Acquired Immune Deficiency Syndrome
Learning Objectives
• Recognize the origin of human immunodeficiency virus 1 (HIV-1) and HIV-2
• Identify the HIV-1 clade predominant in most of the world
• Describe the structure of HIV-1
• Compare and contrast HIV-1 binding to lymphocytes and monocytes
• Explain the mechanisms involved in the fusion of HIV and the host cell membrane
• Explain the reverse transcription of HIV
• Understand the two-step process used to integrate viral deoxyribonucleic acid (DNA)
• Identify the process by which HIV exits host cells
• Discuss the roles of follicular dendritic cells in the pathogenesis of HIV
• List the possible mechanisms in the killing of CD4 cells
• Identify the possible mechanisms involved in CD8 dysfunction
• Identify HIV target cells in the brain
• Differentiate between the three states of HIV infection
• Compare and contrast HIV infection and acquired immune deficiency syndrome (AIDS)
• List the AIDS-defining medical conditions
• Understand the rationale for using highly active anti-retroviral therapy (HAART)
• Explain the mechanism of action of fusion inhibitors
• Compare and contrast nucleoside and non-nucleoside reverse transcriptase inhibitors
Key Terms
Acquired immune deficiency syndrome (AIDS)
Capsid
Chemokine
Clade
Env
Follicular dendritic cells
Gag
Gp120/gp41
Iccosomes
Integrase
Matrix
Nef
Pol
Protease
Rev
Reverse transcriptase
Simian immunodeficiency virus (SIV)
Syncytia
Tat
Vif
Vpr
Vpu
Introduction
Human immunodeficiency virus (HIV) is closely related to simian immunodeficiency viruses (SIVs), which are found in equatorial Africa. SIVs are found in nine classes of monkeys and chimpanzees. Genetic sequencing strongly suggests that an SIV endemic to a chimpanzee subspecies (Pan troglodytes troglodytes) first crossed the species barrier in 1930. This virus is the ancestral HIV-1. In Gabon and Cameroon, the Sooty Mangabey monkey population is infected with a different SIV. Sometime between 1955 and 1972, the monkey SIV crossed the species barrier in several direct jumps to humans. The genome of this virus differs from HIV-1 by 30% to 40% and is designated HIV-2.
Subsequent studies linked HIV with the destruction of the immune system and acquired immune deficiency syndrome (AIDS). The syndrome is characterized by infections with unusual opportunistic pathogens (e.g., Pneumocystis jiroveci) and the development of aggressive sarcomas.
HIV-1 causes over 90% of human infections in the world. In the United States, 1.1 million individuals are infected with HIV-1, and 56,000 new cases are reported each year. HIV-2 is responsible for at least six epidemics in Africa. The World Health Organization (WHO) estimates that 4.3 million individuals are newly infected with HIV each year. Over 95% of these infections are occurring in the developing countries and Sub-Saharan Africa.
Human Immunodeficiency Virus Groups and Clades
According to the frequency of infections and the genetic diversity, HIV-1 is subdivided into groups and clades. HIV-1 is initially grouped into M (major), O (outlying), and N (new) groups. The HIV-1 M group has spread throughout the world. Groups N and O are restricted to West Equatorial Africa. Mutations within the M group, environmental pressure, and therapeutic selection have given rise to HIV clades. By definition, a clade is a virus subgroup that arose from a single common ancestor. The HIV-1 M group has nine major genetically diverse clades (A, B, C, D, F, G, H, J, and K). In the Western world, the B clade is the predominant virus. Transmission of HIV-1 B is rapid and associated with sex between men, intravenous drug use, and transmission of the virus from mothers to babies. HIV-1 A and C clades are found in southern and eastern Africa. Recombinant A-G and A-E clades also are emerging as infectious agents in Africa and Asia. In areas where subtypes C or A-E are common, heterosexual transmission is more frequent. Heterosexual transmission occurs because these clades replicate more efficiently in the Langerhans and dendritic cells of the vagina, and the virus is then shed into vaginal fluids. In the United States, the emerging trend in heterosexual HIV transmission has occurred because of the introduction of the A-E or C clade into North America.
