3 Biochemistry and physiology of receptor activation
Case 3.2
Questions
• 3.2.1 What are the various second messenger pathways that can be activated by receptor stimulation in a neuron?
• 3.2.2 What interactions result from receptor stimulation on the expression DNA in the neuron?
• 3.2.3 How does the neuron generate enough energy to support its responses to receptor stimulation?
• 3.2.4 What is the central integrative state of a neuron and how is it influenced by receptor activation?
Introduction
Synaptic transmission has been conceptualised as a set of processes by which neurotransmitters, acting through their receptors, cause changes in the conductance of specific ion into and out of the neuron to produce excitatory or inhibitory postsynaptic potentials. However, it has become evident that neurotransmitters elicit diverse and complicated effects in target neurons. This has led to the development of a much more complex view of synaptic transmission (Huganir & Greengard 1990).
Phenotypic and functional development of neurons is accomplished through genetic lineage and environmentally induced gene activation
The initial modulation of genetic events by the environment takes place during the differentiation of stem cells into neurons. Neurons are neurons because they produce the proteins and enzymes necessary to carry out the functions of neurons. They can manufacture axons and dendrites because they are rich in tubulin and microtubule-associated proteins (Black & Baas 1989). They can maintain a membrane potential that varies within a specific range of values depending on the environmental circumstances because they manufacture gated plasma ion channels (Hess 1990). They can communicate with other neurons because they have neurotransmitter-specific enzymes to produce neurotransmitters (Snyder 1992). In other words, all or most of the necessary functions of a neuron are made possible by the activation of the genes that code for the proteins necessary to subserve those functions and the suppression of those genes that do not.
The gene repertoire available for transcription in a neuron is determined by the lineage of the neuron and the stage of commitment and differentiation that the neuron has achieved (Van den Berg C 1986; Pleasure 1992). There are usually temporally and environmentally dependent branch points in the development of a neuron lineage that can determine a particular course of differentiation or development that the neuron will take (Lillien & Raff 1990). For instance, during one critical phase or branch point in the development of a neuron the type of neurotransmitters that the neuron will be producing is determined by the environment in which the axons have come into contact. The determinant factors encountered by the axons are transported via retrograde axonal flow to the nucleus where the signals are interpreted and the appropriate genes activated to manufacture the enzymes necessary to produce the specific neurotransmitters signalled for.
Environmental stimulus is conveyed to the nucleus of the neuron via specialised receptor systems on the membrane
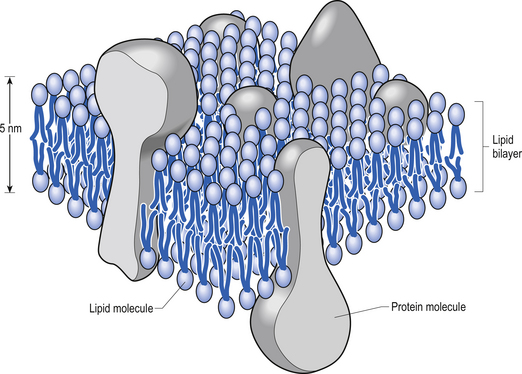
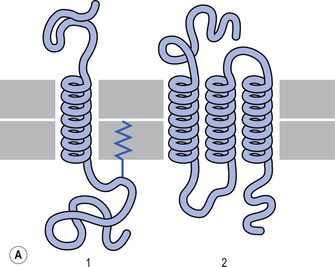
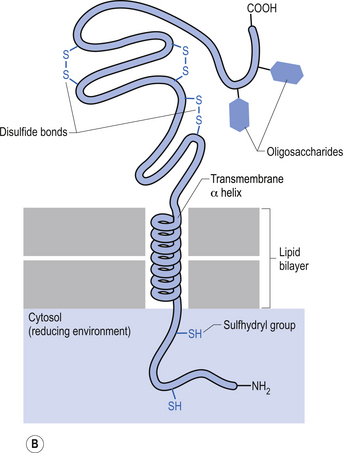
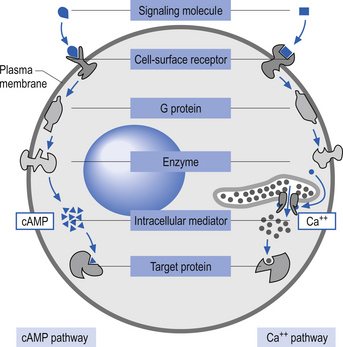
The plasma membrane of a neuron is essential for the survival of the neuron. It encloses the cell and maintains essential differences in composition and ion concentrations between the cytosol of the neuron and the external environment. The plasma membrane is essentially composed of proteins floating in a thin bilayered lipid structure held together by non-covalent interactions. This unique structure forms a relatively impermeable barrier to the passage of most water-soluble compounds. Some of the proteins in the lipid bilayered structure act as structural support, while others act as receptors and transducers that relay information across the membrane about the neuron’s environment (Fig. 3.1). Proteins that span the membrane usually assume an α-helical structure as they pass through the lipid portions of the membrane. This configuration is the most thermodynamically stable, due to interactions with the polar peptide bonds of the polypeptide and the hydrophobic nature of the lipid environment. The transmembrane portion of the protein can pass through the membrane only once, resulting in a single-pass transmembrane protein, or multiple times, resulting in the formation of a multipass transmembrane protein (Fig. 3.2). Multipass transmembrane proteins can form channels in the membrane that can be regulated by a variety of mechanisms (Alberts et al. 1994). Some channels are intimately associated with specialised proteins that act as receptors. The neuron has specific receptor proteins for a variety of chemical compounds known as transmitters.
