Cleavage and Implantation
Cleavage
Morphology
Compared with most other species, mammalian cleavage is a leisurely process measured in days rather than hours. Development proceeds at the rate of roughly one cleavage division per day for the first 2 days (Figs. 3.1 and 3.2). After the 2-cell stage, mammalian cleavage is asynchronous, with 1 of the 2 cells (blastomeres) dividing to form a 3-cell embryo. When the embryo consists of approximately 16 cells, it is called a morula (derived from the Latin word meaning “mulberry”).
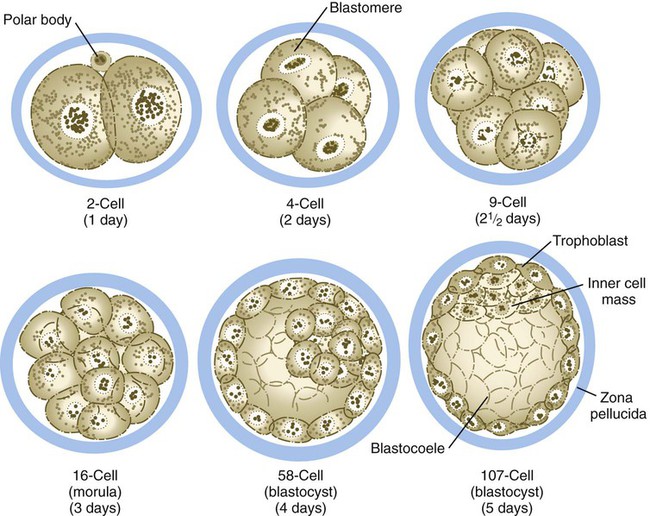
The drawings of the 58-cell and 107-cell stages represent bisected embryos.
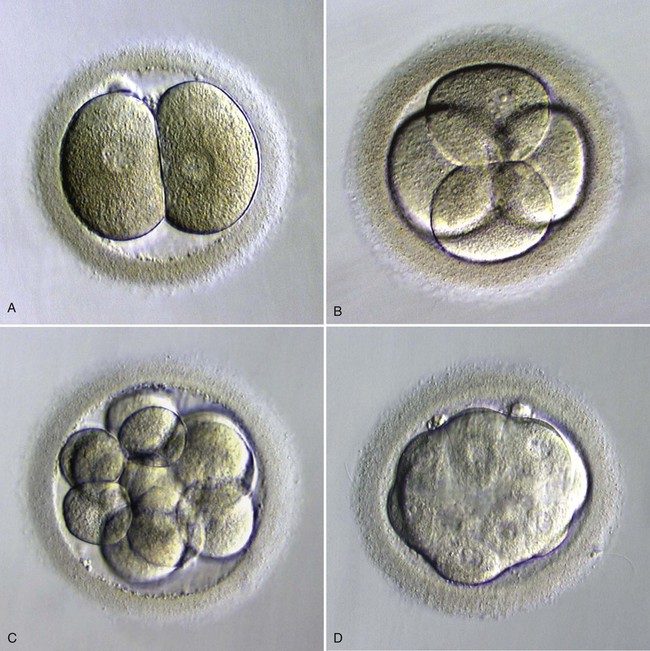
A, Two blastomeres. B, Four blastomeres. C, Twelve blastomeres. D, Morula in late stage of compaction (5 days). Note the indistinct cell outlines. (From Veeck LL, Zaninovic N: An atlas of human blastocysts, Boca Raton, Fla, 2003, Parthenon.)
Starting after the eight-cell stage, the embryos of placental mammals enter into a phase called compaction, during which the individual outer blastomeres tightly adhere through gap and tight junctions and lose their individual identity when viewed from the surface. Compaction is mediated by the concentration of calcium (Ca++)–activated cell adhesion molecules, such as E-cadherin, in a ring around the apical surface of the blastomeres. Through the activity of a sodium (Na+), potassium (K+)–adenosine triphosphatase (ATPase)–based Na+ transport system, Na+ and water (H2O) move across the epitheliumlike outer blastomeres and accumulate in spaces among the inner blastomeres. This process, which occurs about 4 days after fertilization, is called cavitation, and the fluid-filled space is known as the blastocoele (blastocyst cavity). At this stage, the embryo as a whole is known as a blastocyst (Fig. 3.3).
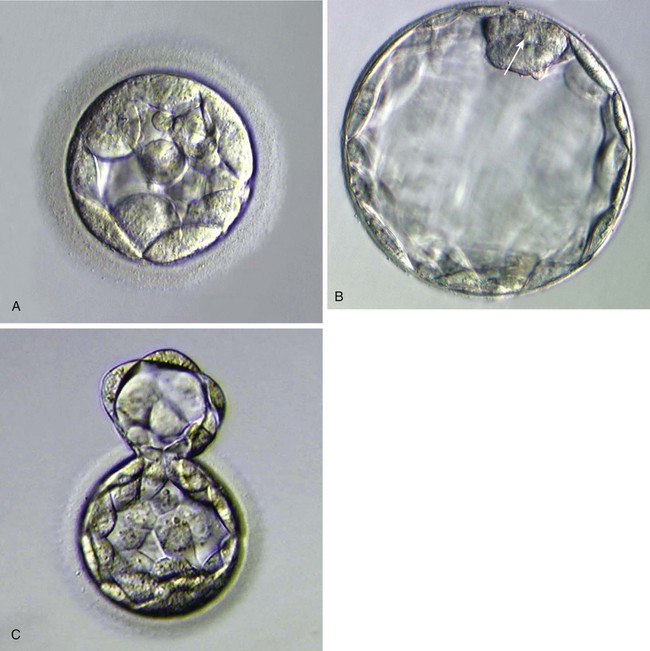
A, Morula, showing the beginning of cavitation. B, Blastocyst, showing a well-defined inner cell mass (arrow) and blastocoele. At this stage, the zona pellucida is very thin. C, A hatching blastocyst beginning to protrude through the zona pellucida. (From Veeck LL, Zaninovic N: An atlas of blastocysts, Boca Raton, Fla, 2003, Parthenon.)