Structure of the Human Immunodeficiency Virus
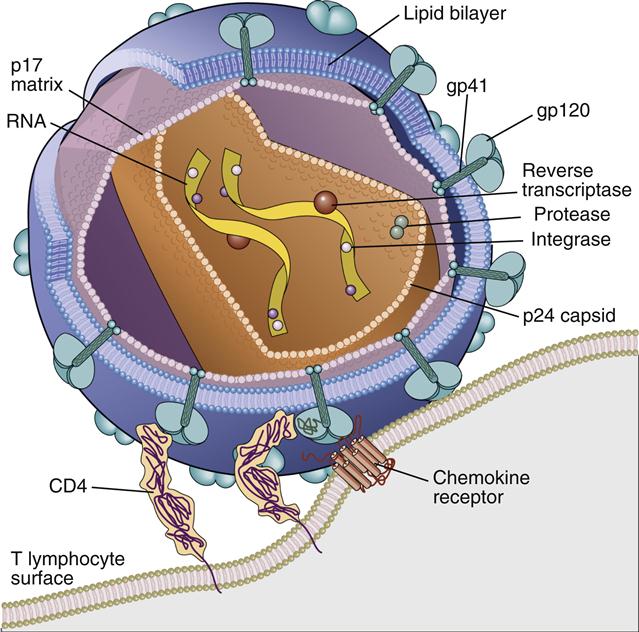
HIV virus comprises an inner core, a capsid, a matrix shell, and a host cell–derived outer membrane. The inner core contains two single strands of ribonucleic acid (RNA). One RNA strand, called the plus (+) or coding strand, contains viral genes that are transcribed. A second strand, the negative (–) strand, is complementary to the plus strand. The RNA and several proteins necessary for viral replication are encased in a cone-shaped capsid (Figure 26-1).
Additional protection for the capsid core is provided by an icosahedral shell or matrix. The shell anchors a lipid membrane to the outer surface of the virus. Protruding from the HIV membrane is the gp120 viral protein, which is anchored by a transmembrane gp41 protein. Three molecules of the gp120–gp41 aggregate to form a trimolecular structure necessary for docking to HIV target cells.
Human Immunodeficiency Virus Genes
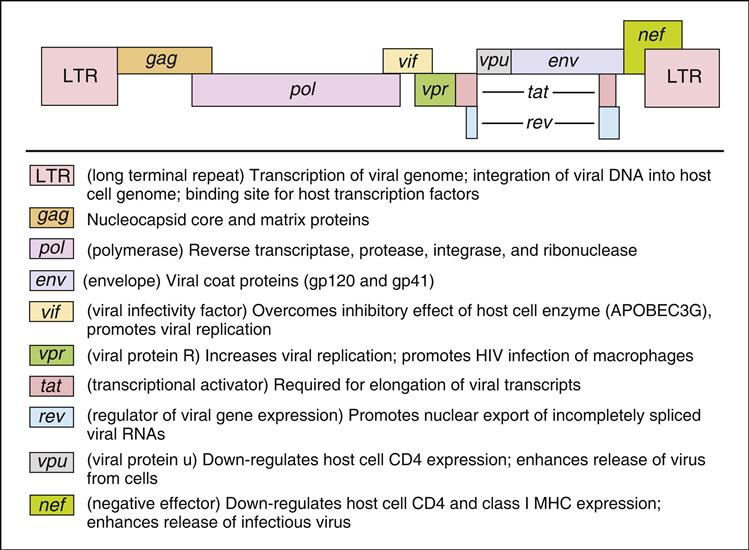
The HIV-1 genome codes for all the gene products necessary for viral replication. The genome is divided into structural genes (gag, pol, and env), regulatory genes (tat and rev) and accessory genes (vpr, nef, vpu, and vif). The organization of the HIV genome and the function of individual genes are shown in Figure 26-2.
Life Cycle of the Human Immunodeficiency Virus
The life cycle of HIV is divided into several stages, which include attachment, membrane fusion, uncoating, reverse transcription of viral RNA into DNA, and integration into the host cell genome. When HIV is integrated into the host cell DNA, it is called a latent provirus. Latency is characterized by the lack of viral gene expression. At some point in time, intracellular tat levels increase, and viral RNA is transcribed. After synthesis of multiple copies, the double-stranded RNA is packaged and exported from the cell.
