Viruses
After reading this chapter, the student will be able to:
• Describe the two commonly used methods of classifying viruses
• Define and describe capsid, capsomeres, nucleocapsid, virion, and envelope
• Describe and differentiate the types of viral genomes
• Describe and explain the steps in the multiplication of bacteriophages and animal viruses
• Discuss the possible effects of viral infections
• Describe the different kinds of host cell damage caused by viral infections
• Name the DNA viruses infecting and subsequently causing diseases in humans
• Name the RNA viruses infecting and subsequently causing disease in humans
• Compare and contrast viroids and virusoids
• Describe prions and the infections/diseases caused by them
General Structure and Classification
Classification
For the determination of order, the type of nucleic acid DNA or RNA, single or double stranded, and the presence or absence of an envelope are used. In addition, other characteristics such as the type of host, the capsid shape, immunological properties, and the type of disease the virus causes are also considered for further detailed virus classification. An additional classification system is the Baltimore classification system, devised by David Baltimore. This system places the virus into one of seven groups distinguishing the viruses on the basis of the relationship between the viral genome and the messenger RNA (Box 7.1). In modern virus classification the ICTV system is used in conjunction with the Baltimore classification system. Although animal and plant virologists both use these systems, the actual application in the process of classification is quite different because of the diversity of viruses.
HEALTHCARE APPLICATION
Medically Important Viruses
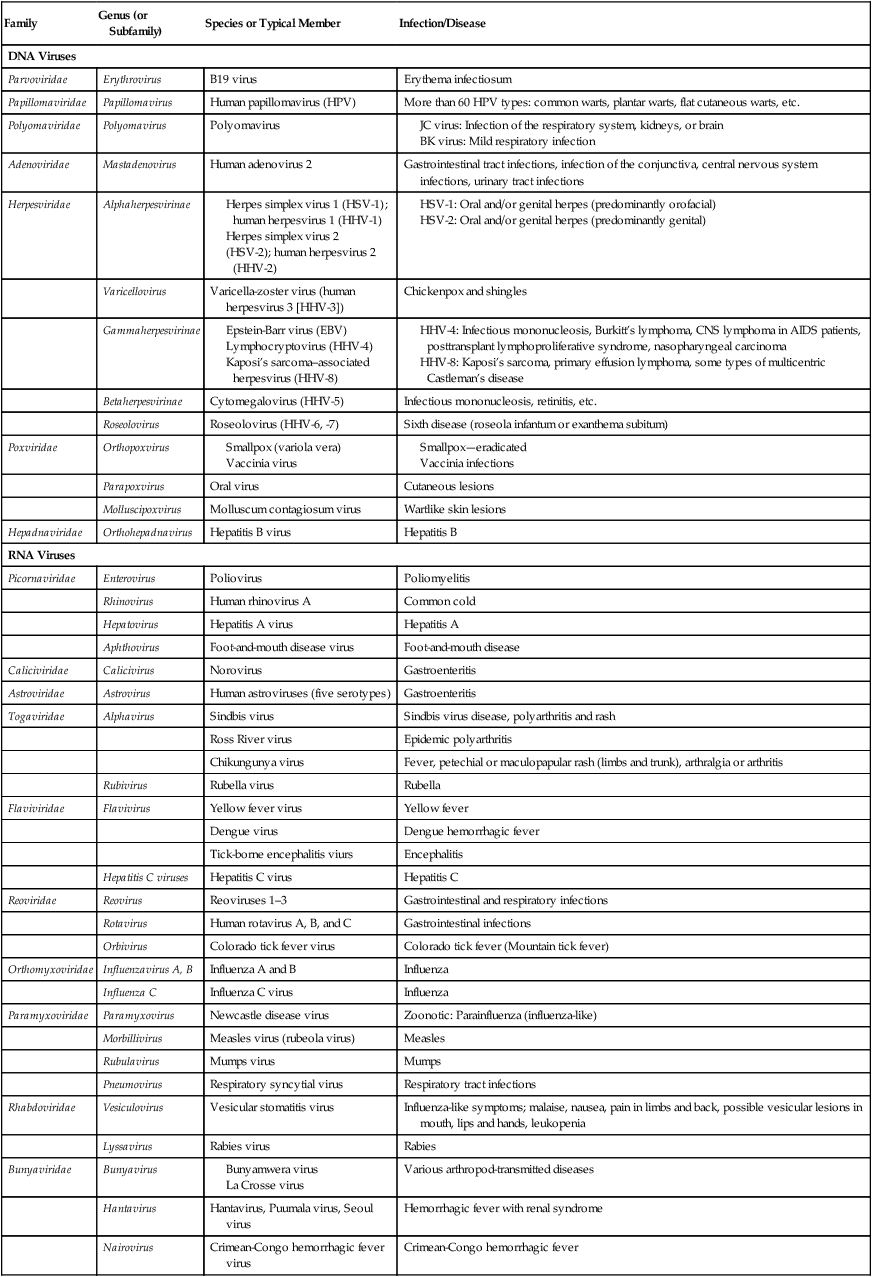
| Family | Genus (or Subfamily) | Species or Typical Member | Infection/Disease |
| DNA Viruses | |||
| Parvoviridae | Erythrovirus | B19 virus | Erythema infectiosum |
| Papillomaviridae | Papillomavirus | Human papillomavirus (HPV) | More than 60 HPV types: common warts, plantar warts, flat cutaneous warts, etc. |
| Polyomaviridae | Polyomavirus | Polyomavirus | |
Morphology
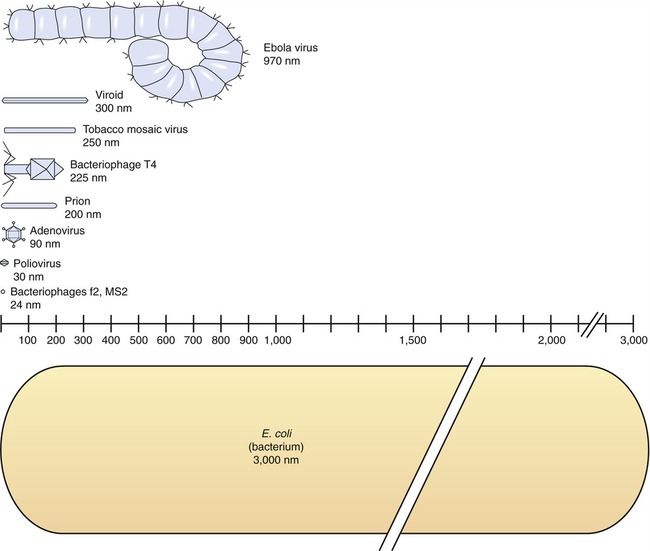
The size of viruses varies from very small, such as the parvovirus at about 20 nm and the poliovirus at 30 nm, to fairly large, such as the vaccinia virus at 400 nm and the poxviruses, which can be up to 450 nm. Most viruses cannot be visualized by light microscopy; therefore electron microscopy is used to examine their structure (see Chapter 1, Scope of Microbiology). However, some viruses are as large as or larger than the smallest bacteria and can be visualized by high optical magnification (Figure 7.1).