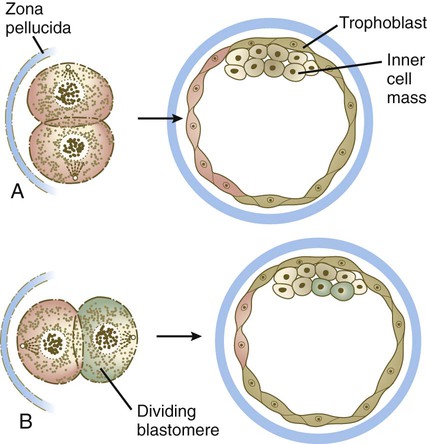
At the blastocyst stage, the embryo, which is still surrounded by the zona pellucida, consists of two types of cells: an outer epithelial layer (the trophoblast) that surrounds a small inner group of cells called the inner cell mass (see Fig. 3.1). Each blastomere at the two-cell and the four-cell stage contributes cells to both the inner cell mass and the trophoblast. The end of the blastocyst that contains the inner cell mass is known as the embryonic pole, and the opposite end is called the abembryonic pole. The appearance of these two cell types reflects major organizational changes that have occurred within the embryo and represents the specialization of the blastomeres into two distinct cell lineages. Cells of the inner cell mass give rise to the body of the embryo itself in addition to several extraembryonic structures, whereas cells of the trophoblast form only extraembryonic structures, including the outer layers of the placenta. There is increasing evidence that fibroblast growth factor-4, a growth factor secreted by cells of the inner cell mass, acts to maintain mitotic activity in the overlying trophoblast.
Molecular, Genetic, and Developmental Control of Cleavage
Along with the increase in cell numbers, mammalian cleavage is a period dominated by several critical developmental events. The earliest is the transition from maternally to zygotically produced gene products. Another is the polarization of individual blastomeres, which sets the stage for the developmental decision that results in the subdivision of the cleaving embryo into two distinct types of cells: the trophoblast and the inner cell mass (see Fig. 3.1). Most studies of the molecular biology and genetics of early mammalian development have been done on mice. Until more information on early primate embryogenesis becomes available, results obtained from experimentation on mice must be used as a guide.
Because of the lack of massive storage of maternal ribosomes and RNAs during oogenesis, development of the mammalian embryo must rely on the activation of zygotic gene products at a very early stage. Most maternal transcription products are degraded by the two-cell stage (Fig. 3.4). Some of these, however, stimulate the activation of the embryonic genome, which begins producing RNAs from a significant number of genes (>1500) by the time cleavage has advanced to the four-cell stage. There does not seem to be a sharp transition between the cessation of reliance on purely maternal gene products and the initiation of transcription from the embryonic genome. Some paternal gene products (e.g., isoforms of β-glucuronidase and β2-microglobulin) appear in the embryo very early, while maternal actin and histone mRNAs are still being used for the production of corresponding proteins. As an indication of the extent to which the early embryo relies on its own gene products, development past the two-cell stage does not occur in the mouse if mRNA transcription is inhibited. In contrast, similar treatment of amphibian embryos does not disrupt development until late cleavage, at which time the embryos begin to synthesize the mRNAs required to control morphogenetic movements and gastrulation.
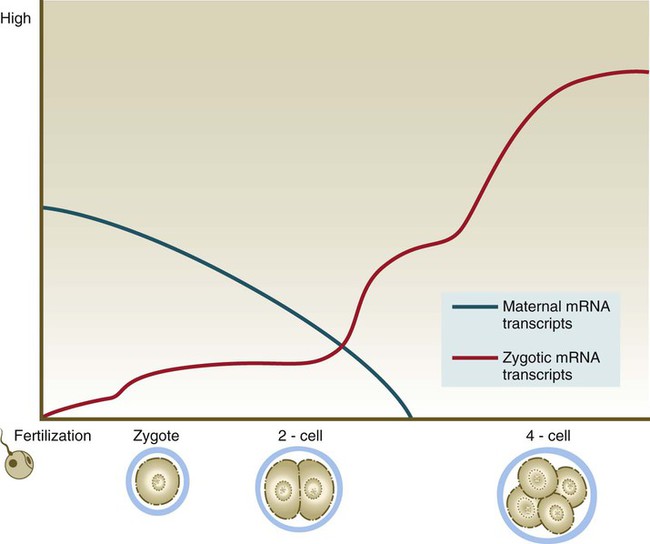
Blue line, maternal mRNAs; red line, zygotic mRNAs.
Mature eggs and sperms are transcriptionally inactive. A major reason for this is that their DNA is highly methylated. Methylation, which occurs on CpG dinucleotides, normally inactivates the associated gene. Such inactivation is often called epigenetic regulation because it does not alter the fundamental DNA sequence. Methylation can inactivate informational genes or their regulators (e.g., enhancers or promoters). Pronounced cycles of global methylation and demethylation occur during the life span of an individual (Fig. 3.5). Within 4 hours after fertilization, the paternally derived genome undergoes rapid, massive demethylation. Demethylation of the maternally derived genome occurs more gradually until the early morula, at which stage all the DNA is maximally demethylated. Remethylation ensues in the inner cell mass, until by the late blastocyst stage it returns to maximal levels. Within the germ cell line, the high methylation levels characteristic of the early embryo fall after the primordial germ cells have entered the genital ridge. During later gametogenesis, remethylation occurs. This remethylation imprints (see p. 43) maternal or paternal characteristics on the gametes and for some genes has profound effects on the embryos produced from these gametes. Epigenetic control is not confined to methylation patterns. Even as early as the zygote, different patterns of histone association with the chromatin account for pronounced differences in gene expression between the male and female pronuclei.
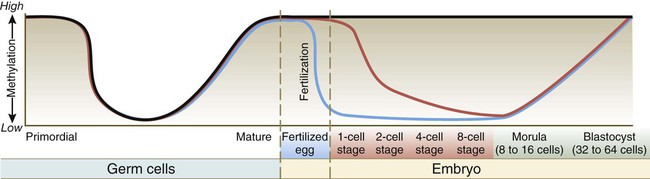
Migrating primordial germ cells are highly methylated, but they lose their methylation on entering the primitive gonad. Methylation is then lost and later reacquired during late stages of gamete maturation. After fertilization, methylation remains high in imprinted genes (black line), but DNA in the male pronucleus undergoes rapid enzymatically mediated demethylation in the zygote (blue line), whereas demethylation in the female chromosomes occurs more slowly (over several days) (red line). This, in addition to changes in histone patterns, accounts for the greater levels of transcription in the paternal genome during very early development. By the blastocyst stage, high methylation levels have returned. (Modified from Santos F, Dean W: Reproduction 127:643-651, 2004.)