Attachment
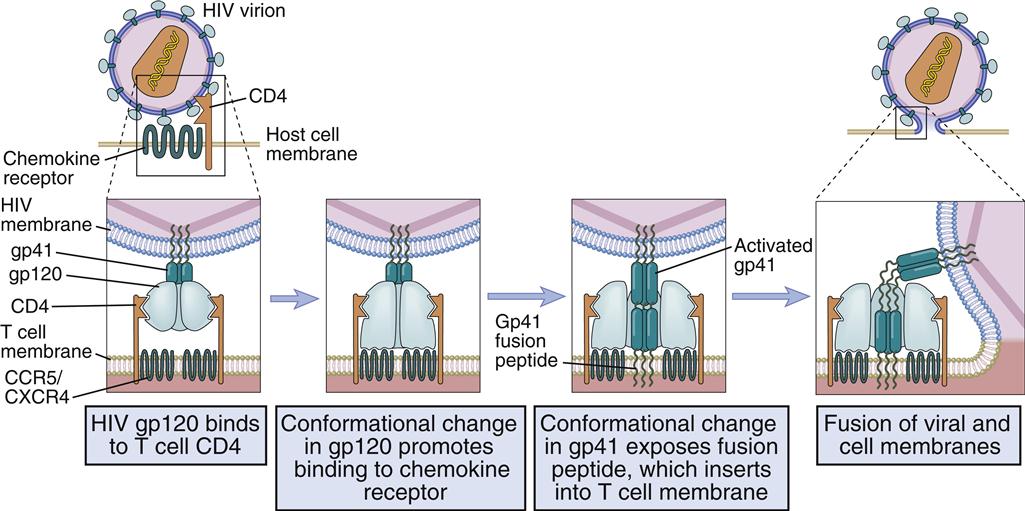
Different receptors are used to attach the virus gp120–gp41 to lymphocytes or monocytes. Attachment to T cells requires both CD4 and the chemokine receptor (CXCR4). Monocyte attachment points for HIV gp120–gp41 are a cell surface heparin molecule and a macrophage-specific chemokine (CCR5) receptor. Interaction with two receptors is necessary for the fusion of HIV with the host cell membrane (Figure 26-3).
Membrane Fusion
After interaction with gp120, the membrane attachment component (gp41) disassociates itself and undergoes a conformational change to become a long, cylindrical fusion protein, which bridges the divide between the virus and the cell membrane. After inserting itself into the host cell membrane, the gp41 fusion protein refolds itself using two specialized molecular H-1 and H-2 sequences called heptadrepeats. The H-1 and H-2 regions fold and form a six-helix fusion protein bundle, which brings the virus and host cell membrane into close contact (see Figure 26-3). Several six-helix fusion proteins form a narrow fusion pore in the membrane, which expands to allow viral entry into the cell.
After uncoating of the virus in the host cell cytoplasm, the nef gene is translated; it accelerates the endocytosis of the CD4 virus complex and downregulates the expression of class I molecules, which decreases the risk of the CD8-mediated destruction of infected cells.
Reverse Transcription (RT)
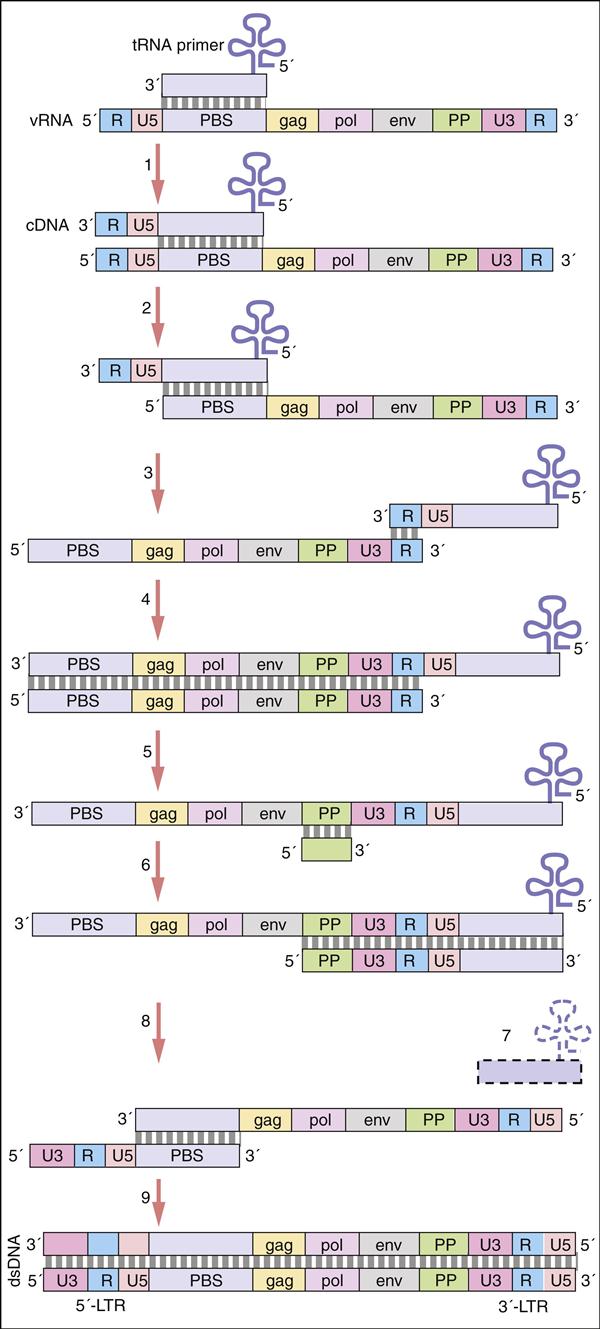
Pol gene products are critical for the synthesis and integration of viral DNA into the host genome and the generation of capsid proteins. Pol gene products include the HIV-1 reverse transcriptase, an integrase, and the late-phase protease. The pol gene proteins complex with RNase and vpr in the cell membrane to form the