islet cell differentiation and pancreatic physiological function in the mouse. Mech Dev. 2007;124:898–910.
118. Smith, SB, Watada, H, Scheel, DW, et al. Autoregulation and maturity onset diabetes of the young transcription factors control the human PAX4 promoter. J Biol Chem. 2000;275:36910–36919.
119. Smith, SB, Gasa, R, Watada, H, et al. Neurogenin3 and hepatic nuclear factor 1 cooperate in activating pancreatic expression of Pax4. J Biol Chem. 2003;278:38254–38259.
120. Watada, H, Scheel, DW, Leung, J, et al. Distinct gene expression programs function in progenitor and mature islet cells. J Biol Chem. 2003;278:17130–17140.
121. Sussel, L, Kalamaras, J, Hartigan-O’Connor, DJ, et al. Mice lacking the homeodomain transcription factor Nkx2.2 have diabetes due to arrested differentiation of pancreatic beta cells. Development. 1998;125:2213–2221.
122. Sander, M, Sussel, L, Conners, J, et al. Homeobox gene Nkx6.1 lies downstream of Nkx2.2 in the major pathway of beta-cell formation in the pancreas. Development. 2000;127:5533–5540.
123. Ahlgren, U, Pfaff, SL, Jessell, TM, et al. Independent requirement for ISL1 in formation of pancreatic mesenchyme and islet cells. Nature. 1997;385:257–260.
124. Sander, M, Neubuser, A, Kalamaras, J, et al. Genetic analysis reveals that PAX6 is required for normal transcription of pancreatic hormone genes and islet development. Genes Dev. 1997;11:1662–1673.
125. St-Onge, L, Sosa-Pineda, B, Chowdhury, K, et al. Pax6 is required for differentiation of glucagon-producing alpha-cells in mouse pancreas. Nature. 1997;387:406–409.
126. Collombat, P, Mansouri, A, Hecksher-Sorensen, J, et al. Opposing actions of Arx and Pax4 in endocrine pancreas development. Genes Dev. 2003;17:2591–2603.
127. Hussain, MA, Lee, J, Miller, CP, et al. POU domain transcription factor brain 4 confers pancreatic alpha-cell-specific expression of the proglucagon gene through interaction with a novel proximal promoter G1 element. Mol Cell Biol. 1997;17:7186–7194.
127a. Johansson, KA, Dursun, U, Jordan, N, et al. Temporal control of neurogenin3 activity in pancreas progenitors reveals competence windows for the generation of different endocrine cell types. Dev Cell. 2007;12:457–465.
128. Miralles, F, Battelino, T, Czernichow, P, et al. TGF-beta plays a key role in morphogenesis of the pancreatic islets of Langerhans by controlling the activity of the matrix metalloproteinase MMP-2. J Cell Biol. 1998;143:827–836.
129. Hynes, R. Integrins. Bidirectional, allosteric signaling machines. Cell. 2002;110:673.
130. Cirulli, V, Beattie, GM, Klier, G, et al. Expression and function of alpha(v)beta(3) and alpha(v)beta(5) integrins in the developing pancreas. Roles In the adhesion and migration of putative endocrine progenitor cells [In Process Citation]. J Cell Biol. 2000;150:1445–1460.
131. Kaido, T, Perez, B, Yebra, M, et al. Alphav-integrin utilization in human beta-cell adhesion, spreading, and motility. J Biol Chem. 2004;279:17731–17737.
132. Deltour, L, Leduque, P, Paldi, A, et al. Polyclonal origin of pancreatic islets in aggregation mouse chimaeras. Development. 1991;112:1115–1121.
133. Butler, AE, Janson, J, Bonner-Weir, S, et al. Beta-cell deficit and increased beta-cell apoptosis in humans with type 2 diabetes. Diabetes. 2003;52:102–110.
134. Yamagata, K, Furuta, H, Oda, N, et al. Mutations in the hepatocyte nuclear factor-4alpha gene in maturity-onset diabetes of the young (MODY1). Nature. 1996;384:458–460.
135. Yamagata, K, Oda, N, Kaisaki, P, et al. Mutations in the hepatocyte nuclear factor-1alpha gene in maturity-onset diabetes of the young (MODY3). Nature. 1996;384:455–458.