The relationship between the position of the blastomeres and their ultimate developmental fate was incorporated into the inside-outside hypothesis. The essence of this hypothesis is that the fate of a blastomere derives from its position within the embryo, rather than from its intrinsic properties. The outer blastomeres ultimately differentiate into the trophoblast, whereas the inner blastomeres form the inner cell mass. If marked blastomeres from disaggregated embryos are placed on the surface of another early embryo, they typically contribute to the formation of the trophoblast. Conversely, if the same marked cells are introduced into the interior of the host embryo, they participate in the formation of the inner cell mass (Fig. 3.6).
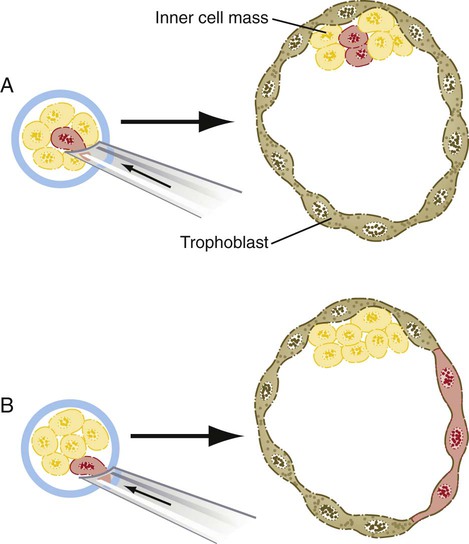
A, If a marked blastomere is inserted into the interior of a morula, it and its progeny become part of the inner cell mass. B, If a marked blastomere is placed on the outside of a host morula, it and its descendants contribute to the trophoblast.
The cell polarity model offers an alternative explanation for the conversion of generic blastomeres to trophoblast or inner cell mass. According to this hypothesis, if the plane of cell division of a blastomere at the eight-cell stage is parallel to the outer surface of the embryo, the outer daughter cell develops a polarity, with its apical surface facing the zona pellucida (Fig. 3.7). The inner daughter cell remains apolar and goes on to form part of the inner cell mass. Experimental evidence suggests that a key element underlying a daughter cell’s becoming an outer cell is inheritance of a patch of outer cell membrane containing microvilli and the actin microfilament-stabilizing protein, ezrin. The proteins that produce polarity in the outer cells are postulated to direct their differentiation toward the trophoblastic lineage. Common to the inside-outside hypothesis and the cell polarity model is the recognition that a cell that does not contact the surface does not differentiate into trophoblast, but rather becomes part of the inner cell mass.
A, If the plane of cleavage of a blastomere is perpendicular to the surface of the embryo, each daughter cell becomes trophoblast. B, If the plane of cleavage is parallel to the surface, the daughter blastomere located at the surface becomes trophoblast, whereas the daughter cell located on the interior becomes part of the inner cell mass.
Oct-4 is expressed in all blastomeres up to the morula stage. As various differentiated cell types begin to emerge in the embryo, the levels of oct4 gene expression in these cells decrease until it is no longer detectable. Such a decrease is first noted in cells that become committed to forming extraembryonic structures and finally in cells of the specific germ layers as they emerge from the primitive streak (see Chapter 5). Even after virtually all cells of the embryo have ceased to express the oct4 gene, it is still detectable in the primordial germ cells as they migrate from the region of the allantois to the genital ridges. Because of its pattern of distribution, oct-4 protein is suspected to play a regulatory role in maintenance of the undifferentiated state and in establishing and maintaining the pluripotency of the germ cells.
Parental Imprinting
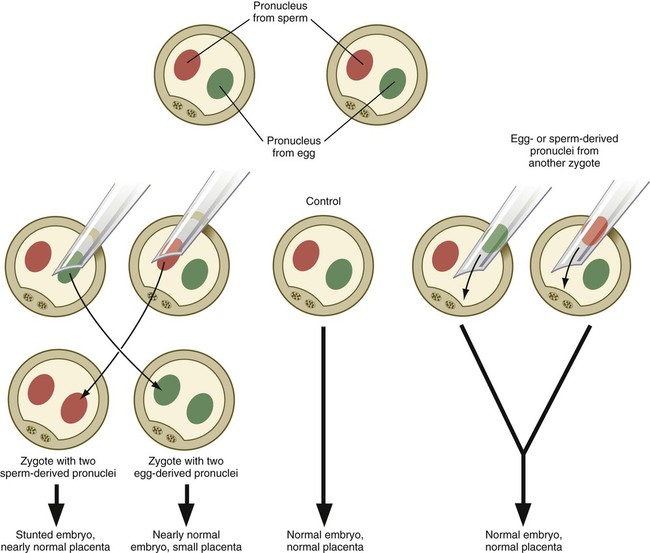
Experimentation, coupled with observations on some unusual developmental disturbances in mice and humans, has shown that the expression of certain genes derived from the egg differs from the expression of the same genes derived from the spermatozoon. Called parental imprinting, the effects are manifest in different ways. It is possible to remove a pronucleus from a newly inseminated mouse egg and replace it with a pronucleus taken from another inseminated egg at a similar stage of development (Fig. 3.8). If a male or female pronucleus is removed and replaced with a corresponding male or female pronucleus, development is normal. If a male pronucleus is removed and replaced with a female pronucleus (resulting in a zygote with two female pronuclei), however, the embryo itself develops fairly normally, but the placenta and yolk sac are poorly developed. Conversely, a zygote with two male pronuclei produces a severely stunted embryo, whereas the placenta and yolk sac are nearly normal.
Not all genes are parentally imprinted. Present estimates suggest that up to 2100 human genes are imprinted. Clinical Correlation 3.1 discusses some conditions and syndromes associated with disturbances in parental imprinting.