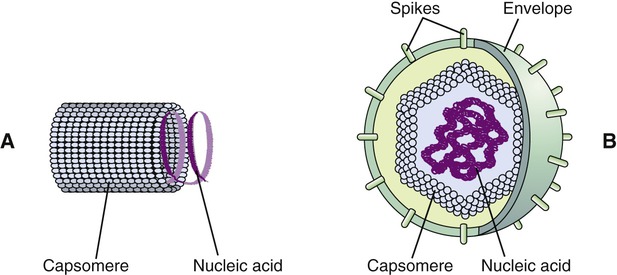
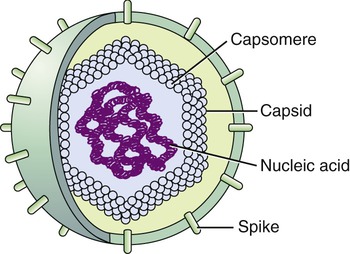
Viruses consist of genetic material carried in a “shell” called the viral coat or capsid. The capsid consists of proteins that are coded by the viral genome. The capsid is a complex and highly organized entity that gives form to the virus and serves as the basis for morphological distinction. Subunits referred to as protomeres assemble to form capsomeres, which in turn spontaneously aggregate to form the capsid. Proteins associated with nucleic acids are called nucleoproteins and the association of viral capsid proteins with viral nucleic acid is referred to as a nucleocapsid. At the junction points of the capsomeres, long projections from the nucleocapsids, called spikes, are frequently attached (Figure 7.2). These spikes aid in attachment to the host cell membrane receptors. Some viruses have an envelope around the coat and once the virus is fully assembled it is called a virion. Depending on the presence or absence of an envelope, viruses are referred to as naked or enveloped viruses (Figure 7.3).
Helical Viruses
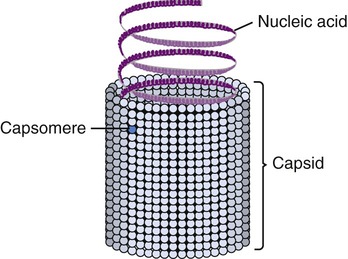
Helical capsids have rod-shaped capsomeres, connected along their long axis, resembling a wide ribbon stacked around a central axis forming a helical structure with a central cavity, a hollow tube. As a result, these virions are rod shaped or filamentous and can be short and highly rigid, as in many plant viruses, or long and very flexible, as in many animal viruses. The genetic material can be single-stranded RNA (ssRNA) or single-stranded DNA (ssDNA) and is bound into the protein helix by the interactions between the negative charge on the nucleic acid and the positive charge on the proteins. The nucleocapsid of a naked helical virus is rigid and tightly wound into a cylinder-shaped structure. An example is the well-studied tobacco mosaic virus, an ssRNA virus that infects primarily tobacco plants and other members of the Solanaceae family (Figure 7.4). Enveloped helical nucleocapsids are more flexible and this type is found in several human viruses such as influenza and measles viruses.
Icosahedral Viruses
Capsids of many virus families are multifaced structures known as icosahedrons. These are three-dimensional, geometric figures with 12 corners, 20 triangular faces, and 30 edges (Figure 7.5). This icosahedral capsid symmetry results in a spherical appearance of viruses at low magnification. The arrangement of the capsomeres varies between the viruses and the shape and dimension of the icosahedron depend on the characteristics of its protomeres. Although the basic symmetry of icosahedral viruses is the same there are major variations in the number of capsomeres, and therefore differences in the size of the viruses. The basic triangular face of a small virus is constructed of 3 protomeres, with 60 of these subunits forming the whole capsid. For example, the poliovirus has 32 capsomeres, whereas an adenovirus has 240 capsomeres. During the assembling of an icosahedral virus the nucleic acid is packed into the center, forming a nucleocapsid. Icosahedral viruses can be naked, such as the adenovirus, or enveloped, as the herpes simplex virus (Figure 7.6). Included in this morphological group are the Iridoviridae (infect mainly invertebrates), Herpesviridae, Adenoviridae, Papovaviridae, and Parvoviridae.
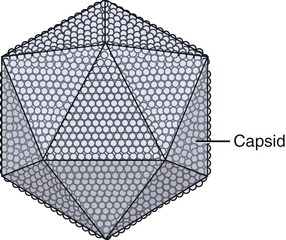
The capsids of these viruses are called icosahedrons: three-dimensional, geometric figures with 12 corners, 20 triangular faces, and 30 edges.
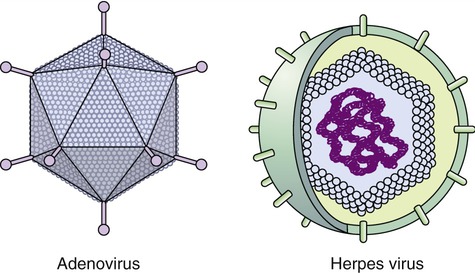
Enveloped Viruses
Many viruses have an outer structure, the viral envelope, which surrounds the nucleocapsid. Enveloped viruses typically obtain their envelope by budding though a host plasma membrane (see Plasma Membrane in Chapter 3, Cell Structure and Function) but may also include some viral glycoproteins. In some cases the viral envelope is derived from other membranes in the host cell, such as from the membranes of the endoplasmic reticulum or the nuclear membrane. The creation of the viral envelope through the budding process allows the viral particles to leave the host cell without disrupting the plasma membrane and therefore without killing the cell. As a result, some budding viruses can set up persistent infections.