136. Stoffers, DA, Ferrer, J, Clarke, WL, et al. Early-onset type-II diabetes mellitus (MODY4) linked to IPF1 [letter]. Nat Genet. 1997;17:138–139.
137. Horikawa, Y, Iwasaki, N, Hara, M, et al. Mutation in hepatocyte nuclear factor-1 beta gene (TCF2) associated with MODY [letter]. Nat Genet. 1997;17:384–385.
138. Malecki, MT, Jhala, US, Antonellis, A, et al. Mutations in NEUROD1 are associated with the development of type 2 diabetes mellitus. Nat Genet. 1999;23:323–328.
139. Shimomura, H, Sanke, T, Hanabusa, T, et al. Nonsense mutation of islet-1 gene (Q310X) found in a type 2 diabetic patient with a strong family history. Diabetes. 2000;49:1597–1600.
140. Jensen, J, Heller, RS, Funder-Nielsen, T, et al. Independent development of pancreatic alpha- and beta-cells from neurogenin3-expressing precursors: a role for the notch pathway in repression of premature differentiation. Diabetes. 2000;49:163–176.
141. Wang, J, Cortina, G, Wu, SV, et al. Mutant neurogenin-3 in congenital malabsorptive diarrhea. N Engl J Med. 2006;355:270–280.
142. Lee, JE, Hollenberg, SM, Snider, L, et al. Conversion of Xenopus ectoderm into neurons by NeuroD, a basic helix-loop-helix protein. Science. 1995;268:836–844.
143. Naya, FJ, Huang, HP, Qiu, Y, et al. Diabetes, defective pancreatic morphogenesis, and abnormal enteroendocrine differentiation in BETA2/neuroD-deficient mice. Genes Dev. 1997;11:2323–2334.
144. Mutoh, H, Naya, FJ, Tsai, MJ, et al. The basic helix-loop-helix protein BETA2 interacts with p300 to coordinate differentiation of secretin-expressing enteroendocrine cells. Genes Dev. 1998;12:820–830.
145. Qiu, Y, Sharma, A, Stein, R. p300 mediates transcriptional stimulation by the basic helix-loop-helix activators of the insulin gene. Mol Cell Biol. 1998;18:2957–2964.
146. Sellick, GS, Barker, KT, Stolte-Dijkstra, I, et al. Mutations in PTF1A cause pancreatic and cerebellar agenesis. Nat Genet. 2004;36:1301–1305.
147. Pin, CL, Rukstalis, JM, Johnson, C, et al. The bHLH transcription factor Mist1 is required to maintain exocrine pancreas cell organization and acinar cell identity. J Cell Biol. 2001;155:519–530.
148. Pin, CL, Bonvissuto, AC, Konieczny, SF. Mist1 expression is a common link among serous exocrine cells exhibiting regulated exocytosis. Anat Rec. 2000;259:157–167.
149. Lemercier, C, To, RQ, Swanson, BJ, et al. Mist1: a novel basic helix-loop-helix transcription factor exhibits a developmentally regulated expression pattern. Dev Biol. 1997;182:101–113.
150. Waeber, G, Thompson, N, Nicod, P, et al. Transcriptional activation of the GLUT2 gene by the IPF-1/STF-1/IDX-1 homeobox factor. Mol Endocrinol. 1996;10:1327–1334.
151. Ahlgren, U, Jonsson, J, Jonsson, L, et al. beta-cell-specific inactivation of the mouse Ipf1/Pdx1 gene results in loss of the beta-cell phenotype and maturity onset diabetes. Genes Dev. 1998;12:1763–1768.
152. Watada, H, Kajimoto, Y, Umayahara, Y, et al. The human glucokinase gene beta-cell-type promoter: an essential role of insulin promoter factor 1/PDX-1 in its activation in HIT-T15 cells. Diabetes. 1996;45:1478–1488.
153. Watada, H, Kajimoto, Y, Miyagawa, J, et al. PDX-1 induces insulin and glucokinase gene expressions in alphaTC1 clone 6 cells in the presence of betacellulin. Diabetes. 1996;45:1826–1831.