All receptors for chemical transmitters have three things in common:
1. They are membrane-spanning proteins in which the external portion of the protein recognises and binds a specific neurotransmitter. Some common neurotransmitters include acetylcholine (ACh), norepinephrine (NE), epinephrine (E), serotonin or 5-hydroxytryptophan (5-HT), and dopamine (DA).
2. They carry out an effector function within the target cell. This function may include regulation of specific ion channels, release or activation of second messenger compounds, or modulation of activity levels of intracellular enzymes.
3. It is the receptor that determines the action of the transmitter based on the activity it produces inside the cell. This is an important point to remember. Many neurotransmitters are classified as excitatory or inhibitory to certain cellular functions; however, it is the internal wiring of the receptors that determine the response of a transmitter. For example, acetylcholine has an inhibitory or slowing effect on the heart rate but an excitatory effect on skeletal muscle.
Receptors can be either directly or indirectly linked to ion channels
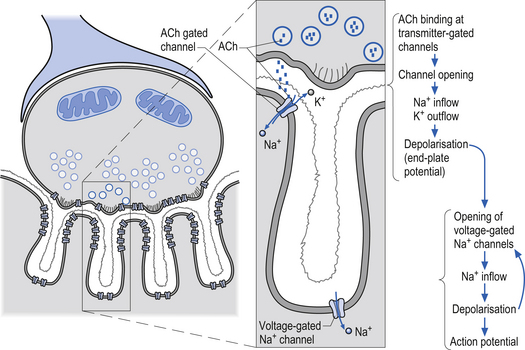
Receptors that gate ion channels directly are called inotropic receptors. Upon binding of a transmitter, the receptor undergoes a conformational change that allows the ion channel to open. The receptor is part of the same molecular structure that composes the channel. The activation of inotropic receptors produces fast synaptic actions (milliseconds in duration), e.g. ACh receptor at the neuromuscular junction (Fig. 3.3).
Receptors that indirectly gate ion channels are called metabotropic receptors. These types of receptors act through a special series of interlinked proteins called G-proteins. G-proteins are so named because of their ability to bind the guanine nucleotides, guanosine triphosphate (GTP), and guanosine diphosphate (GDP). Four major types of G-proteins are involved in transduction of signals produced by neurotransmitter binding, Gs, Gi/o, Gq, and G12, and multiple subtypes exist for each.
Activation of these types of receptors often activates a second messenger such as cyclic AMP (cAMP) or diacylglycerol in the cytoplasm of the neuron. Other prominent second messengers in brain include cyclic GMP (cGMP), calcium, phosphatidylinositol (PI), inositol triphosphate (IP3), arachidonic acid, and nitric oxide (NO) (Duman & Nestler 2000) (Fig. 3.3).
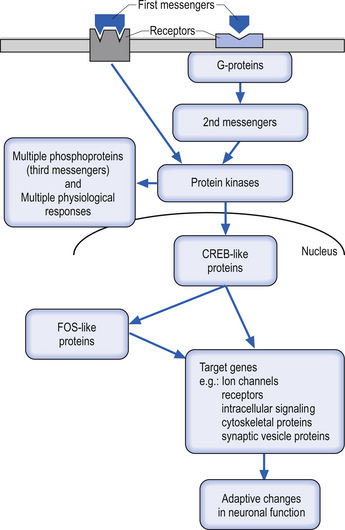
Many second messengers then act on a variety of intracellular kinases or enzymes to promote or inhibit cellular functions. Such intracellular processes can produce rapid changes in neuronal function such as changes in ionic conductance across the membrane. These second messenger processes can also produce short- to medium-term modulatory effects on neuronal function, such as regulation of the responsiveness of the neuron to the same or different neurotransmitters (transmitter modulation) via changes in receptor sensitivity. Relatively long-term modulatory effects on neuronal function, including changes achieved through the regulation of gene expression, can also be regulated by the actions of second messengers on other intracellular components called third messengers. Such changes can last seconds to minutes and include altered synthesis of receptors, ion channels, and other cellular proteins, or for much longer periods ultimately resulting in forms of learning and memory (Fig. 3.4).
Free levels of intracellular Ca++ can act as a potent second messenger in a number of different pathways
Under normal conditions the intercellular concentration of free Ca++ ions is under strict regulatory control. Receptor activation can result in increases in free intercellular Ca++ ion levels in two different ways involving two types of mechanisms that operate to different extents in different cell types. Neurotransmitter receptor activation can alter the flux of extracellular Ca++ ions into neurons or can regulate release of Ca++ ions from intracellular stores. Once released, Ca++ ions can exert multiple actions on neuronal function via intracellular regulatory proteins. Receptors can directly regulate the conductance of specific voltage-gated Ca++ channels via coupling with G-proteins. In addition, activation of other second messenger systems can alter Ca++ channel conductance. Depolarisation of a neuron by any means will activate voltage-gated Ca++ channels, which will lead to the flux of Ca++ into the cells. Finally, extracellular Ca++ can pass through some ligand-gated channels, such as the nicotinic cholinergic and glutamate N-methyl-D-aspartate (NMDA) receptors (Duman & Nestler 2000).