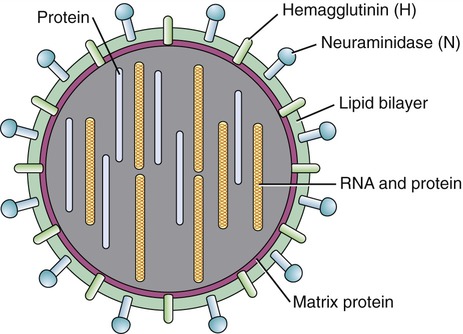
During the formation of the envelope some or all of the host membrane proteins are replaced by viral proteins and some form a layer between the capsid and the envelope. These envelope proteins are glycoproteins; they are exposed to the outside of the envelope as spikes. The lipid bilayer of the envelope is exclusively host specific whereas the majority or all of the glycoproteins are virus specific. The glycoproteins on the surface of the envelope serve to identify and bind to receptor sites on the plasma membrane of the host. Parts of the viral capsid and/or envelope are also responsible for stimulating the immune system of the host to produce antibodies that can neutralize the virus and prevent further infection (see Chapter 20, The Immune System). The viral envelope can give a virion protection from certain enzymes and chemicals; however, because enveloped viruses depend on their intact envelope to be infectious, agents that damage the envelope, such as alcohol and detergents, greatly reduce the infectivity of the virus (see Chapter 19, Physical and Chemical Methods of Control). An example of an enveloped virus is the influenza virus shown in Figure 7.7.
Complex Viruses
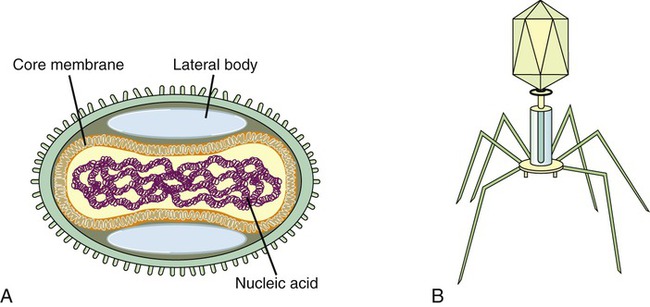
Complex viruses are a special group of viruses that consist of a capsid that is neither purely helical nor completely icosahedral and that has extra structures such as protein tails or a complex outer wall. The poxviruses are large, complex DNA viruses with an unusual morphology. These viruses lack a regular capsid and the viral genome is associated with proteins within a central disk structure, the nucleoid, which is surrounded by a membrane and two lateral bodies. The nucleoid is surrounded by several layers of lipoproteins and a coarse surface of fibrils (Figure 7.8, A).
An even more complex group of viruses are bacteriophages, viruses that infect bacteria. In general, bacteriophages have an icosahedral head bound to a polyhedral tail, which is attached to a base plate with many protruding protein tail fibers (Figure 7.8, B). Most bacteriophages contain double-stranded DNA (dsDNA); however, ssDNA- and RNA-type phages also exist. There are at least 12 separate groups of bacteriophages (Table 7.1), which are structurally and genetically diverse. Probably the most widely studied bacteriophages are the ones that infect Escherichia coli, a bacterium found in the normal flora of the intestinal tract (see Chapter 9, Infection and Disease). These bacteriophages are called coliphages and include the lambda phage and the T phages. Bacteriophages exist in different sizes and shapes, but the basic structural features are the same (Figure 7.9):
TABLE 7.1
| Family or Group | Genus | Type Member | Particle Morphology | Envelope | Genome |
| Corticoviridae | Corticovirus | PM2 | Isometric | No | Supercoiled dsDNA |
| Cystoviridae | Cystovirus | ϕ6 | Isometric | Yes | Three segments, dsRNA |
| Inoviridae | Inovirus | Coliphages fd | Rod | No | Circular ssDNA |
| Plectrovirus | Acholeplasma phage | ||||
| Leviviridae | Levivirus | Coliphage MS2 | Icosahedral | No | 1(+)strand (+)ssRNA |
| Allolevirus | Coliphage Qbeta | ||||
| Lipothrixiviridae | Lipothrixvirus | Thermoproteus phage 1 | Rod | Yes | Linear dsDNA |
| Microviridae | Microvirus | Coliphages ϕX174 | Icosahedral | No | Circular ssDNA |
| Spirovirus | Spiroplasma phages | ||||
| Mac-1 phage | |||||
| Myoviridae | Coliphages T4 | Tailed phage | No | Linear dsDNA | |
| Plasmaviridae | Plasmavirus | Acholeplasma phage | Pleomorphic | Yes | Circular dsDNA |
| Podoviridae | Coliphage T7 | Tailed phage | No | Linear dsDNA | |
| Siphoviridae | Lambda phage group | Coliphage lambda | Tailed phage | No | Linear dsDNA |
| Sulfolobus Shibatae virus | SSV-1 | Lemon shaped | No | Circular dsDNA | |
| Tectiviridae | Tectivirus | Phage PRD1 | Icosahedral | No | Linear dsDNA |
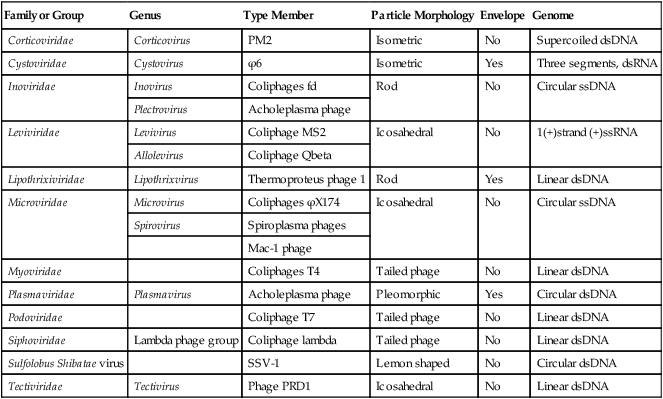
dsDNA, Double-stranded DNA; dsRNA, double-stranded RNA; ssDNA, single-stranded DNA.
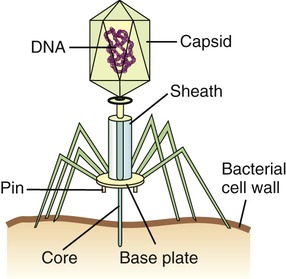
Bacteriophages, although different in size and shape, all have a capsid that contains the nucleic acid. Many bacteriophages also have a complex tail that enables the bacteriophage to attach to the host and inject the nucleic acid.
• Head or Capsid: All bacteriophages possess a head structure that varies in size and shape. It is made of many copies of one or more different proteins and the nucleic acid is located within this structure. The head or capsid therefore acts as the protective covering for the nucleic acid.
• Tail: Many bacteriophages have tails attached to the head, whereas others do not. The tail is a hollow tube that serves as a channel for the nucleic acid to pass through during the process of infecting a bacterium. Its size also varies depending on the phage type. Some of the more complex phages have a tail that is surrounded by a sheath, which contracts during infection of the bacterium. These phages also have a base plate at the end of the tail, with one or more tail fibers attached to it. The base plate and tail fibers play a major role in binding of the bacteriophage to the bacterium. Phages without base plate and tail fibers have other structures involved in binding of the phage to the bacterium.