154. Watada, H, Kajimoto, Y, Kaneto, H, et al. Involvement of the homeodomain-containing transcription factor PDX-1 in islet amyloid polypeptide gene transcription. Biochem Biophys Res Commun. 1996;229:746–751.
155. Schwartz, PT, Perez-Villamil, B, Rivera, A, et al. Pancreatic homeodomain transcription factor IDX1/IPF1 expressed in developing brain regulates somatostatin gene transcription in embryonic neural cells. J Biol Chem. 2000;275:19106–19114.
156. Perez-Villamil, B, Schwartz, PT, Vallejo, M. The pancreatic homeodomain transcription factor IDX1/IPF1 is expressed in neural cells during brain development. Endocrinology. 1999;140:3857–3860.
157. Harrison, KA, Druey, KM, Deguchi, Y, et al. A novel human homeobox gene distantly related to proboscipedia is expressed in lymphoid and pancreatic tissues. J Biol Chem. 1994;269:19968–19975.
158. Torres, M, Gómez-Pardo, E, Gruss, P. Pax2 contributes to inner ear patterning and optic nerve trajectory. Development. 1996;122:3381–3391.
159. Ritz-Laser, B, Estreicher, A, Gauthier, B, et al. The paired homeodomain transcription factor Pax-2 is expressed in the endocrine pancreas and transactivates the glucagon gene promoter. J Biol Chem. 2000;275:32708–32715.
160. Smith, SB, Ee, HC, Conners, JR, et al. Paired-homeodomain transcription factor PAX4 acts as a transcriptional repressor in early pancreatic development. Mol Cell Biol. 1999;19:8272–8280.
161. Sosa-Pineda, B, Chowdhury, K, Torres, M, et al. The Pax4 gene is essential for differentiation of insulin-producing beta cells in the mammalian pancreas. Nature. 1997;386:399–402.
162. Shimajiri, Y, Sanke, T, Furuta, H, et al. A missense mutation of Pax4 gene (R121W) is associated with type 2 diabetes in Japanese. Diabetes. 2001;50:2864–2869.
163. Hill, ME, Asa, SL, Drucker, DJ. Essential requirement for Pax6 in control of enteroendocrine proglucagon gene transcription. Mol Endocrinol. 1999;13:1474–1486.
164. Bienvenu, T, Poirier, K, Friocourt, G, et al. ARX, a novel Prd-class-homeobox gene highly expressed in the telencephalon, is mutated in X-linked mental retardation. Hum Mol Genet. 2002;11:981–991.
165. Rudnick, A, Ling, TY, Odagiri, H, et al. Pancreatic beta cells express a diverse set of homeobox genes. Proc Natl Acad Sci U S A. 1994;91:12203–12207.
166. Watada, H, Mirmira, RG, Leung, J, et al. Transcriptional and translational regulation of beta-cell differentiation factor Nkx6.1. J Biol Chem. 2000;275:34224–34230.
167. Jensen, J, Serup, P, Karlsen, C, et al. mRNA profiling of rat islet tumors reveals nkx 6.1 as a beta-cell-specific homeodomain transcription factor. J Biol Chem. 1996;271:18749–18758.
168. Petri, A, Ahnfelt-Ronne, J, Frederiksen, KS, et al. The effect of neurogenin3 deficiency on pancreatic gene expression in embryonic mice. J Mol Endocrinol. 2006;37:301–316.
169. Lebel, M, Agarwal, P, Cheng, CW, et al. The Iroquois homeobox gene Irx2 is not essential for normal development of the heart and midbrain-hindbrain boundary in mice. Mol Cell Biol. 2003;23:8216–8225.
170. German, MS, Wang, J, Chadwick, RB, et al. Synergistic activation of the insulin gene by a LIM-homeodomain protein and a basic helix-loop-helix protein: building a functional insulin minienhancer complex. Genes & Dev. 1992;6:2165–2176.
171. Jin, T, Trinh, DK, Wang, F, et al. The caudal homeobox protein cdx-2/3 activates endogenous proglucagon gene expression in InR1-G9 islet cells. Mol Endocrinol. 1997;11:203–209.