X-Chromosome Inactivation
Genetic studies show a complex ontogenetic history of X-chromosome inactivation (Fig. 3.9). In the female zygote, both X chromosomes are transcriptionally inactive, although not through the actions of XIST, because of the global inactivation of transcription in the early cleaving embryo. By the four-cell stage and into the morula stage, the paternally derived X chromosome becomes inactivated as the result of parental imprinting. Then, as the embryo forms the blastocyst, the paternally derived X chromosomes in the trophoblast and the hypoblast (see Fig. 5.1) remain inactivated, but within the cells of the inner cell mass both X chromosomes become active. As the cells of the inner cell mass begin to differentiate, the somatic cells undergo random permanent XIST-based X-chromosome inactivation of either the maternal or the paternal X chromosome. Within the germ cell line, activation of both X chromosomes occurs during the first meiotic division.
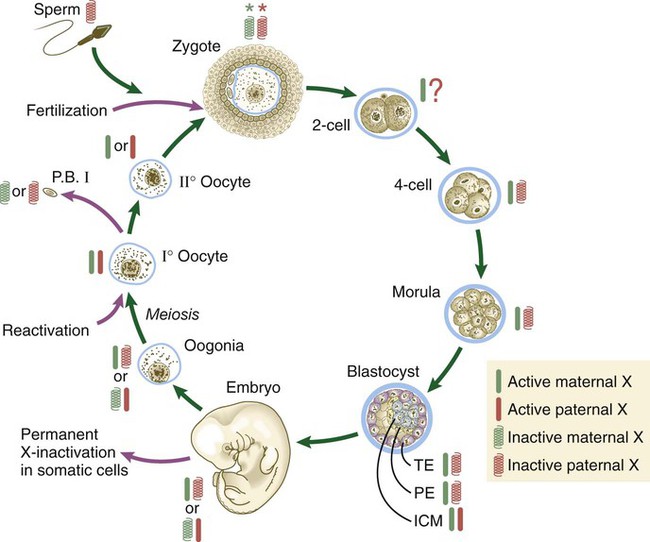
The red and green symbols refer to inactivated paternally (red) and maternally (green) derived X chromosomes. ICM, inner cell mass; P.B. I, first polar body; PE, primitive (extraembryonic) endoderm; TE, trophectoderm. (Based on Gartler SM, Riggs AD: Annu Rev Genet 17:155-190, 1983; and Thorvaldsen JL, Verona RI, Bartolomei MS: Dev Biol 298:344-353, 2006.)
Developmental Properties of Cleaving Embryos
Early mammalian embryogenesis is considered to be a highly regulative process. Regulation is the ability of an embryo or organ primordium to produce a normal structure if parts have been removed or added.* At the cellular level, this means that the fates of cells in a regulative system are not irretrievably fixed, and the cells can still respond to environmental cues. Because the assignment of blastomeres to different cell lineages is one of the principal features of mammalian development, identifying the environmental factors that are involved is important.
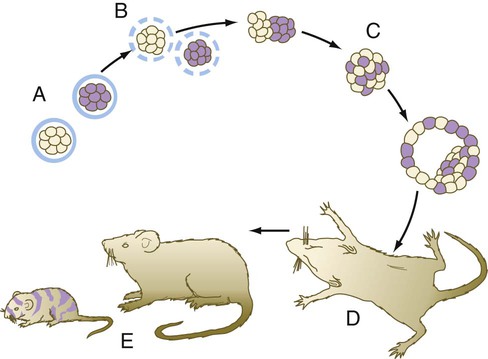
Another means of showing the regulative properties of early mammalian embryos is to dissociate mouse embryos into separate blastomeres and to combine the blastomeres of two or three embryos (Fig. 3.10). The combined blastomeres soon aggregate and reorganize to become a single large embryo, which goes on to become a normal-appearing tetraparental or hexaparental mouse. By various techniques of making chimeric embryos, it is possible to combine blastomeres to produce interspecies chimeras (e.g., a sheep-goat). It is likely that many human genetic mosaics (chimeras), most commonly recognized when some regions of the body are male and others are female, are the result of the fusion of two early fraternal twin embryos. Other possibilities for chimerism involve the exchange of cells through common vascular connections.
Experimental Manipulations of Cleaving Embryos
Much of the knowledge about the developmental properties of early mammalian embryos is the result of more recently devised techniques for experimentally manipulating them. Typically, the use of these techniques must be combined with other techniques that have been designed for in vitro fertilization, embryo culture, and embryo transfer (see Chapter 2).
Blastomere deletion and addition experiments (Fig. 3.11) have convincingly shown the regulative nature (i.e., the strong tendency for the system to be restored to wholeness) of early mammalian embryos. Such knowledge is important in understanding why the exposure of early human embryos to unfavorable environmental influences typically results in either death or a normal embryo.
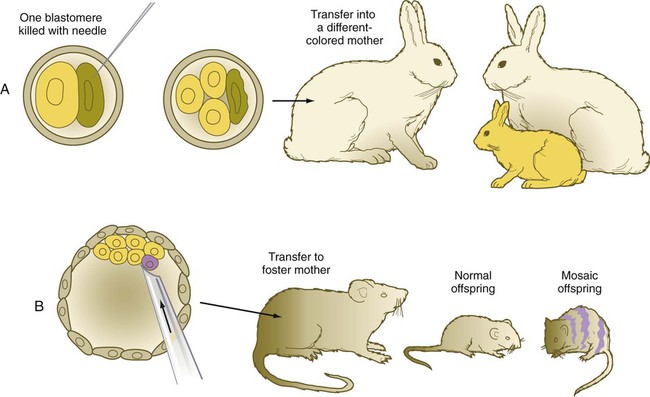
A, If one blastomere is killed with a needle, and the embryo is transferred into a different-colored mother, a normal offspring of the color of the experimentally damaged embryo is produced. B, If a blastomere of a different strain is introduced into a blastocyst, a mosaic offspring with color markings characteristic of the strain of the introduced blastomere is produced.
One of the most powerful experimental techniques has been the injection of genetically or artificially labeled cells into the blastocyst cavity of a host embryo (see Fig. 3.11B). This technique has been used to show that the added cells become normally integrated into the body of the host embryo, thus providing additional evidence for embryonic regulation. An equally powerful use of this technique has been in the study of cell lineages in the early embryo. By identifying the progeny of the injected marked cells, investigators have been able to determine the developmental potency of the donor cells.