Viral Multiplication
Multiplication of Bacteriophages
The events that occur in the multiplication cycle of bacteriophages include adsorption, penetration, replication, assembly, maturation, and release. Adsorption represents the recognition process between phage and bacterium, followed by attachment and then penetration of the phage DNA into the bacterium. Replication represents copying of phage nucleic acid, followed by the assembly of phage parts and maturation of the newly formed bacteriophages. The mature phages are then released from the host cell and are ready to infect others. These events are shown in Figure 7.10.
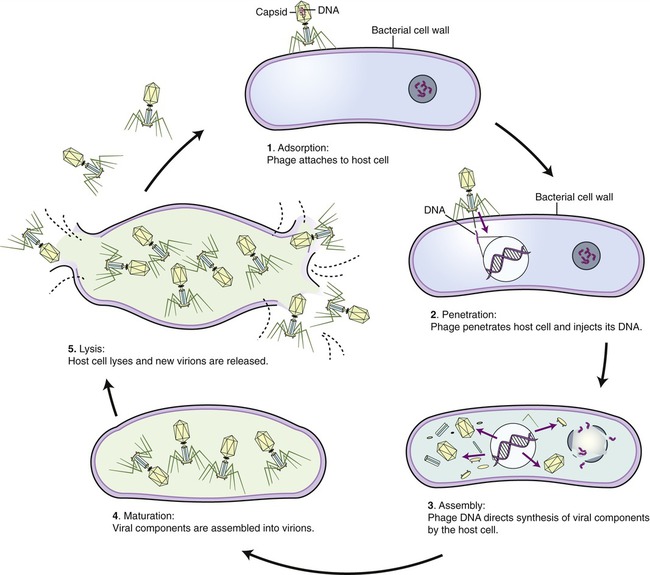
Assembly
• Proteins that seal the damaged bacterial cell wall to prevent premature bacterial cell death
• Enzymes that are necessary to copy the genome of the bacteriophage
• Proteins that are needed for the formation of the capsid head and part of the tail fibers
• Enzymes required for the weakening of the bacterial cell wall in order to lyse the bacterial cell and release the newly assembled bacteriophages
• Lytic or virulent phages are phages that after multiplication destroy the bacterium by lysis at the end of the multiplication cycle, releasing the newly formed phages into the environment. This lytic life cycle can be divided into three phases: the eclipse period, the intracellular accumulation phase, and the lysis and release phase.
• Eclipse period: In this phase no infectious phage particles can be found in the host cell. The viral synthesis process utilizes the resources of the host and the phage nucleic acid takes over the biosynthetic machinery to produce phage specific mRNA’s to make proteins. In the early phase of this period, early mRNA’s code for proteins which are needed for the synthesis of phage DNA and synthesis of host DNA, RNA and proteins come to an end. After the production of phage DNA late mRNAs and their corresponding proteins are made. These late proteins are structural proteins for the phages and also proteins necessary for the lysis of the host cell.
• Intracellular accumulation phase: The nucleic acids and the structural proteins made in the previous phase are assembled into complete infectious phages that accumulate in the host cell.
• Lysis and release phase: During this phase the bacteria begin to lyse, due to the accumulation of phage lysis proteins, and the phages are released into the environment surrounding the bacterium. Infected bacteria may release up to 1000 bacteriophages.
• Lysogenic or temperate phages are those that can either multiply by the lytic cycle or undergo adsorption and penetration into the bacterial host but are not replicated or released at that time. Instead the phage genes are not transcribed, and the phage genome exists in a repressed state. The phage DNA in this stage is referred to as a prophage, because it is not a phage but has the potential to produce a phage. In general, the bacteriophage DNA is integrated into the host chromosome and passed on to the daughter cells of the bacterium. The host cell carrying a prophage might not be adversely affected at all and the lysogenic state might persist indefinitely or eventually result in lysis. Lysogeny is a less deadly form of parasitism, allowing the virus to spread without killing the host. On occasion the prophage in a lysogenic cell can be activated, which ultimately results in viral replication and the lytic cycle.
Multiplication of Animal Viruses
The basic stages of multiplication of animal viruses are similar to those seen in bacteriophages. However, several differences are present between the host cells, their viruses, and also in some steps of the multiplication cycle. The stages of the multiplication cycle for animal viruses include adsorption, penetration, uncoating, replication, assembly, and finally release from the host cell. The duration of the multiplication cycle varies among animal viruses. The basic stages of the cycle in naked and enveloped viruses are shown in Figure 7.11.
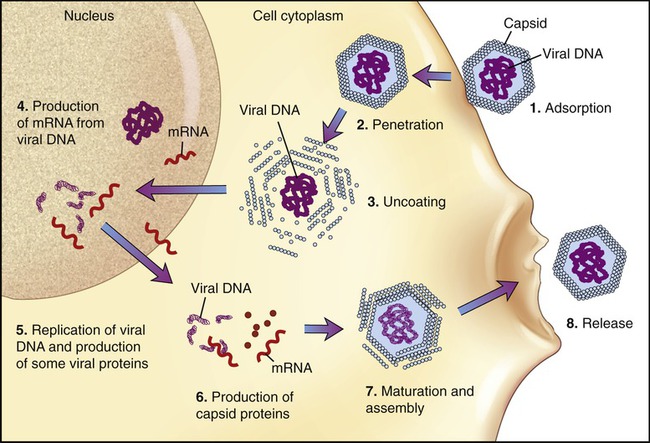
Adsorption
The first stage of multiplication is the attachment (adsorption) of the virus to a susceptible host cell. Like bacteriophages, animal viruses are specific to their host cells and they infect only one or a restricted range of host species. This limitation is referred to as the host range. For example, hepatitis viruses attack cells in the liver, the mumps virus attacks the salivary glands, and HIV attaches to the CD4 protein of helper T cells (see Chapter 20, The Immune System). They adsorb to specific receptor sites embedded in the plasma membrane of their host, which are generally glycoproteins the host cell requires for its normal cellular functions. The attachment sites on the virus (viral receptors) are distributed over the capsid or envelope of the virus and are generally glycoproteins or proteins. Animal cells that lack receptors for a specific virus are resistant and cannot be infected by that virus. Naked and enveloped viruses differ in their mode of attachment to the host cell. Naked viruses have surface molecules that adhere to the membrane receptor, whereas enveloped viruses possess glycoprotein spikes that bind to the plasma membrane receptors of the host cells (Figure 7.12). The attachment of a virus to its host can be prevented by antibody molecules that are capable of attaching to the viral attachment sites or to the host cell receptors. The presence of these antibodies in the host organism is the most important basis for immunity against viral infections (see Chapter 20).