The structure of chromatin can regulate gene transcription induced by receptor stimulation
In human neurons, DNA is contained in the nucleus of the cell. The nucleus is also the site of DNA replication and transcription. Chromatin is formed from subunits of nucleosomes, which are chromosomes intimately associated with histone proteins. A chromosome is composed of extremely long molecules of DNA. The DNA not actively involved in transcription processes is stored in supercoiled structures that drastically reduce the space requirements in the nucleus for the storage of DNA. Chromatin is not only structurally important in this storage role but also acts in the regulation of gene expression by inhibiting transcription factors access to binding sites on DNA. Activation of a gene requires that the chromatin or nucleosomal structure be modified to allow the binding of regulatory proteins to the appropriate subset of genes. This is accomplished by a specialised group of proteins referred to as activator proteins that remodel the chromatin and expose core promoter sites on the appropriate genes. This permits the binding of yet another complex of proteins called general transcription factors to the core promoter site on the DNA. This complex of general transcription factors can then recruit and bind with RNA polymerase to enter the transcription initiation phase of the replication process (Workman & Kingston 1998).
In humans, three different types of RNA polymerases, I, II, and III, are involved with the transcription of different types of DNA. Polymerase I is involved in the transcription of ribosomal RNAs (rRNA). Polymerase II is involved in the transcription of messenger RNA (mRNA) and another subset of RNAs known as snRNA, which are involved in splicing RNA segments. Polymerase III is involved in the transcription of a number of smaller RNA types including transfer RNA (tRNA) (Struhl 1999) (Fig. 3.5).
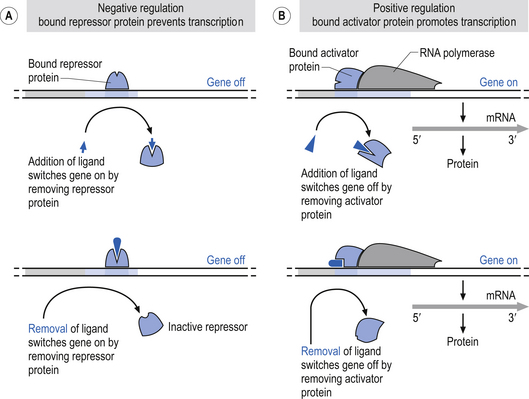
Figure 3.5 This illustrates the negative and positive regulatory processes modulating transcription polymerase.
In humans, the expression of highly complex genes requires that additional transcriptional activators are necessary for the transcriptional process to function. These additional transcriptional activation complexes are referred to as functional regulatory elements or transcription factors that bind to specialised sites within the structure of the gene. These functional transcription factors determine the unique pattern of expression for each gene, both in the normal course of development and in response to environmental stimuli. Aspects of gene expression under the control of various transcription factors include the cell type in which the gene is expressed, the time during development when the gene is expressed, and the level to which it will be expressed (Collingwood et al. 1999).
Several families of transcription factors as well as several modes of activation or inhibition of these factors have been identified. For example, the cAMP response element binding protein (CREB) family of transcription factors activate transcription of genes to which they are linked when they are phosphorylated by cAMP-dependent protein kinase (protein kinase A). Protein kinase A is activated in the presence of cAMP (De Cesare & Sassone-Corsi 2000).
The CREB family of transcription factors can also be activated by other second messengers such as Ca++ bound by calmodulin that can activate a variety of protein kinases upon entering the nucleus of the neuron. These kinases can in turn phosphorylate CREB, resulting in the activation of transcription of the specific CREB-linked gene (Nestler & Hyman 2002).
Dissemination of the receptor stimulus throughout the neuron
The environmental stimulus – whether it be a growth hormone, a neurotransmitter, or a hormone – must get its signal from the receptors on the neuron cell membrane to the transcriptional controlling factors in the nucleus in order for production of the necessary proteins that it calls for.
Gene expression
• Gene regulatory proteins or transcription factors are activated in the cytosol by second messengers and secondary effectors. This follows the activation of membrane receptors by neurotransmitters and peptides, and changes to intracellular ion concentrations.
• Immediate early gene responses can occur within nanoseconds of cell activation and represent a learned response of the cell. They can lead to protein synthesis or activation, and can also trigger transcription within the nucleus of the cell.
• DNA code is transcribed to produce a single mRNA molecule, which then leaves the nucleus via nuclear pores.
• mRNA passes this code to a ribosome, with the help of rRNA, which reads the base-sequence of the mRNA and translates the code into an amino acid sequence.
• tRNA transfers the appropriate amino acids in the cytosol to their designated site in the sequence being formed by the rRNA.
Some signalling molecules such as hydrophobic hormones (glucocorticoids, oestrogen, and testosterone) can gain direct access to the nuclear apparatus by their lipid-soluble chemical structure that allows them the ability to transverse the highly hydrophobic bilayered lipid plasma membrane dependent mainly on their concentration gradients. These hormones then bind with intracellular hormone receptors that carry them through the cytoplasm and across the nuclear membrane where they bind and alter the conformation of transcriptional factors (Evans & Arriza 1989; Lin et al. 1998).
Other signalling molecules such as Ca++ ions gain access through specific ion channels present in the neuron plasma membrane (Hess 1990).
Protein hormones, growth factors, peptide neuromodulators, and neurotransmitters must act on their transcription protein targets indirectly by either inducing a change in a transmembrane protein channel related to their receptor proteins (Lester & Jahr 1990) or by inducing a change in linked intramembrane proteins. Changes in these linked intramembranous proteins called G-proteins eventually result in the release of intracellular ions or the generation of intracellular second messenger such as cAMP, diacylglycerol, and inositol triphosphate (Berridge & Irvine 1989; Huang 1989), which then activate directly or through other intermediates the transcription factors in the nucleus such as CREB (Fig. 3.6).