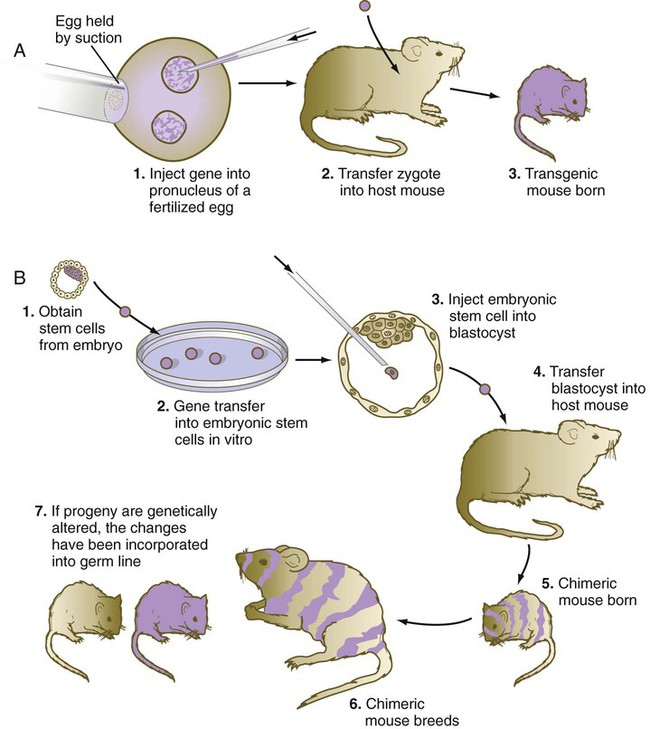
A technique that provides great insight into the genetic control mechanisms of mammalian development is the production of transgenic embryos. Transgenic embryos (commonly mice) are produced by directly injecting foreign DNA into the pronuclei of zygotes (Fig. 3.12A). The DNA, usually recombinant DNA for a specific gene, can be fused with a different regulatory element that can be controlled by the investigator.
Transgenic mice can be created by injecting the rat growth hormone gene coupled with a metallothionein promoter region (MT-I) into the pronuclei of mouse zygotes. The injected zygotes are transplanted into the uteri of foster mothers, which give birth to normal-looking transgenic mice. Later in life, when these transgenic mice are fed a diet rich in zinc, which stimulates the MT-I promoter region, the rat growth hormone gene is activated and causes the liver to produce large amounts of the polypeptide growth hormone. The function of the transplanted gene is obvious; under the influence of the rat growth hormone that they are producing, the transgenic mice grow to a much larger size than their normal littermates (Fig. 3.13).

The one on the left (normal mouse) weighs 21.2 g. The one on the right (a transgenic littermate of the normal mouse) carries a rat gene coding for growth hormone. It weighs 41.2 g. (From Palmiter RD and others: Nature 300:611-615, 1982.)
Some types of twinning represent a natural experiment that shows the highly regulative nature of early human embryos, as described in Clinical Correlation 3.2.
Stem Cells and Cloning
A major development in biomedical research at the turn of the 21st century was the realization that certain cells (stem cells) in both human embryos and adults have the capacity to develop into a variety of cell and tissue types in response to specific environments. In embryos, stem cells can be derived from the inner cell mass (embryonic stem cells [ES cells]) or primordial germ cells (embryonic germ cells). In adults, stem cells have been isolated from tissues as diverse as bone marrow, skeletal muscle, brain tissue, and fat. Regardless of their origin, stem cells are maintained and propagated in an undifferentiated state in culture. Characteristically, stem cells express oct4, Sox2, and Nanog (see p. 42), which are involved in maintaining the undifferentiated state.
Genetic engineering of specific genes is possible in ES cells. When such genetically manipulated cells are introduced into blastocysts, they can become incorporated into the host embryo (see Fig. 3.12B). If the progeny of a genetically engineered ES cell become incorporated into the germline, the genetic trait can be passed to succeeding generations.
Embryo Transport and Implantation
Transport Mechanisms by the Uterine Tube
The entire period of early cleavage occurs while the embryo is being transported from the place of fertilization to its implantation site in the uterus (see Fig. 2.2). It is increasingly apparent that the early embryo and the female reproductive tract influence one another during this period of transport. One such influence is early pregnancy factor, a molecule of the heat shock protein family and homologous to chaperonin 10, an intramitochondrial protein. Early pregnancy factor, which is detectable in maternal blood within 36 to 48 hours after fertilization, is an immunosuppressant and is postulated to provide immunological protection to the embryo. Although this factor is produced by the embryo, its presence in serum seems to result from its synthesis and secretion by the ovary. Because the assay for this protein is cumbersome, it has not found wide use in pregnancy testing.
Zona Pellucida
During the entire period from ovulation until entry into the uterine cavity, the ovum and the embryo are surrounded by the zona pellucida. During this time, the composition of the zona changes, through contributions from the blastomeres and the maternal reproductive tissues. These changes facilitate the transport and differentiation of the embryo. After the embryo reaches the uterine cavity, it begins to shed the zona pellucida in preparation for implantation. This is accomplished by a process called blastocyst hatching. A small region of the zona pellucida, usually directly over the inner cell mass in the primate, dissolves, and the blastocyst emerges from the hole. In rodents, blastocyst hatching is accomplished through the action of cysteine protease enzymes that are released from long microvillous extensions (trophectodermal projections) protruding from the surfaces of the trophoblastic cells. Over a narrow time window (4 hours in rodents), the zona pellucida in this area is digested, and the embryo begins to protrude. In the uterus, the trophectodermal projections then make contact with the endometrial epithelial cells as the process of implantation begins. Enzymatic activity around the entire trophoblast soon begins to dissolve the rest of the zona pellucida. Only a few specimens of human embryos have been taken in vivo from the period just preceding implantation, but in vitro studies on human embryos suggest a similar mechanism, which probably occurs 1 to 2 days before implantation (see Fig. 3.3C). Box 3.1 summarizes the functions of the zona pellucida.