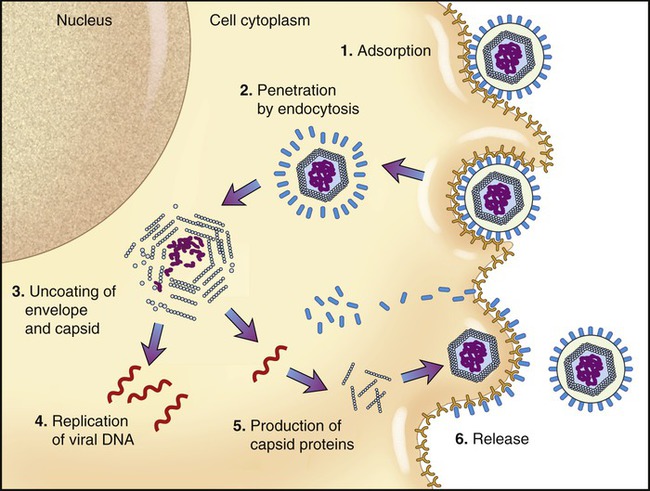
Uncoating
When the nucleocapsid is in the host cell cytoplasm the viral nucleic acid is released by a process called uncoating. The uncoating process varies between viruses. If penetration occurs by receptor-mediated endocytosis (see Chapter 3, Cell Structure and Function) the entire virus is engulfed by the host cell and becomes enclosed in a vacuole or vesicle. The enzymes in the vacuole or vesicle then dissolve the envelope and the capsid, and the viral genome is released. Another method of entry occurs by direct fusion of the viral envelope with the plasma membrane of the host cell, thereby releasing the nucleocapsid into the cell cytoplasm. Some antiviral drugs (see Chapter 22, Antimicrobial Drugs) exert their effect by preventing the uncoating of the nucleocapsid.
Replication
• An ssRNA with a positive-sense configuration can be used directly as mRNA.
• An ssRNA with a negative-sense configuration must first be transcribed into a positive-sense strand by viral polymerase and can then be used as mRNA.
• Retroviruses use a different route. These viruses have a positive-sense ssRNA that is made into a negative-sense ssDNA, using viral reverse transcriptase present in the nucleocapsid. This process is followed by the formation of dsDNA, which enters the nucleus and becomes integrated into the host DNA. The integrated viral DNA is then transcribed by host polymerase into mRNA.
Viral Infections
• Abortive infections are infections of nonpermissive cells where no viral production occurs.
• Lytic or cytocidal infections kill the host cell. This type of infection occurs in permissive cells, cells that provide the biosynthetic machinery to support the complete replication cycle of the virus.
• Persistent infections are infections that do not cause cell death. They can be:
• Chronic infections that are not lytic but productive. The cellular effects of chronic infections are mostly the same as in acute cytocidal infections with the exception that the production of viruses may be slower, intermittent or limited to only a few cells.
• Latent infections with limited macromolecular synthesis but no virus synthesis. In other words, only restricted expression of the viral genome occurs. The viral gene products are associated with few if any changes in the infected cell.
• Slow infections, characterized by a prolonged incubation period without prominent morphological and physiological changes to the host cell. Although the progression of cellular damage may take years, it is followed by extensive cellular injury and disease.
• Transforming infections, in which the viral nucleic acid may remain in specific host cells indefinitely, and a virus may or may not be produced. The term oncogenic transformation refers to the genetic modification of cell proliferation control so that the cell becomes cancerous. DNA or RNA tumor viruses may initiate several changes that convert normal cells into malignant tumor cells (see Genotoxic Effects later in this chapter). Infected cells may also undergo other types of inheritable changes resulting in morphological, physiological, and biochemical alterations that are nononcogenic. In other words, if a virus infects a cell and alters the properties of that cell, the cell is referred to as transformed.
Host Cell Damage
Morphological Effects
• Altered shape (i.e., rounding of cells)
• Detachment from tissue surface
• Altered membrane permeability
• Inclusion bodies (either altered host cell structure or accumulation of viral components)
The specific cytopathic effects caused by different viruses depend on the viruses and the cells they infect. The observations of cytopathic effects can be used in clinical virology to aid in the identification of a viral isolate. Cytopathic changes of selected viruses are listed in Table 7.2.
TABLE 7.2
Cytopathic Effects of Selected Viruses
| Virus | Cytopathic effect |
| Adenovirus | Nuclear inclusions; cell clumping |
| Herpes simplex virus | Cell fusion leading to multinucleated giant cells; nuclear inclusions |
| HIV | Multinucleated giant cells |
| Influenza virus | Rounding of cells |
| Poliovirus | Cell lysis |
| Rabies virus | Cytoplasmic inclusions: Negri bodies |
| Reovirus | Cell enlargement; vacuoles and inclusion bodies in cytoplasm |
| Smallpox virus | Rounding of cells; cytoplasmic inclusions |
Genotoxic Effects
Genotoxic substances damage DNA either directly, by binding to the DNA, or indirectly leading to DNA damage by affecting enzymes involved in DNA replication. These substances therefore cause mutations, which may or may not lead to cancer. After a viral infection, breakage, fragmentation, rearrangement, or other changes in the host DNA may occur. Double-stranded DNA viruses interact directly with the host DNA and in some cases the viral DNA becomes integrated at a particular site in the host genome. All parts of the viral genome are expressed in permissive cells, which leads to viral replication, lysis, and cell death. When the viral DNA becomes integrated into the host genome at random sites in nonpermissive cells, only part of the viral genome is expressed. In this case the early control functions of the virus are expressed, while viral structural proteins are not produced. Several DNA viruses such as the hepatitis B virus, the herpesviruses, and the papillomaviruses are potentially initiators of cancer. RNA tumor viruses (retroviruses) usually change cells into malignant cells by integrating their own nucleic acid into the host cell genome and may even produce infectious new viruses. Because RNA is the genome of the mature viral particle it must first be copied to DNA (see Viral Multiplication earlier in this chapter) before integration into the host DNA.