viral reverse transcription complex. The complex docks with the actin microfilaments in the cell and begins the RT process. After degradation of the negative RNA strand, the plus, or coding, strand is prepared for transcription. The plus RNA stand consists of the following: (1) R regions, which are direct repeats at both ends of the genome, (2) a U5 noncoding region, which is initially transcribed and forms the 3′ end of new DNA; (3) a leucine transfer RNA (tRNA) primer binding site (PBS), which is used to initiate reverse transcription; (4) a leader sequence, which is located downstream of the transcription start site and upstream of the structural genes; (5) structural, regulatory, and accessory genes; (6) a polypurine tract (PPT), which is responsible for initiating plus strand synthesis; and (7) a U3 noncoding region, which contains the promoter elements of the single-stranded DNA and a noncoding 5′ end of the genome (Figure 26-4).
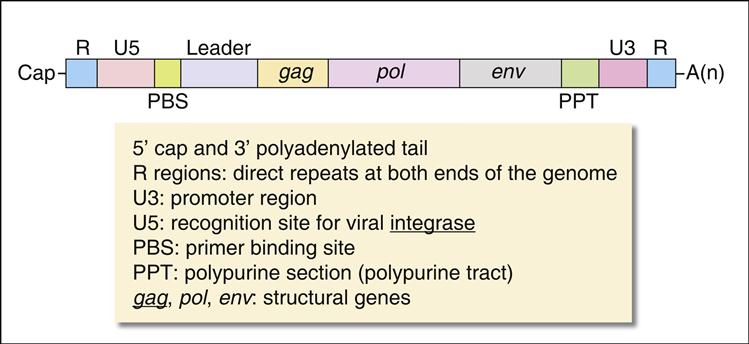

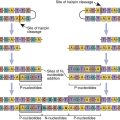
Reverse transcription begins from the 5′ terminus and proceeds to the 3′ terminus. In the initial stages of reverse transcription, the leucine tRNA binds to the PBS near the 5′ end of the viral RNA. Complementary DNA binds to the U5 and R sites and the tRNA, creating a complex consisting of a small fragment of DNA linked to the viral RNA. Subsequently, the 5′ end of the viral RNA in the complex is degraded by the enzyme RNase to create a DNA primer. The primer consists of the tRNA and DNA complementary U5 and R regions (see Figure 26-4). Since the reverse transcriptase synthesizes DNA from the 5′ end to the 3′ prime end, the primer must jump to the other end of the viral DNA, where it binds to the R region. The reverse transcriptase is activated and elongates a plus, or sense, DNA strand.