172. Laser, B, Meda, P, Constant, I, et al. The caudal-related homeodomain protein Cdx-2/3 regulates glucagon gene expression in islet cells. J Biol Chem. 1996;271:28984–28994.
173. Jin, T, Drucker, DJ. Activation of proglucagon gene transcription through a novel promoter element by the caudal-related homeodomain protein cdx-2/3. Mol Cell Biol. 1996;16:19–28.
174. Chawengsaksophak, K, James, R, Hammond, VE, et al. Homeosis and intestinal tumours in Cdx2 mutant mice. Nature. 1997;386:84–87.
175. Vallejo, M, Penchuk, L, Habener, JF. Somatostatin gene upstream enhancer element activated by a protein complex consisting of CREB, Isl-1-like, and alpha-CBF-like transcription factors. J Biol Chem. 1992;267:12876–12884.
176. Leonard, J, Serup, P, Gonzalez, G, et al. The LIM family transcription factor Isl-1 requires cAMP response element binding protein to promote somatostatin expression in pancreatic islet cells. Proc Natl Acad Sci USA. 1992;89:6247–6251.
177. Thor, S, Ericson, J, Brannstrom, T, et al. The homeodomain LIM protein isl-I is expressed in subsets of neurons and endocrine cells in the adult rat. Neuron. 1991;7:1–9.
178. Dong, J, Asa, SL, Drucker, DJ. Islet cell and extrapancreatic expression of the LIM domain homeobox gene isl-1. Mol Endocrinol. 1991;5:1633–1641.
179. Karlsson, O, Thor, S, Norberg, T, et al. Insulin gene enhancer binding protein Isl-1 is a member of a novel class of proteins containing both a homeo- and a Cys-His domain. Nature. 1990;344:879–882.
180. Millonig, JH, Millen, KJ, Hatten, ME. The mouse Dreher gene Lmx1a controls formation of the roof plate in the vertebrate CNS [see comments]. Nature. 2000;403:764–769.
181. Phippard, D, Heydemann, A, Lechner, M, et al. Changes in the subcellular localization of the Brn4 gene product precede mesenchymal remodeling of the otic capsule. Hear Res. 1998;120:77–85.
182. Emens, LA, Landers, DW, Moss, LG. Hepatocyte nuclear factor 1 alpha is expressed in a hamster insulinoma line and transactivates the rat insulin I gene. Proc Natl Acad Sci USA. 1992;89:7300–7304.
183. Noguchi, T, Yamada, K, Yamagata, K, et al. Expression of liver type pyruvate kinase in insulinoma cells: involvement of LF-B1 (HNF1). Biochem Biophys Res Com. 1991;181:259–264.
184. Pontoglio, M, Sreenan, S, Roe, M, et al. Defective insulin secretion in hepatocyte nuclear factor 1alpha-deficient mice. J Clin Invest. 1998;101:2215–2222.
185. Coffinier, C, Thepot, D, Babinet, C, et al. Essential role for the homeoprotein vHNF1/HNF1beta in visceral endoderm differentiation. Development. 1999;126:4785–4794.
186. Coffinier, C, Barra, J, Babinet, C, et al. Expression of the vHNF1/HNF1beta homeoprotein gene during mouse organogenesis. Mech Dev. 1999;89:211–213.
187. Lemaigre, FP, Durviaux, SM, Truong, O, et al. Hepatocyte nuclear factor 6, a transcription factor that contains a novel type of homeodomain and a single cut domain. Proc Natl Acad Sci U S A. 1996;93:9460–9464.
188. Rausa, FM, Ye, H, Lim, L, et al. In situ hybridization with 33P-labeled RNA probes for determination of cellular expression patterns of liver transcription factors in mouse embryos [published erratum appears in Methods: 1998.Nov;16(3):359–360]. Methods. 1998;16:29–41.
189. Shih, DQ, Navas, MA, Kuwajima, S, et al. Impaired glucose homeostasis and neonatal mortality in hepatocyte nuclear factor 3alpha-deficient mice. Proc Natl Acad Sci U S A. 1999;96:10152–10157.