• Temporally and spatially graded rises in second messenger levels (Berridge & Irvine 1989);
• Recruitment of various combinations of second messengers after a single stimulus (Nishizuka 1988); or
• Regional variations in the intracellular targets on which the second messengers act (Nishizuka 1988; Huang 1989).
Prolonged activity of second messengers can lead to a variety of damaging effects on a neuron, ranging from inappropriate activity to transformation and cell death. This has resulted in the formation of a variety of regulatory mechanisms in the neuron to control the concentration and temporal activity of second messengers. Most commonly, the second messengers are sequestered by intracellular proteins, or degraded by intracellular enzymes within milliseconds of release in the neuron (Boekhoff et al. 1990). Because of their size and short span of activity, second messengers are limited in their ability to act over the long term and limited in their specificity for precise and effective modulation of protein transcription. These shortcomings have led to the search for a third messenger system within the neuron that can function very specifically and over long periods to modulate gene expression.
Extracellular signals result in activation of a third messenger system coded for by immediate early genes
Third messengers are groups of nuclear proteins known as translational factors that are induced by a variety of extracellular signals. These proteins bind to specific nucleotide sequences in the promoter and enhancer regions of genes (Mitchell & Tjian 1989). The third messengers are coded by immediate early genes (IEGs), also referred to as primary response genes or competence genes. The proteins encoded for by immediate early genes in concert with other transcriptional factors exert powerful excitatory or inhibitory effects on the initiation of RNA synthesis (Pleasure 1992). Many of the IEGs were initially recognised because they are the normal nuclear homologues of transforming retroviral oncogenes, which are the class of gene released by viruses immediately upon entering a host cell.
The most fully studied IEG is c-fos. The c-fos gene has three binding sites for CREB and is activated by neurotransmitters or other stimuli that stimulate the production of cAMP in the neuron (Ahn et al. 1998). Changes in the tertiary structure of c-fos gene are detectable within one minute after cell stimulation, first appearing in regulatory regions of the gene and then propagating to decoding regions of the gene. The half-life of the c-fos gene and the protein it codes for are very short, in the range of 20–30 minutes. This time frame of activation is much shorter than other proteins of a structural or enzymatic nature but is many orders of magnitude greater than the half-life of the second messengers.
How are neurons stimulated?
• A receptor converts an impulse of energy into an electrical impulse that travels along the nerve.
• They are derived from mesoderm – therefore, they do not need to be activated in order to stay alive, as is the case with nerve cells.
• The receptor potential is the first representation of an internal or external event to be coded in the nervous system.
Features of receptor potentials
• It is graded (depending on intensity and duration) and unpropagated (amplitude decreases progressively) and needs to be amplified; thus the initial response of a receptor is its greatest.
• Degradation occurs quickly due to Na+ being drawn out, but the receptor potential may trigger an action potential due to the higher concentration of voltage-gated sodium channels where the axon meets the receptor.
• There is a one-to-one relationship between the environmental stimulus and the receptor potential.
Fine-tuning of the effects of third messengers is accomplished through a complex network of controls. Since there are now over a hundred IEGs and corresponding proteins composing third messengers, a very complex matrix of interactivity which would allow complicated but minor variances in linear and temporal combinations of third messengers for various functions can be developed (Pettersson & Schaffner 1990; Ptashne & Gann 1990) (Table 3.1). For example, when fos binds to DNA as a heterodimer complex with another third messenger protein called jun the transcription of the target protein – usually tyrosine hydroxylase, neurotensin, neuromedin, or a proenchephalin – is dramatically increased (Gizang-Ginsberg & Ziff 1990; Kislauskis & Dobner 1990). However when fos binds to DNA on its own it inhibits the transcription of c-fos, its coding gene (Gius et al. 1990).
| Class | Proto-oncogene nomenclature | Homologue |
|---|---|---|
| Receptor ligand |
Reproduced with permission from Discussions in Neuroscience, 7(4) (August 1991), Elsevier Science Publishers BV.
The fos family of genes may act as a molecular switch within the neuron
Under resting conditions the concentration of c-fos mRNA and protein in the neuron are extremely low, but c-fos expression can be dramatically increased by a variety of stimuli (Correa-Lacarcel et al. 2000). For example, experimental induction of a grand mal seizure causes marked increases in c-fos mRNA within the brain within 30 minutes and induces the formation of c-fos protein within 2 hours (Sonnenberg et al. 1989). The fos-like proteins are highly unstable and return to normal values within 4–6 hours. Administration of other substances such as cocaine or amphetamine causes a similar pattern of expression in the striatum (Graybiel et al. 1990; Hope et al. 1994). With repeated activation, the c-fos family of genes become refractory to the stimulus, and other isoforms of the fos proteins which express very long half-lives in brain tissue are expressed and accumulate in specific neurons in response to repeated stimulus (Pennypacker et al. 1995; Chen et al. 1997). The accumulation of these proteins remains in the neurons long after the stimulus has ceased. The prolonged presence of the fos-like proteins may act as a molecular switch inside the cell, shutting off or modulating responses to repeated stimulus. The true functional significance of the sustained presence of these fos-like proteins in neurons remains unknown but may have a mediating effect on the development of various striatal-based movement disorders (Kelz & Nestler 2000).