Subsequently, most of the RNA is degraded by the RNase leaving only the PPT. Using a DNA polymerase and the polypurine primer, a small piece of negative, or antisense, DNA is synthesized in a 5′ end to 3′ end direction. The DNA fragment codes for the U3, R, and U5 segments. This fragment jumps to the 5′ of the plus strand DNA and binds to the PBS tract. A DNA polymerase extends the plus strand and assists in the synthesis of double-stranded DNA (Figure 26-5).
Preintegration Complex
Following the synthesis of double-stranded viral DNA, a preintegration complex (PIC) forms for the transport of DNA to the nucleus. The PIC consists of DNA, viral protein R (vpr), and the integrase. The PIC switches attachment from actin microfilaments to microtubules, which actively transports HIV DNA to the nucleus. To increase the efficiency of viral DNA into the host chromosome, vpr arrests proliferating cells in the G2 phase of cell division.
Integrase
Double-stranded viral DNA is integrated into the host genome by a two-step process that occurs in the cytoplasm and the nucleus. In the cytoplasm, the integrase binds to the blunt end of the DNA strands and removes two nucleotides from the 3′-OH end of each viral DNA strand in a process called 3′-end processing. Processed viral DNA is transferred to the nucleus and the exposed 3′-OH ends ligated to the cellular DNA (3′ end joining). Host cell DNA repair enzymes fill in any gaps in DNA and remove unpaired nucleotides. Viral DNA is synthesized and inserted within 6 hours of infection.
Viral Assembly and Release
Tat gene translation is essential for viral replication. Tat promotes full length synthesis of viral RNA by stabilizing the RNA polymerase and preventing the termination of protein synthesis. Interactions between Tat and the polymerase increase the production of viral RNA and messenger RNA (mRNA) approximately 100-fold.
Rev binds to mRNA, which contains a Rev response element (RRE). This promotes stabilization and export of mRNA from the nucleus to the cytoplasm of unspliced and single-spliced RNA. Under normal circumstances, only unspliced RNA is exported. The ability to transport both types of genetic material is necessary for viral replication. Early in the replication process, unspliced RNA that codes for RNase, nef, tat, and rev exits the nucleus. As rev levels increase, single-spliced RNA coding for matrix and capsid proteins are exported from the nucleus. The vif protein protects RNA from degradation by cytoplasmic enzymes.
Translation of the gag and env genes is necessary for the formation of HIV capsid and matrix proteins. Gag gene transcription generates a large p55 precursor protein, which is cleaved by the protease into small p24 (capsid) and p17 (nucleocapsid) proteins (see Figure 26-1). The env gene complex codes for gp120–gp41, regulatory, and accessory proteins. With the assistance of specialized proteins, the assembled virus passes through the host cell lipid bi-layer membrane and exits the cell by a process called budding. As a consequence, the virus is coated with a cholesterol-rich membrane. Since the budding process does not kill the host cell, viral synthesis is continual.
Human Immunodeficiency Virus and the Immune System
Follicular Dendritic Cells and Human Immunodeficiency Virus
During the initial phases of infection, HIV is concentrated in lymph node germinal centers. Follicular dendritic cells (FDCs) use dendritic extensions called iccosomes to trap viable viruses. Some CD4 cells are infected by contact with the iccosomes. Within 6 weeks, FDCs express FasR molecules on the cell surface and are killed by CD8 cells.
FDC destruction has far-reaching effects on T cell development. Interleukin 12 (IL-12) produced by dendritic cells is necessary for the differentiation of CD4Th1 cells and maintenance of CD4 memory cells. As a consequence of FDC destruction and a lack of IL-12, CD4Th1 inflammatory responses cannot be implemented, thus increasing the risk of infection with intracellular pathogens such as mycobacteria, Histoplasma, and Pneumocystis. In addition, the dysfunctional memory cells cannot facilitate isotypic switching required for an anamnestic response to extracellular pathogens such as Staphylococcus aureus, Haemophilus influenzae, and Streptococcus pneumoniae.