Implantation into the Uterine Lining
Successful implantation requires a high degree of preparation and coordination by the embryo and the endometrium (Table 3.1). The complex hormonal preparations of the endometrium that began at the close of the previous menstrual period all are aimed at providing a suitable cellular and nutritional environment for the embryo. Even before actual contact is made between the embryo and endometrium, the uterine epithelium secretes into the uterine fluid certain cytokines and chemokines that facilitate the implantation process. At the same time, cytokine receptors appear on the surface of the trophoblast. Dissolution of the zona pellucida signals the readiness of the embryo to begin implantation.
Table 3.1
| Age (Days) | Developmental Event in Embryos |
| 5 | Maturation of blastocyst |
| 5 | Loss of zona pellucida from blastocyst |
| 6? | Attachment of blastocyst to uterine epithelium |
| 6-7 | Epithelial penetration |
 |
Trophoblastic plate formation and invasion of uterine stroma by blastocyst |
| 9-11 | Lacuna formation along with erosion of spiral arteries in endometrium |
| 12-13 | Primary villus formation |
| 13-15 | Secondary placental villi, secondary yolk sac formation |
| 16-18 | Branching and anchoring villus formation |
| 18-22 | Tertiary villus formation |
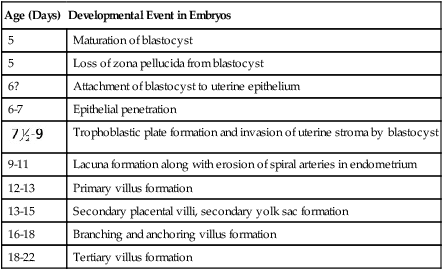
Modified from Enders AC: Implantation, embryology. In Encyclopedia of human biology, vol 4, New York, 1991, Academic Press.
The next stage of implantation is penetration of the uterine epithelium. In primates, the cellular trophoblast undergoes a further stage in its differentiation just before it contacts the endometrium. In the area around the inner cell mass, cells derived from the cellular trophoblast (cytotrophoblast) fuse to form a multinucleated syncytiotrophoblast. Although only a small area of syncytiotrophoblast is evident at the start of implantation, this structure (sometimes called the syntrophoblast) soon surrounds the entire embryo. Small projections of syncytiotrophoblast insert themselves between uterine epithelial cells. They spread along the epithelial surface of the basal lamina that underlies the endometrial epithelium to form a flattened trophoblastic plate. Within a day or so, syncytiotrophoblastic projections from the small trophoblastic plate begin to penetrate the basal lamina. The early syncytiotrophoblast is a highly invasive tissue, and it quickly expands and erodes its way into the endometrial stroma (Fig. 3.18A and B). Although the invasion of the syncytiotrophoblast into the endometrium is obviously enzymatically mediated, the biochemical basis in humans is not well understood. By 10 to 12 days after fertilization, the embryo is completely embedded in the endometrium. The site of initial penetration is first marked by a bare area or a noncellular plug and is later sealed by migrating uterine epithelial cells (Fig. 3.18C and D).
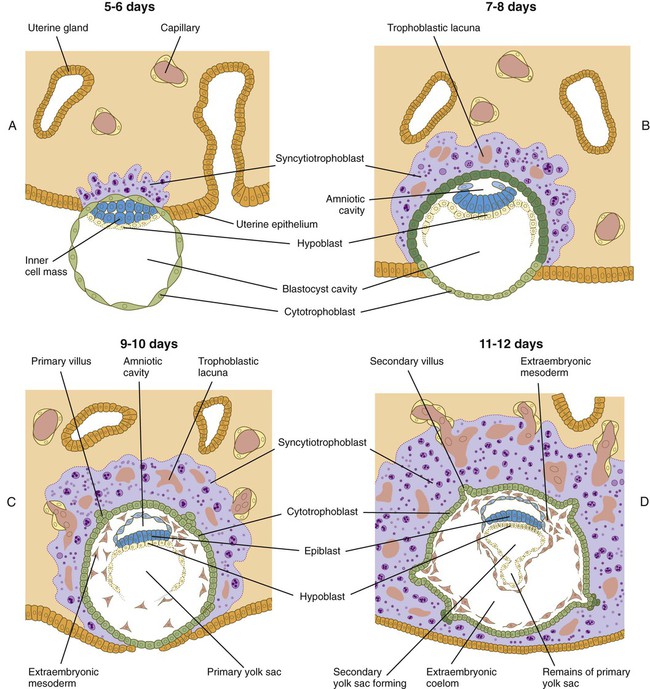
A, The syncytiotrophoblast is just beginning to invade the endometrial stroma. B, Most of the embryo is embedded in the endometrium; there is early formation of the trophoblastic lacunae. The amniotic cavity and yolk sac are beginning to form. C, Implantation is almost complete, primary villi are forming, and the extraembryonic mesoderm is appearing. D, Implantation is complete; secondary villi are forming.
As early implantation continues, projections from the invading syncytiotrophoblast envelop portions of the maternal endometrial blood vessels. They erode into the vessel walls, and maternal blood begins to fill the isolated lacunae that have been forming in the trophoblast (see Fig. 3.18C and D). Trophoblastic processes enter the blood vessels and even share junctional complexes with the endothelial cells. By the time blood-filled lacunae have formed, the trophoblast changes character, and it is not as invasive as it was during the first few days of implantation. Leakage of blood from the uterus at this stage can produce “spotting,” which is sometimes misinterpreted to be an abnormal menstrual period.
While the embryo burrows into the endometrium, and some cytotrophoblastic cells fuse into syncytiotrophoblast, the fibroblastlike stromal cells of the edematous endometrium swell, with the accumulation of glycogen and lipid droplets (see Fig. 7.6). These cells, called decidual cells, are tightly adherent and form a massive cellular matrix that first surrounds the implanting embryo and later occupies most of the endometrium. Concurrent with the decidual reaction, as this transformation is called, the leukocytes that have infiltrated the endometrial stroma during the late progestational phase of the endometrial cycle secrete interleukin-2, which prevents maternal recognition of the embryo as a foreign body during the early stages of implantation. An embryo is antigenically different from the mother and consequently should be rejected by a cellular immune reaction similar to the type that rejects an incompatible heart or kidney transplant. A primary function of the decidual reaction apparently is to provide an immunologically privileged site to protect the developing embryo from being rejected, but a real understanding of how this is accomplished has resisted years of intensive research.