We will now look at a variety of receptors and their respective neurotransmitters.
Acetylcholine (cholinergic) receptors
Acetylcholine is essential for the communication between nerve and muscle at the neuromuscular junction. ACh is also involved in direct neurotransmission in the autonomic ganglia and is also active in cortical processing, arousal, and attention activity in the brain (Karczmar 1993) (Fig. 3.7).
Cholinergic transmission can occur through G-protein coupled mechanisms via muscarinic receptors or through inotropic nicotinic receptors. The activity of ACh is terminated by the enzyme acetylcholinesterase, which is located in the synaptic clefts of cholinergic neurons. To date, seventeen different subtypes of nicotinic receptors and five different subtypes of muscarinic receptors have been identified (Nadler et al. 1999; Picciotto et al. 2000).
Cholinergic, nicotinic receptors are also present at the neuromuscular junctions of skeletal muscle.
Adrenergic receptors
Alpha-2 receptors have been demonstrated in wide areas of the central nervous system (CNS) including multiple nuclei of the brainstem and pons, the midbrain, the hypothalamus, the septal region, amygdala, olfactory system, hippocampus, cerebral cortex, spinal cord, and cerebellum and in many neuroendocrine cells (Wang et al. 1996). Alpha-2 receptors are mediated by the GTP-binding proteins subfamily and affect three different routes of inhibitory activation:
• Inhibition of adenyl cyclase and thereby inhibition of the production of cAMP;
• Suppression of voltage-activated calcium channels, thus reducing the flow of extracellular Ca++ into target cells; and
• Increased conductance of K+ ions through the membranes of target cells.
All three of these activities (inhibition of adenyl cyclase, suppression of voltage-sensitive calcium channels, and stimulation of potassium channels) can contribute to the inhibition of the target cell, and to reduction of neurotransmitter release in neurons or hormone release in neuroendocrine cells (Langer 1974).
The beta-adrenergic receptors have three well-recognised subclasses: beta-1, beta-2, and beta-3 receptors. All three subtypes are coupled to adenyl cyclase activation via stimulatory G-proteins (Barnes 1995).
Beta-1 and -2 receptors have been demonstrated in the lungs, including airway smooth muscle, epithelium, cholinergic and sensory nerves, submucosal glands, and pulmonary vessels, and are also found in the heart; here beta-1 receptors are predominantly in the myocytes, while the beta-2 receptors are on innervating neurons. Beta-2 receptors are also present in saphenous vein, mast cells, macrophages, eosinophils, and T lymphocytes (Ruffolo et al. 1995). Beta-3 receptors are primarily expressed in brown and white adipose tissue, although some studies have also reported the presence of beta-3 receptors in oesophagus, stomach, ileum, gallbladder, colon, skeletal muscle, liver, and cardiovascular system (Krief et al. 1993; Berlan et al. 1995).
Glutamate receptors
The transmitter L-glutamate or L-glutamic acid (Glu) is the major excitatory transmitter in the brain and spinal cord (Hollmann & Heinemann 1994).
The role of Glu in the function of the nervous system is much more diverse and complex than a simple excitatory neurotransmitter. It also plays a major role in brain development, neuronal migration, differentiation, and axon development and maintenance (Komuro & Rakic 1993; Wilson & Keith 1998). In the mature nervous system, Glu is essential in the processes involving stimulus-dependent modifications of synapses necessary for neural plasticity to occur.
NMDA receptors vs non-NMDA receptors – clinical considerations
• They are unique receptors as they depend on membrane potential and activation by glutamate and co-factor glycine. Cell needs to be depolarised first (≈20 mV) in order for magnesium plug to be expelled, such as:
• Activation of NMDA receptors results in massive influx of calcium, which results in biochemical cascade of events leading to IEG response. NMDA receptors can promote changes in protein production or phosphorylation of intracellular domain of membrane channels.
• NMDA receptor activity promotes synaptogenesis and survivability of the neuron.
Persistent or overwhelming stimulation of glutamate receptors can result in neuronal degeneration or, in some circumstances, neuronal death by necrosis or apoptosis. This process is referred to as excitotoxicity and has been linked to the development of a range of disorders including Huntington’s disease, Alzheimer’s disease, amyotrophic lateral sclerosis, and stroke (Choi 1988; Ankarcrona et al. 1995; Olney et al. 1997).
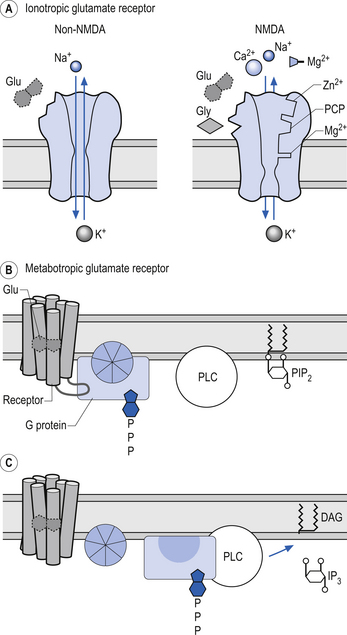
Because AMPA and kainate receptors are similar in structure and not voltage-dependent they are sometimes referred to as non-NMDA receptors (Fig. 3.8).