Destruction of CD4 Cells
The mechanisms used by HIV to destroy CD4 cells are unclear. CD4 cells may be killed in an innocent bystander reaction. HIV synthesizes a truncated gp120, which is released into the cytoplasm. Some noninfected CD4 cells bind the soluble gp120 and are killed by cytotoxic T cells. Infected CD4 cells also form syncytia with monocytes or other T cells and undergo rapid apoptosis. Evidence also indicates that HIV acts as a superantigen that stimulates a massive polyclonal proliferation of CD4 cells. Proliferation and activation are followed by HIV infection and apoptosis of large numbers of CD4 cells. The destruction of CD4 cells results in an inversion in the normal C4–CD8 ratio in peripheral blood.
The ultimate collapse of the CD4 population is caused by stem cell exhaustion. Approximately 2.0 × 109 CD4 cells must be produced each day to match the number of dying cells. Over time, the number of stem cells able to produce CD4 cells is exhausted and the number of CD4 cells is permanently reduced.
Loss of CD8 Cell Function
Cytotoxic CD8 cells are important in containing HIV infections; individuals with normal CD8 numbers have longer survival rates. In children and adults, a loss of CD8 function correlates with increased viremia, spread of the virus within the host, and the onset of clinical symptoms. Decreased CD8 function, which can be restored by IL-2, is associated with reduced expression of T cell receptors and CD28.
Human Immunodeficiency Virus and Neurologic Effects
In 10% of patients with HIV, the virus crosses the blood–brain barrier and infects macrophages, microglial cells, and astrocytes. These cells produce proinflammatory cytokines, neurotoxins, and reactive oxygen species. Additional neurotoxins are produced by monocytes or macrophages recruited into the brain from peripheral blood. Toxin damage to neuronal cells is reflected in cognitive dysfunction and symptoms associated with the AIDS–dementia complex.
Human Immunodeficiency Virus Infection and Acquired Immune Deficiency Syndrome
In 2008, the Centers for Disease Control and Prevention (CDC) revised its acquired immune deficiency syndrome (AIDS) classification system for adults and adolescents into a single case definition for HIV infection. The case definition is based on laboratory data and the presence of an AIDS-defining condition (Box 26-1).
Presumptive laboratory evidence of HIV infection includes a positive result in an HIV antibody screening test that is confirmed by an alternative HIV test (Western blot or immunofluorescence). The presence of HIV nucleic acids, p24 capsid antigen, or isolation of HIV from blood or bodily fluids is also considered evidence of HIV infection. If the patient meets the laboratory criteria for HIV infection and one of the three stages of HIV infection (stages 1–3), HIV infection is considered confirmed.
Treatment of Human Immunodeficiency Virus Infections
Highly active anti-retroviral therapy (HAART) is a treatment strategy that combines drugs that inhibit viral replication at different steps. These drugs are classified into five categories: (1) attachment and fusion inhibitors, (2) reverse transcriptase inhibitors (nucleoside and non-nucleoside inhibitors), (3) protease inhibitors, (4) integrase inhibitors, and (5) combination drugs. At the present time, over 30 drugs have been approved by the U.S. Food and Drug Administration (FDA) for the treatment of HIV infection.
Attachment and Fusion Inhibitors
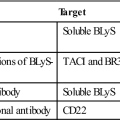
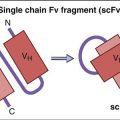
Attachment and fusion inhibitors prevent the initial interactions between the virus and target cell membranes. At this point in time, two fusion inhibitors are licensed in the United States: (1) Maraviroc prevents the binding of gp120 to the CCR5 co-receptor for HIV. It is most effective when used to treat individuals infected with macrophage-tropic HIV strains and does not inhibit HIV attachment of CXCR4 lymphocyte-tropic HIV-1 strains. (2) Enfuvirtide is a synthetic 36-amino-acid segment of the HR2 region of gp41, which prevents the formation of a six-helix fusion protein necessary for membrane fusion.
Nucleoside Reverse Transcriptase Inhibitors
The reverse transcriptase (RT) is a major target in anti-retroviral therapy. RT inhibitors are classified as nucleoside reverse transcriptase inhibitors (NRTIs) or non-nucleoside reverse transcriptase inhibitors (NNRTIs). Nucleosides (adenine, cytosine, guanine, and thymidine) consist of a phosphate group and a sugar molecule, either deoxyribose (DNA) or ribose (RNA). In the synthesis of DNA, nucleosides are linked together using 5′-3′-hydroxyl linkages between sugars. In many of the NRTIs, the hydroxyl group at the 3′ position in the sugar is replaced by another molecule. The NRTIs can be inserted into new DNA, but chain synthesis is terminated (Table 26-1).