HEALTHCARE APPLICATION
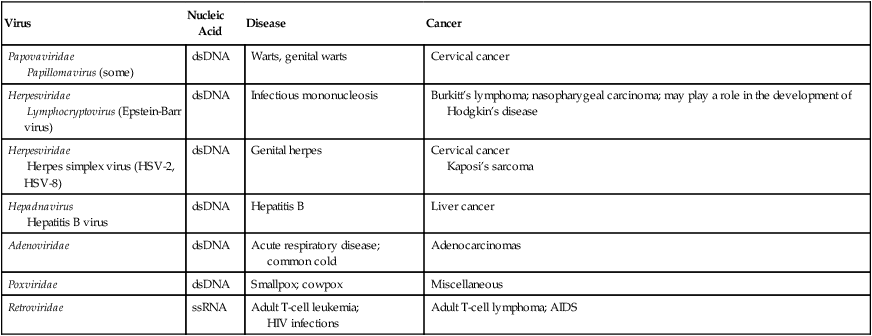
Oncogenic Viruses and Associated Cancers
| Virus | Nucleic Acid | Disease | Cancer |
| Papovaviridae Papillomavirus (some) |
dsDNA | Warts, genital warts | Cervical cancer |
| Herpesviridae Lymphocryptovirus (Epstein-Barr virus) |
dsDNA | Infectious mononucleosis | Burkitt’s lymphoma; nasopharygeal carcinoma; may play a role in the development of Hodgkin’s disease |
| Herpesviridae Herpes simplex virus (HSV-2, HSV-8) |
dsDNA | Genital herpes | Cervical cancer Kaposi’s sarcoma |
| Hepadnavirus Hepatitis B virus |
dsDNA | Hepatitis B | Liver cancer |
| Adenoviridae | dsDNA | Acute respiratory disease; common cold | Adenocarcinomas |
| Poxviridae | dsDNA | Smallpox; cowpox | Miscellaneous |
| Retroviridae | ssRNA | Adult T-cell leukemia; HIV infections |
Adult T-cell lymphoma; AIDS |
Major Groups of Viruses in Vertebrates
DNA Viruses
Adenoviruses
Hepadnaviruses
Hepadnaviruses are part of the family Hepadnaviridae and cause hepatitis in humans and animals. These viruses are enveloped virions with a small circular genome of partially double-stranded and partially single-stranded DNA. When the virus enters the host cell, the single-stranded portion is restored to a double strand by a DNA polymer present in the core. One member of the group is the hepatitis B virus (HBV), which infects hepatocytes where it can persist, causing hepatitis, and can also be a factor in liver cell carcinoma. HBV can cause acute or chronic, symptomatic or asymptomatic disease (see Chapter 12, Infections of the Gastrointestinal System).
Herpesviruses
Herpesviruses (Herpesviridae) are a large family of DNA viruses that cause disease in humans and animals. The human herpesviruses are grouped into three subfamilies: Alphaherpesvirinae, Betaherpesvirinae, and Gammaherpesvirinae. This classification is based on differences in viral characteristics such as genome structure, tissue tropism, cytopathology, and site of latent infection, as well as pathogenesis of the disease (Table 7.3). The human herpesviruses are as follows:
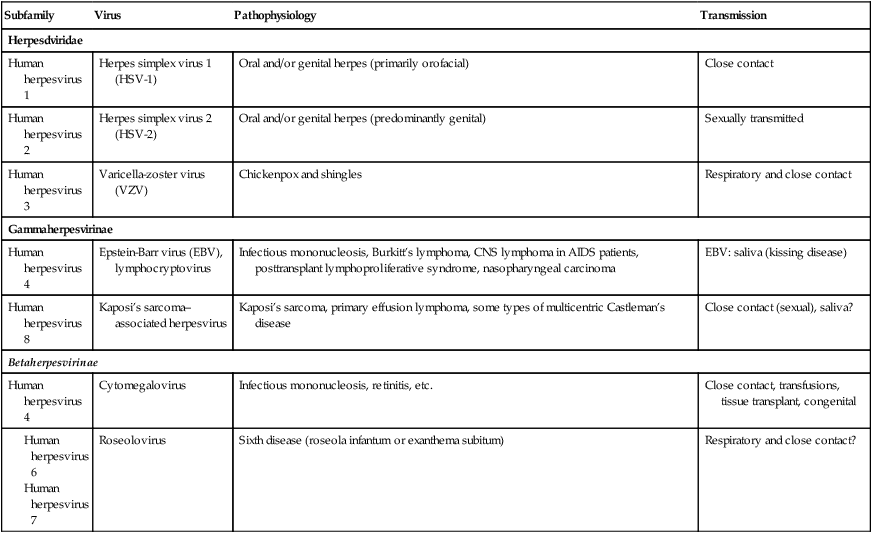
TABLE 7.3
Characteristics of Herpesviridae
| Subfamily | Virus | Pathophysiology | Transmission |
| Herpesdviridae | |||
| Human herpesvirus 1 | Herpes simplex virus 1 (HSV-1) | Oral and/or genital herpes (primarily orofacial) | Close contact |
| Human herpesvirus 2 | Herpes simplex virus 2 (HSV-2) | Oral and/or genital herpes (predominantly genital) | Sexually transmitted |
| Human herpesvirus 3 | Varicella-zoster virus (VZV) | Chickenpox and shingles | Respiratory and close contact |
| Gammaherpesvirinae | |||
| Human herpesvirus 4 | Epstein-Barr virus (EBV), lymphocryptovirus | Infectious mononucleosis, Burkitt’s lymphoma, CNS lymphoma in AIDS patients, posttransplant lymphoproliferative syndrome, nasopharyngeal carcinoma | EBV: saliva (kissing disease) |
| Human herpesvirus 8 | Kaposi’s sarcoma–associated herpesvirus | Kaposi’s sarcoma, primary effusion lymphoma, some types of multicentric Castleman’s disease | Close contact (sexual), saliva? |
| Betaherpesvirinae | |||
| Human herpesvirus 4 | Cytomegalovirus | Infectious mononucleosis, retinitis, etc. | Close contact, transfusions, tissue transplant, congenital |
The herpesviruses are large enveloped dsDNA (linear) viruses that acquire their envelope by budding from nuclear and plasma membranes of the host cell. They replicate in the nucleus of infected cells and characteristically demonstrate latency, which allows them to exist in a dormant stage for the duration of the host tissue’s lifetime. When activated they cause recurrent disease in their hosts (e.g., varicella-zoster virus; see Chapter 10, Infections of the Integumentary System, Soft Tissue, and Muscle). Reactivation of the viruses can occur in response to environmental changes such as ultraviolet light, x-rays, heat, cold, hormonal changes, stress (both physical and emotional), and immune deficiencies.
RNA Viruses
Bunyaviridae
Bunyaviridae species are enveloped viruses with a large negative-stranded RNA as their nucleic acid. They are zoonoses and while generally found in animals, certain members in this family can infect humans (Table 7.4). Bunyaviridae comprises a group of at least 200 viruses broken down into five genera based on structural and biochemical features: Bunyavirus, Phlebovirus, Uukuvirus, Nairovirus, and Hantavirus. With the exception of the hantavirus, which is carried by rodents, Bunyaviridae species are arthropod-borne.