Glutamate NMDA receptors
The NMDA receptor is a complex receptor that has three exceptional qualities:
1. The receptor controls a gated channel permeable to Na, K, and Ca (non-NMDA receptors are not permeable to Ca).
2. The channel will only function if glycine is also present as a co-factor.
3. The function of the channel is dependent on a specific membrane voltage being reached as well as the presence of a chemical transmitter.
Glutamate NMDA receptor activation can modulate genetic expression in the neuron via Ca++-induced second messenger systems
When glutamate Kainate and AMPA receptors are activated by glutamate binding, the result is an influx of Na+ ions into the neuron, which depolarises the neuron, bringing its membrane potential towards threshold. Simultaneously, glutamate will also bind to the NMDA receptors on the neuron membrane. Recall that in order to activate NMDA receptors, which allow Ca++ to move into the neuron, the membrane potential must meet certain criteria. The membrane potential necessary to activate NMDA receptors is usually sufficient to bring the postsynaptic neuron to threshold so that an action potential is initiated. Thus glutamate-induced Ca++ influx is associated with action potential initiation in the postsynaptic neuron. Ca++ influx into the postsynaptic neuron results in the activation of a variety of second and third messenger systems that result in modulation of mRNA and protein production in the neuron (see below).
Glycine receptors
Glycine is a less common inhibitory transmitter that acts on inotropic channels that gate Cl. The inhibition that these receptors produce is due to the increased conductance of Cl ions. Since the concentration of Cl outside the cell is much greater than that inside the cell, Cl flows down its concentration gradient into the cell, making the inside more negative and hyperpolarising the cell. This results in a decreased probability that the neuron will produce an action potential per unit stimulus and is referred to as inhibition of the neuron.
Serotonin or 5-hydroxy-tryptophan (5-HT) receptors
Serotonin has been implicated in a staggering array of physiological and behavioural activities including affect, aggression, appetite, cognition, emesis, gastrointestinal function, perception, sensory function, sex, and sleep (Bloom & Kupfer 1995).
To date, five different subtypes of 5-HT receptor have been isolated. All members of the 5-HT type-1 receptor family tend to modulate inhibitory effects through either presynaptic or postsynaptic action. All members of the 5-HT type 2 subgroup modulate excitatory activity (Aghajanian & Sanders-Bush 2002).
Dense concentrations of 5-HT receptors are found in the dorsal raphe nucleus, hippocampus, and cerebral cortex. The facial nerve and other cranial motor nuclei also have high densities of 5-HT receptors (Chalmers & Watson 1991).
Dopamine receptors
There are five types of dopamine (DA) receptors that are classified into two major categories. The five receptor types are D1–D5. These receptors fall into two class categories, the D1-like receptor class and the D2-like receptor class (Spano et al. 1978; Su et al. 1996). The D1-like receptor class includes D1 and D5 receptor types. The D2-like receptor class includes D2, D3, and D4 receptor types (Sunahara et al. 1990; Sumiyoshi et al. 1995).
Dopamine receptors are present in most parts of the CNS but in particular they are found in three main projection systems: the nigrostriatal pathway, comprising the neurons of the substantia nigra pars compacta, which project to neurons of the neostriatum; the mesolimbic pathway comprising neurons of the ventral tegmental area of the mesencephalon, which project to widespread areas of the limbic system; and the mesocortical pathways, which involve neurons in the substantia nigra and the ventral tegmental areas of the mesencephalon that project to the prefrontal cortical areas of the brain (Bjorklund & Lindvall 1984; Blumenfeld 2002).
Attempts to understand the actions of the different DA receptors are often stymied by the complex array of activities that these receptors can produce. DA receptors have been shown to interact via both inotropic and G-protein coupled mechanisms. DA has also been shown to have a modulatory affect on other transmitters in the region of its activity and to alter the actions of groups of neurons through modulation of gap junction activities between the neurons (Grace 2002). DA can also regulate its own levels of interaction by activating autoreceptors sensitive to local DA concentrations. These autoreceptors have been found on the soma of dopaminergic neurons as well as at the dopaminergic nerve terminal synapses.
Several studies have shown the presence of dopaminergic neurons in the substantia nigral and ventral tegmental areas of the mesencephalon. The majority of these neurons seem to exhibit spontaneous action potential generation driven by an endogenous pacemaker conductance activity (Grace & Bunney 1984). The rate of this pacemaker generation is normally closely regulated by feedback from autoreceptors located on the soma and synaptic areas of the neurons (Harden & Grace 1995).
The complexity of the interactions involving dopaminergic receptors can be illustrated in the following example. D1 receptor activation in dorsal striatum neurons results in further inhibition of previously hyperpolarised neurons. However, with repeated stimulation of D1 receptors in previously hyperpolarised neurons excitation can occur. When D1 and D2 receptors are stimulated simultaneously the result is a synergistic inhibition of the neuron. However, the D1-mediated inhibition previously described can be reversed by subsequent stimulation of D2 receptors on the neuron (Cepeda et al. 1995; Hernandez-Lopez et al. 1997; Onn et al. 2000).