Table 26-1
Nucleoside or Nucleotide Reverse Transcriptase Inhibitors
| Drug | Chemical Structure |
| Zidovudine | Thymidine analog, 3’ azido group (–N3) replaces the 3’ hydroxyl group |
| Abacavir | Synthetic nucleoside, which lacks the 3’-OH group on sugar groups |
| Didanosine | Adenosine analog, which lacks the 3’-OH group on sugar groups |
| Emtricitabine | Synthetic cytidine analog, which has a fluorine in the 5’ position |
| Stavudine | Synthetic thymidine analog, which lacks hydroxyl radical at the 3’ position |
| Tenofovir | Analog of adenosine 5’ phosphate. Converted to an acrylic nucleoside phosphonate. |
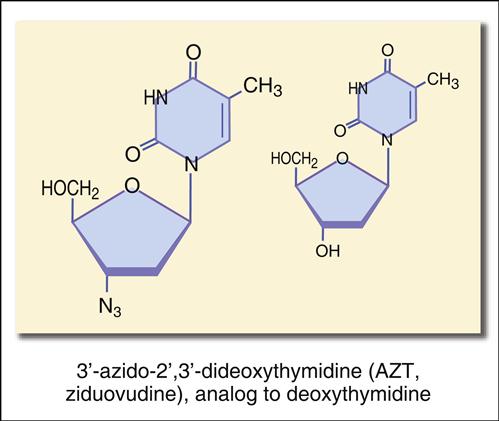
The most studied reverse transcriptase inhibitor is zidovudine or 3′-azido-2′,3′-dideoxythymidine (Figure 26-6).
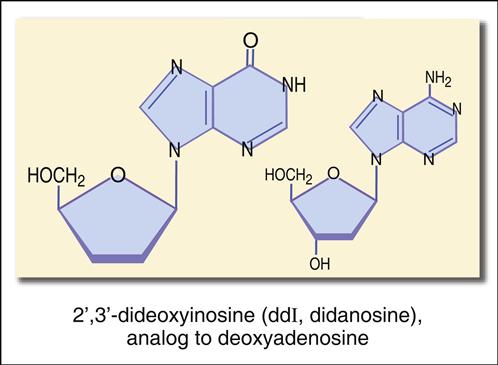
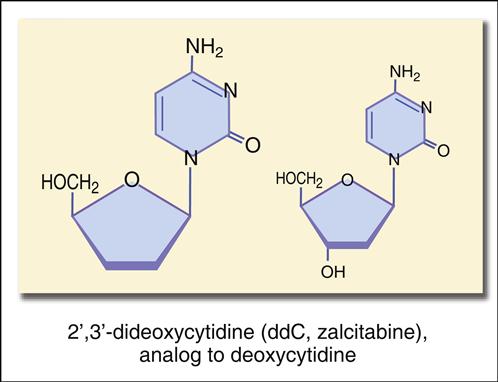
The structure mimics thymidine. However, a 3′ azido group (N3) replaces the 3′ hydroxyl group on the nucleotide. Other synthetic analogs are dehydroxylated forms of purines and pyrimidines. For example, didanosine is an adenosine analog, and zalcitabine is a cytosine analog (Figures 26-7 and 26-8).
Non-Nucleoside Reverse Transcriptase Inhibitors
Although the NNRTIs have different chemical structures, they are all highly specific to the HIV-1 RT. They bind to the RT and inhibit RNA and DNA polymerase activity. Delavirdine, etravirine, and nevirapine are NNTRIs that are currently licensed in the United States.
Protease Inhibitors
Protease inhibitors prevent the cleavage of the large gag–pol polyproteins into active proteins and enzymes. Protease inhibition results in disorganized capsid or matrix formation and the production of noninfectious viruses. Box 26-2 shows the currently licensed agents that are competitive inhibitors of the HIV protease.
Integrase Inhibitors
Integrase is an attractive drug target because integration of viral DNA into the host genome is critical to the infective process, and integrase has no functional analog in humans. Raltegravir is the only nontoxic, selective integrase inhibitor currently licensed by the FDA. This drug acts by inhibiting single-strand transfer of viral DNA to the host genome.
Summary