TABLE 7.4
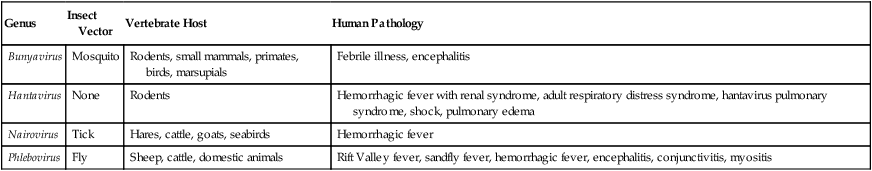
Examples of Bunyaviridae Capable of Infecting Humans
| Genus | Insect Vector | Vertebrate Host | Human Pathology |
| Bunyavirus | Mosquito | Rodents, small mammals, primates, birds, marsupials | Febrile illness, encephalitis |
| Hantavirus | None | Rodents | Hemorrhagic fever with renal syndrome, adult respiratory distress syndrome, hantavirus pulmonary syndrome, shock, pulmonary edema |
| Nairovirus | Tick | Hares, cattle, goats, seabirds | Hemorrhagic fever |
| Phlebovirus | Fly | Sheep, cattle, domestic animals | Rift Valley fever, sandfly fever, hemorrhagic fever, encephalitis, conjunctivitis, myositis |

Hepatitis Viruses
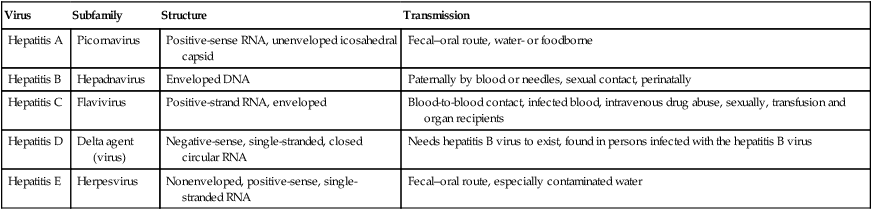
Many of the liver diseases, collectively called hepatitis, are caused by viruses, coming from a wide range of viral families (Table 7.5). Each of the hepatitis viruses infects and damages the liver, causing jaundice and the release of liver enzymes. Viral hepatitis is a cause of considerable morbidity and mortality in the human population, both from acute and chronic infection, including chronic active hepatitis, cirrhosis, and hepatocellular carcinoma. Viral hepatitis is a major public health problem throughout the world, affecting several hundreds of millions of people.
TABLE 7.5
| Virus | Subfamily | Structure | Transmission |
| Hepatitis A | Picornavirus | Positive-sense RNA, unenveloped icosahedral capsid | Fecal–oral route, water- or foodborne |
| Hepatitis B | Hepadnavirus | Enveloped DNA | Paternally by blood or needles, sexual contact, perinatally |
| Hepatitis C | Flavivirus | Positive-strand RNA, enveloped | Blood-to-blood contact, infected blood, intravenous drug abuse, sexually, transfusion and organ recipients |
| Hepatitis D | Delta agent (virus) | Negative-sense, single-stranded, closed circular RNA | Needs hepatitis B virus to exist, found in persons infected with the hepatitis B virus |
| Hepatitis E | Herpesvirus | Nonenveloped, positive-sense, single-stranded RNA | Fecal–oral route, especially contaminated water |
Orthomyxoviruses
Orthomyxoviruses belong to the family Orthomyxoviridae and include the influenza A, B, and C viruses. The viruses are enveloped with a segmented negative-sense RNA genome. Influenza A and C viruses infect multiple species, whereas influenza B infects mostly humans. Significant human disease is due only to influenza A and B viruses. The type A viruses are the most virulent human pathogens in this group and cause the most severe disease. Influenza viruses are respiratory viruses (see Chapter 11, Infections of the Respiratory System) responsible for the classic flulike symptoms of fever, malaise, headache, and body aches (myalgias). Influenza is one of the most prevalent of the viral infections. The segmented nature of the influenza virus genome makes the development of new strains possible through mutation and rearrangement of gene segments between human and animal strains of the virus. This genetic instability and the resulting emergence of new strains are responsible for the annual flu epidemics and periodic pandemics (see Chapter 18, Emerging Infectious Diseases). The prophylactic vaccines and antiviral drugs that are now available make it possible to prevent serious disease complications in people at risk. Example of known flu pandemics (worldwide) are given in Table 7.6.
TABLE 7.6
| Name of Pandemic | Date | Estimated Deaths |
| Asiatic (Russian) flu | 1889–1890 | 1 million |
| Spanish flu | 1918–1920 | 40 million |
| Asian flu | 1957–1958 | 1 to 1.5 million |
| Hong Kong flu | 1968–1969 | 0.75 to 1 million |
Picornaviruses
Two main categories causing a variety of human infections are the enteroviruses and the rhinoviruses. They are distinctly different in the stability of their capsid at low pH as well as in their optimal temperature for growth. The capsid of enteroviruses is stable at the pH 3, which allows them to survive under harsh environmental conditions such as the gastrointestinal tract and sewer systems. They are transmitted by the fecal–oral route. In contrast, the rhinoviruses are acid labile; they infect primarily the nose and throat; and they grow best at 33° C (temperature of the nasal cavity). Enteroviruses replicate at 37° C. The portal of entry (see Chapter 9, Infection and Disease) can be the upper respiratory tract, the oropharynx, and the gastrointestinal tract. Most enteroviruses are cytolytic; they replicate rapidly and cause direct damage to the infected cells. In the case of poliovirus, the virus infects skeletal muscle and gains entry to the nervous system by traveling along the innervating nerve fibers (see Chapter 13, Infections of the Nervous System and Senses). Rhinoviruses are the most common initiators of the common cold and upper respiratory tract infections. There are 115 or more serotypes of rhinovirus; their infections are self-limiting and generally do not cause serious disease.
Reoviruses
The name reovirus was given to a group of enteric and respiratory viruses not associated with a particular disease and were considered to be “respiratory, enteric, orphans.” Reoviruses are nonenveloped with an icosahedral nucleocapsid and a dsRNA genome. The virion is rather resistant to undesirable environmental and gastrointestinal conditions such as variations in pH and temperature, detergents, and drying. One of the genera of reoviruses is Rotavirus, the causative agent of human infantile gastroenteritis (see Chapter 12, Infections of the Gastrointestinal System), a common disease in children. The virus accounts for approximately 50% of all case of diarrhea in children requiring hospitalization because of dehydration. Rotaviral infections are a major problem in underdeveloped countries, where the virus is responsible for more than 1 million deaths per year. Rotaviruses are also one of the causes of “traveler’s” diarrhea.