Receptor modulation of neuron bioenergetic processes require ATP for energy
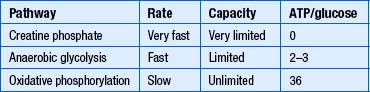
Bioenergetics describes the transfer and utilisation of energy in biological systems. Essential processes like transferring ions across a membrane against their concentration gradients to maintain a membrane potential require energy to operate. In the neuron, most energy-requiring processes are made possible by either direct or indirect coupling with an energy-releasing mechanism involving the hydrolysis of adenosine triphosphate (ATP). The ATP molecule is composed of an adenosine base segment to which three phosphate groups attach. The two terminal phosphate groups release a relatively high amount of energy (7.3 kcal/mol) when they are chemically broken down; these are referred to as high-energy phosphate bonds. The monophosphate bond adjacent to the adenosine base releases about 4.0 kcal/mol when broken down and is referred to as a low-energy phosphate bond. Other compounds such as phosphoenolpyruvate and phosphocreatine contain phosphate bonds which release energy approaching 10 kcal/mol when they are broken down and are referred to as very-high-energy phosphate bonds (Champe & Harvey 1994). These compounds can convert ADP to ATP over short time periods in situations when ATP demands are higher than ATP production. Glucose-6-phosphate and glycerol 3-phosphate both have phosphate bonds that release about 4.0 kcal/mol and are referred to as low-energy phosphate bonds. Thus, ATP is placed in the middle ground between very-high- and low-energy phosphate bonds and acts as a middleman in the transfer of energy between molecules that regulate processes.
Energy transfer in the neuron
• Energy in the electrons of the free hydrogen atoms is carried by NAD and FAD into the electron transport chain – a series of electron carrier molecules on the inner mitochondrial membrane lining the cristae. The electrons fall to successively lower energy levels along the chain as they synthesise more ATP via ATP synthetase.
• Complexes I and II collect electrons from the catabolism of fats, proteins, and carbohydrates and transfer them to ubiquinone (CoQ10). The electrons are then transferred successively to complexes III and IV and to oxygen, which is the final electron acceptor. Three sites along the electron transport chain (complexes I, III, and IV) use the energy released from the transfer of electrons to transport hydrogen ions across the inner mitochondrial membrane to the intermembrane space – thus driving a proton gradient.
• The [H+] is higher in the intermembrane space so it travels back to the matrix via channels that contain ATP synthetase, which is activated by H+ flow.
• ADP + Pi (ATP synthetase) → ATP (32 more molecules per glucose molecule).
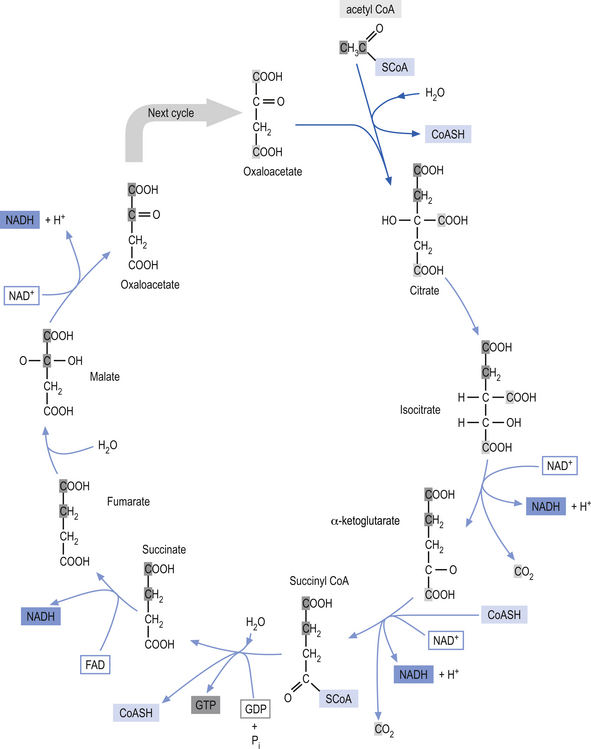
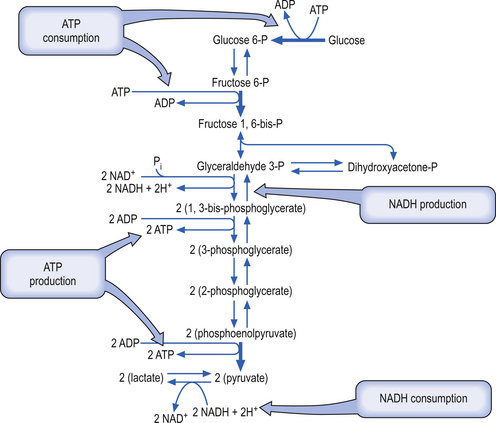
Glycolysis (Embden-Meyerhof pathway) is the metabolism of glucose to pyruvate and lactate. This process results in the net production of only 2 mol of ATP/mol of glucose (Fig. 3.9). On the other hand, pyruvate can pass into the tricarboxylic acid cycle (Krebs cycle) in the mitochondria and via the oxidative phosphorylation cascade produce 30 mol of ATP/mol of glucose (see Fig. 3.3). The energetic benefit of utilising the oxidative phosphorylation route over the glycolytic route is obvious from an energetic perspective (Magistretti et al. 2000).
The glycolytic route may not always be utilised for the production of ATP in neurons because of saturation of enzymes in the Krebs pathway or the oxidative phosphorylation cascades pathways in the mitochondria. Several studies have shown that the ATP production capability of the neuron operating at a basal rate is operating at near maximal capacity. When the neuron undergoes activation it needs to utilise the glycolytic pathway for ATP production, thus producing lactate, which converts to lactic acid under certain conditions (Fox & Raichle 1986; Van den Berg 1986; Fox et al. 1988).