Frequently, a blastocyst fails to attach to the endometrium, and implantation does not occur. Failure of implantation is a particularly vexing problem in in vitro fertilization and embryo transfer procedures, for which the success rate of implantation of transferred embryos remains at about 25% to 30% (see Clinical Correlation 2.1).
Summary
 Early human cleavage is slow, with roughly a single cleavage division occurring per day for the first 3 to 4 days. The cleaving embryo passes through the morula stage (16 cells) and enters a stage of compaction. By day 4, a fluid-filled blastocoele forms within the embryo, and the embryo becomes a blastocyst with an inner cell mass surrounded by trophoblast.
Early human cleavage is slow, with roughly a single cleavage division occurring per day for the first 3 to 4 days. The cleaving embryo passes through the morula stage (16 cells) and enters a stage of compaction. By day 4, a fluid-filled blastocoele forms within the embryo, and the embryo becomes a blastocyst with an inner cell mass surrounded by trophoblast.
 The zygote relies on maternal mRNAs, but by the two-cell stage, the embryonic genome becomes activated. The oct4, Sox2, and Nanog genes are important in very early development, and their expression is associated with the undifferentiated state of cells.
The zygote relies on maternal mRNAs, but by the two-cell stage, the embryonic genome becomes activated. The oct4, Sox2, and Nanog genes are important in very early development, and their expression is associated with the undifferentiated state of cells.
 Through parental imprinting, specific homologous chromosomes derived from the mother and father exert different effects on embryonic development. In female embryos, one X chromosome per cell becomes inactivated through the action of the XIST gene, thus forming the sex chromatin body. The early embryo has distinct patterns of X-chromosomal inactivation.
Through parental imprinting, specific homologous chromosomes derived from the mother and father exert different effects on embryonic development. In female embryos, one X chromosome per cell becomes inactivated through the action of the XIST gene, thus forming the sex chromatin body. The early embryo has distinct patterns of X-chromosomal inactivation.
 The early mammalian embryo is highly regulative. It can compensate for the loss or addition of cells to the inner cell mass and still form a normal embryo. The decision to form trophoblast versus inner cell mass relates to division patterns of polarized cells, starting at the eight-cell stage. According to the inside-outside hypothesis, the position of a blastomere determines its developmental fate (i.e., whether it becomes part of the inner cell mass or the trophoblast).
The early mammalian embryo is highly regulative. It can compensate for the loss or addition of cells to the inner cell mass and still form a normal embryo. The decision to form trophoblast versus inner cell mass relates to division patterns of polarized cells, starting at the eight-cell stage. According to the inside-outside hypothesis, the position of a blastomere determines its developmental fate (i.e., whether it becomes part of the inner cell mass or the trophoblast).
 Transgenic embryos are produced by injecting ribosomal DNA (rDNA) into the pronuclei of zygotes. Such embryos are used to study the effects of specific genes on development. Other techniques involve knocking out genes or interfering with the further processing of gene products.
Transgenic embryos are produced by injecting ribosomal DNA (rDNA) into the pronuclei of zygotes. Such embryos are used to study the effects of specific genes on development. Other techniques involve knocking out genes or interfering with the further processing of gene products.
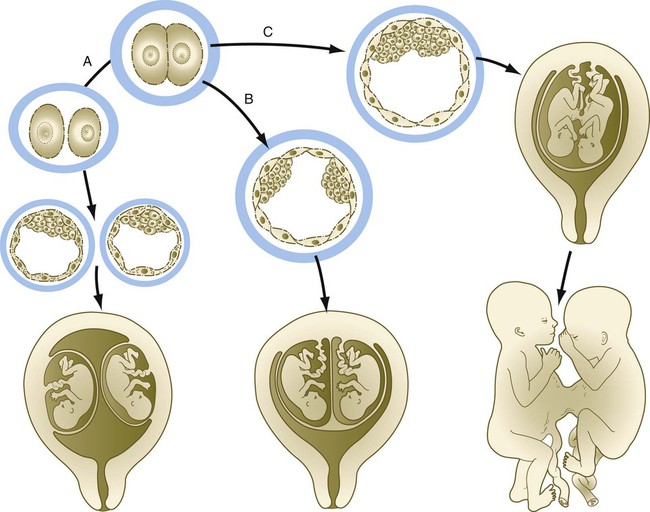
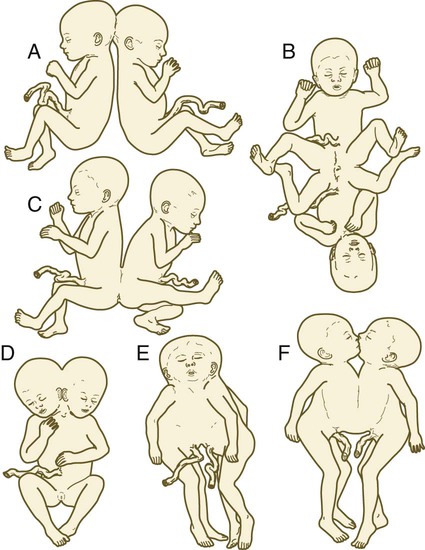
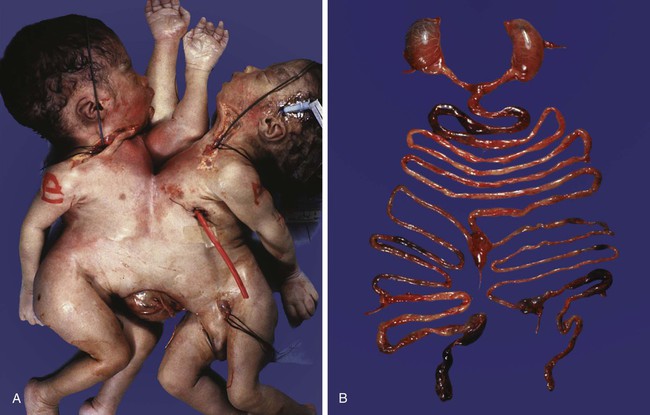
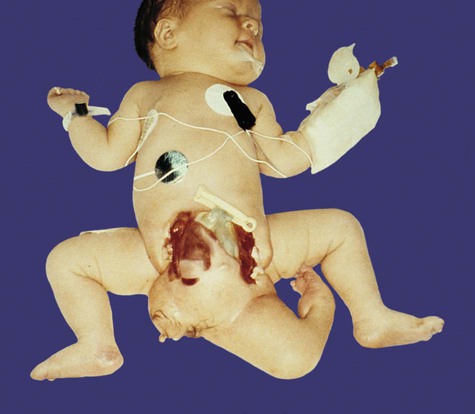
 Monozygotic twinning, usually caused by the complete separation of the inner cell mass, is possible because of the regulative properties of the early embryo. Incomplete splitting of the inner cell mass can lead to the formation of conjoined twins.