Retroviruses
Retroviruses are classified according to the disease they cause, tissue tropism and host range, virion morphology, and complexity of the genome. The subfamilies of the human retroviruses are Oncovirinae, Lentivirinae, and Spumavirinae (Table 7.7). The viruses are enveloped, somewhat spherical in shape, and their genome consists of two identical copies of (+) ssRNA. These viruses have a unique replication cycle: they encode an RNA-dependent DNA polymerase (reverse transcriptase), which allows them to replicate through a DNA intermediate. The DNA copy of the viral genome, called a provirus, is then integrated into the host cell DNA and thereby becomes a cellular gene.
TABLE 7.7
| Subfamily | Characteristics |
| Oncovirinae | Associated with cancer and neurological disorders |
| Type B | Unconventional nucleocapsid in mature virus |
| Type C | Centrally located nucleocapsid core in mature virion |
| Type D | Cylindrical nucleocapsid core |
| Lentivirinae | Slow disease onset: neurologic disorders and immunosuppression; are D-type viruses (cylindrical nucleocapsid core) |
| Spumavirinae | No clinical disease, but characteristic vacuolated “foamy” cytopathology |
| Endogenous viruses | Retrovirus sequence that can be integrated into the human genome |
Subviral Agents
Viroids
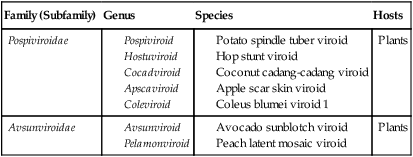
Viroids range in size from 15 to 100 nm in diameter, and their RNA genomes are 246 to 375 nucleotides in length. They are all single-stranded circles; replication does not depend on the presence of a helper virus, and no proteins are encoded. They are common plant pathogens causing serious economic problems. Although they do not have a protective protein coat around the nucleic acid, these naked RNA pathogens can cause disease in susceptible plants. These plants include potatoes, tomatoes, cucumbers, palm trees, and certain fruit trees. The current taxonomy of viroids is shown in Table 7.8.
TABLE 7.8
| Family (Subfamily) | Genus | Species | Hosts |
| Pospiviroidae |
Prions
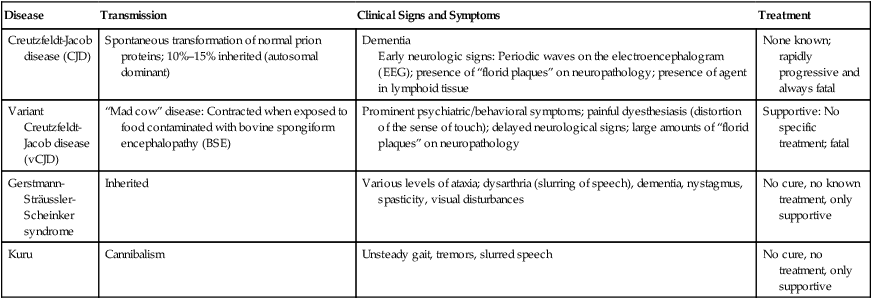
HEALTHCARE APPLICATION
Examples of Human Prion Diseases
| Disease | Transmission | Clinical Signs and Symptoms | Treatment |
| Creutzfeldt-Jacob disease (CJD) | Spontaneous transformation of normal prion proteins; 10%–15% inherited (autosomal dominant) | Dementia Early neurologic signs: Periodic waves on the electroencephalogram (EEG); presence of “florid plaques” on neuropathology; presence of agent in lymphoid tissue |
None known; rapidly progressive and always fatal |
| Variant Creutzfeldt-Jacob disease (vCJD) | “Mad cow” disease: Contracted when exposed to food contaminated with bovine spongiform encephalopathy (BSE) | Prominent psychiatric/behavioral symptoms; painful dyesthesiasis (distortion of the sense of touch); delayed neurological signs; large amounts of “florid plaques” on neuropathology | Supportive: No specific treatment; fatal |
| Gerstmann-Sträussler-Scheinker syndrome | Inherited | Various levels of ataxia; dysarthria (slurring of speech), dementia, nystagmus, spasticity, visual disturbances | No cure, no known treatment, only supportive |
| Kuru | Cannibalism | Unsteady gait, tremors, slurred speech | No cure, no treatment, only supportive |
Summary
• Virus classification is primarily based on morphology, nucleic acid type, mode of replication, host organism, and the type of disease caused. The two classification systems used most often are the ICTV system and the Baltimore system.
• Viruses can infect bacteria, plants, and animals.
• In general, viruses are host specific, as well as specific to tissues and cells.
• According to the shape and arrangement of capsomeres and the presence or absence of an envelope, viruses can be classified into four distinct morphological types: helical, icosahedral, enveloped, and complex.
• The viral genome is represented by nucleic acids, either RNA or DNA, which can be in the form of single- or double-stranded molecules.
• Viruses do not replicate through cell division; they use the host cell machinery and metabolism to manufacture multiple copies of themselves. Although similar, the mechanism of replication varies between bacteriophages and animal viruses.
• Viral infections of animal cells can have different outcomes; they can be abortive, lytic, or persistent infections.
• Viral infections are usually associated with changes in the host cells and include morphological, physiological, and biochemical transformations.
• Subviral agents causing infections in plants, human, and animals are classified as viroids, virusoids (satellites), and prions.
Review Questions
2. The correct hierarchical order for virus classification is:
a. Order, genus, family, subfamily, species
b. Species, genus, family, subfamily, order
3. Viral capsomeres are composed of subunits called:
4. RNA viruses belong to which of the following groups according to the Baltimore classification system?
5. A phage in a repressed stage is referred to as a(n):
6. The correct sequence of stages in the multiplication of animal viruses is:
a. Adsorption, penetration, uncoating, replication, assembly, release
b. Penetration, adsorption, assembly, uncoating, replication, release
c. Adsorption, uncoating, penetration, replication, assembly, release
d. Penetration, uncoating, adsorption, replication, assembly, release
7. A persistent infection that is not lytic but productive is called a _____ infection.
8. All of the following are cytopathic effects as a result of a viral infection except:
9. Which of the following viruses belong to the family Herpesviridae?
10. Transmissible spongiform encephalopathies are caused by:
11. A virus that infects bacteria is referred to as a(n) __________.
12. Kaposi’s sarcoma is caused by human herpesvirus number __________.
13. Members of Picornaviridae are __________ viruses.
14. A fully assembled virus is called a(n) __________.
15. Group VI viruses according to the Baltimore classification system include the families Metaviridae, Pseudoviridae, and __________.
16. Discuss the two methods of viral classification.
17. Describe host cell damage resulting from viral infections.
18. Name and describe the different morphological types of viruses.
19. Describe and contrast the multiplication of bacteriophages versus animal viruses.
20. Explain and discuss the structure and pathogenicity of subviral agents.