190. Kaestner, KH, Katz, J, Liu, Y, et al. Inactivation of the winged helix transcription factor HNF3alpha affects glucose homeostasis and islet glucagon gene expression in vivo. Genes Dev. 1999;13:495–504.
191. Duncan, SA, Navas, MA, Dufort, D, et al. Regulation of a transcription factor network required for differentiation and metabolism. Science. 1998;281:692–695.
192. Rausa, FM, Galarneau, L, Belanger, L, et al. The nuclear receptor fetoprotein transcription factor is coexpressed with its target gene HNF-3beta in the developing murine liver, intestine and pancreas. Mech Dev. 1999;89:185–188.
193. Sund, NJ, Vatamaniuk, MZ, Casey, M, et al. Tissue-specific deletion of Foxa2 in pancreatic beta cells results in hyperinsulinemic hypoglycemia. Genes Dev. 2001;15:1706–1715.
194. Kaestner, KH, Hiemisch, H, Schutz, G. Targeted disruption of the gene encoding hepatocyte nuclear factor 3gamma results in reduced transcription of hepatocyte-specific genes. Mol Cell Biol. 1998;18:4245–4251.
195. Kaestner, KH, Hiemisch, H, Luckow, B, et al. The HNF-3 gene family of transcription factors in mice: gene structure, cDNA sequence, and mRNA distribution. Genomics. 1994;20:377–385.
196. Liu, Y, Shen, W, Brubaker, PL, et al. Foxa3 (HNF-3gamma) binds to and activates the rat proglucagon gene promoter but is not essential for proglucagon gene expression. Biochem J. 2002;366:633–641.
197. Sladek, FM, Zhong, WM, Lai, E, et al. Liver-enriched transcription factor HNF-4 is a novel member of the steroid hormone receptor superfamily. Genes Dev. 1990;4:2353–2365.
198. Miquerol, L, Lopez, S, Cartier, N, et al. Expression of the L-type pyruvate kinase gene and the hepatocyte nuclear factor 4 transcription factor in exocrine and endocrine pancreas. J Biol Chem. 1994;269:8944–8951.
199. Planque, N, Leconte, L, Coquelle, FM, et al. Interaction of Maf transcription factors with Pax-6 results in synergistic activation of the glucagon promoter. J Biol Chem. 2001;276:35751–35760.
200. Olbrot, M, Rud, J, Moss, LG, et al. Identification of beta-cell-specific insulin gene transcription factor RIPE3b1 as mammalian MafA. Proc Natl Acad Sci U S A. 2002;99:6737–6742.
201. Kataoka, K, Han, SI, Shioda, S, et al. MafA is a glucose-regulated and pancreatic beta-cell-specific transcriptional activator for the insulin gene. J Biol Chem. 2002;277:49903–49910.
202. Zhang, C, Moriguchi, T, Kajihara, M, et al. MafA is a key regulator of glucose-stimulated insulin secretion. Mol Cell Biol. 2005;25:4969–4976.
203. Vanhoose, AM, Samaras, S, Artner, I, et al. MafA and MafB regulate Pdx1 transcription through the Area II control region in pancreatic beta cells. J Biol Chem. 2008;283:22612–22619.
204. Nishimura, W, Rowan, S, Salameh, T, et al. Preferential reduction of beta cells derived from Pax6-MafB pathway in MafB deficient mice. Dev Biol. 2008;314:443–456.
205. Artner, I, Blanchi, B, Raum, JC, et al. MafB is required for islet beta cell maturation. Proc Natl Acad Sci U S A. 2007;104:3853–3858.
206. Artner, I, Le Lay, J, Hang, Y, et al. MafB: an activator of the glucagon gene expressed in developing islet alpha- and beta-cells. Diabetes. 2006;55:297–304.
207. Gierl, MS, Karoulias, N, Wende, H, et al. The zinc-finger factor Insm1 (IA-1) is essential for the development of pancreatic beta cells and intestinal endocrine cells. Genes Dev. 2006;20:2465–2478.
208. Gu, G, Wells, JM, Dombkowski, D, et al. Global expression analysis of gene regulatory pathways during endocrine pancreatic development. Development. 2004;131:165–179.