Monozygotic twinning, usually caused by the complete separation of the inner cell mass, is possible because of the regulative properties of the early embryo. Incomplete splitting of the inner cell mass can lead to the formation of conjoined twins.
 After fertilization, the embryo spends several days in the uterine tube before entering the uterus. During this time, it is still surrounded by the zona pellucida, which prevents premature implantation.
After fertilization, the embryo spends several days in the uterine tube before entering the uterus. During this time, it is still surrounded by the zona pellucida, which prevents premature implantation.
 Implantation of the embryo into the uterine lining involves several stages: apposition of the expanded (hatched) blastocyst to the endometrial epithelium, penetration of the uterine epithelium, invasion into the tissues underlying the epithelium, and erosion of the maternal vascular supply. Connective tissue cells of the endometrium undergo the decidual reaction in response to the presence of the implanting embryo. Implantation is accomplished through the invasive activities of the syncytiotrophoblast, which is derived from the cytotrophoblast.
Implantation of the embryo into the uterine lining involves several stages: apposition of the expanded (hatched) blastocyst to the endometrial epithelium, penetration of the uterine epithelium, invasion into the tissues underlying the epithelium, and erosion of the maternal vascular supply. Connective tissue cells of the endometrium undergo the decidual reaction in response to the presence of the implanting embryo. Implantation is accomplished through the invasive activities of the syncytiotrophoblast, which is derived from the cytotrophoblast.
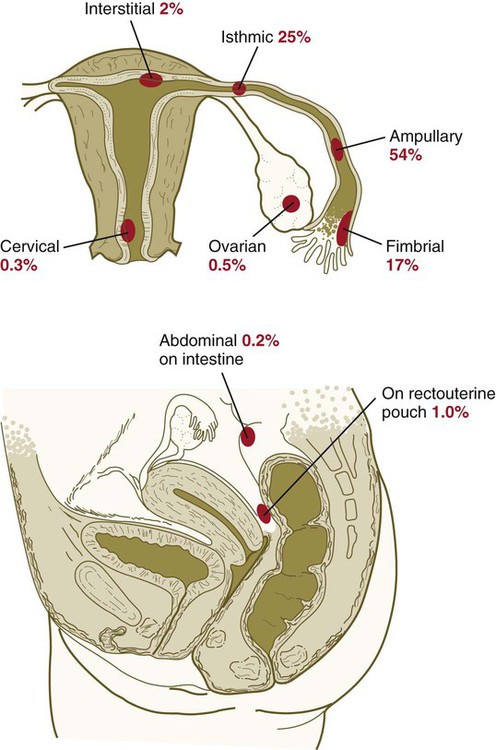
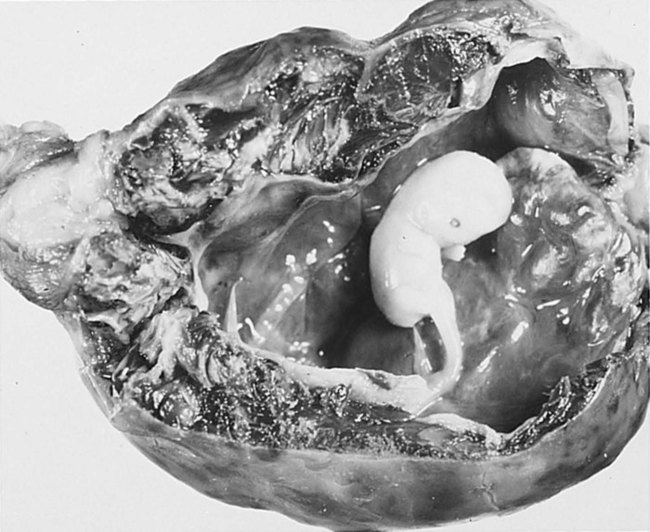
 Implantation of the embryo into a site other than the upper uterine cavity results in an ectopic pregnancy (Clinical Correlation 3.3). Ectopic pregnancy is most often encountered in the uterine tube.
Implantation of the embryo into a site other than the upper uterine cavity results in an ectopic pregnancy (Clinical Correlation 3.3). Ectopic pregnancy is most often encountered in the uterine tube.
 High percentages of fertilized eggs and early embryos do not develop and are spontaneously aborted. Many of these embryos contain major chromosomal abnormalities.
High percentages of fertilized eggs and early embryos do not develop and are spontaneously aborted. Many of these embryos contain major chromosomal abnormalities.
*In contrast to regulative development is mosaic development, which is characterized by the inability to compensate for defects or to integrate extra cells into a unified whole. In a mosaic system, the fates of cells are rigidly determined, and removal of cells results in an embryo or a structure missing the components that the removed cells were destined to form. Most regulative systems have an increasing tendency to exhibit mosaic properties as development progresses.

 months, the implanted embryo and its associated trophoblastic derivatives have grown to the point where the stretching of the tube causes acute abdominal pain. If untreated, a tubal pregnancy typically ends with rupture of the tube and hemorrhage, often severe enough to be life-threatening to the mother.
months, the implanted embryo and its associated trophoblastic derivatives have grown to the point where the stretching of the tube causes acute abdominal pain. If untreated, a tubal pregnancy typically ends with rupture of the tube and hemorrhage, often severe enough to be life-threatening to the mother.