Metabolic demands of the brain require glucose as a substrate for ATP synthesis
The brain, which represents about 2% of the total body weight in humans, consumes 15% of the cardiac output blood flow, 20% of the total body oxygen consumption, and 25% of total body glucose utilisation (Kety & Schmidt 1948). Glucose utilisation calculations need to take into account the fact that glucose can have metabolic fates other than that of ATP production. Glucose can produce metabolic intermediates such as lactate and pyruvate which, when released, do not necessarily enter the tricarboxylic acid cycle but can be removed by the circulation. Glucose can also be incorporated into lipids, proteins, and glycogen and can also be utilised in the formation of a variety of neurotransmitters including GABA, glutamate, and acetylcholine. It is estimated that about 17% of the glucose in neurons is utilised in metabolic processes other than that of ATP production (Magistretti et al. 2000).
Numerous studies have tried to identify molecules other than glucose that could substitute for glucose in brain energy metabolic processes. To date, no other physiologically available substrate has been identified that can substitute for glucose under normal basal conditions. Under a certain set of conditions such as starvation, diabetes, or in breast-fed neonates, acetoacetate and D-3-hydroxybutyrate can be used by the brain as a metabolic substitute for glucose (Owen et al. 1967).
Other cells such as vascular endothelial cells and astrocytes also participate in neural activation processes
Brain metabolism studies in the past have assumed that energy metabolism at the cellular level represented predominantly neuronal activity. However, it is now clear that other types of cells such as neuroglia and vascular endothelial cells not only consume energy but also play a part in the neural activation process and in maintenance of neuron function. In studies spanning a variety of species, the ratio of non-neural to neural substrate is about 50% (Kimelberg & Norenberg 1989). In addition, there is very good evidence to suggest that the astrocyte to neuron ratio increases with increasing brain size.
Two well-established functions of astrocytes include the maintenance of extracellular K+ ion levels within a narrow range and to ensure the reuptake of neurotransmitters. Activation of neurons results in increases of extracellular K+ ion concentrations and increases in concentrations of the synaptic-specific neurotransmitters released by the neuron. For example, at excitatory synapses where glutamate is the activating neurotransmitter, it not only depolarises the postsynaptic neuron but also stimulates the uptake of K+ ions into the surrounding astrocytes (Barres 1991).
Recent studies have also shown that the metabolic activity of astrocytes can be mediated by norepinephrine and other vasoactive substances and that activation of the locus coeruleus in the brainstem prior to activation of neurons may indicate that the metabolic priming of astrocytes is preset prior to neuron activation by the nervous system and is not solely dependent on the generation of local metabolites for activation (Magistretti et al. 1981, 1993; Magistretti & Morrison 1988).
Clinical signs and symptoms of altered brain metabolism can be demonstrated with positron emission tomography (PET) studies
PET studies of patients with Huntington’s disease without hyperactive behaviour have shown normal frontal lobe metabolism in the presence of decreased caudate and putamen metabolism (Kuhl et al. 1982; Young et al. 1986).
PET scans in patients with Parkinson’s disease have shown decreased cortical glucose consumption in the frontal cortex in conjunction with decreased D2 receptor uptake ratios in the nigrostriatal (substantia nigra) regions (Brooks 1984).
Similar findings have been found in children with ADHD or childhood hyperkinetic disorder, where researchers found that after receiving Ritalin, previously normal subjects showed decreased activity in the basal ganglia, and those previously diagnosed with ADHD (originally decreased basal ganglionic activity compared to normal levels) showed an increased activity of basal ganglial areas (Young et al. 1986; Rogers 1998).
Mitochondrial activation and interactions in the neuron
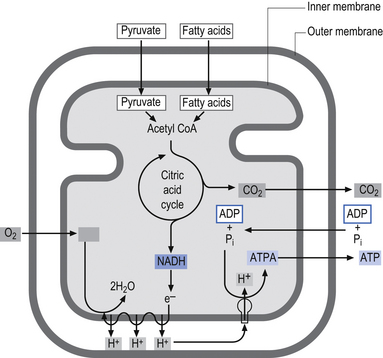
The inner membrane is impermeable to virtually all metabolites and small ions contained in the mitochondria. The inner membrane contains specialised proteins that carry out three main functions including oxidative reactions of the respiratory chain, the conversion of ADP to ATP (ATP synthase), and the transport of specific metabolites and ions in and out of the mitochondrial matrix. The inner membrane space contains several enzymes required for the passage of ATP out of the matrix space and into the cytoplasm of the host cell (Alberts et al. 1994) (Fig. 3.10).
Mitochondrial oxidative phosphorylation (OxPhos) disorders
The respiratory chain is composed of five multienzyme complexes, which include flavin and quinoid compounds, transition metals such as iron–sulphur clusters, hemes, and protein-bound copper compounds. The respiratory chain can be grouped into five complexes that in addition to two small carrier molecules, coenzyme Q and cytochrome c, can be grouped into the following complexes (Mendell & Griggs 1994):
• Complex I—contains NADH and the coenzyme Q oxidoreductase;
• Complex II—contains succinate and the coenzyme Q oxidoreductase;
• Complex III—contains coenzyme Q and cytochrome c oxidoreductase;
Complexes I and II collect electrons from the catabolism of fat, protein, and carbohydrates and transfer these electrons to ubiquinone (CoQ10), and then pass them on through to complexes III and IV before the electrons react with oxygen, which is the final electron receptor in the pathway (Smeitink & Van den Heuvel 1999).
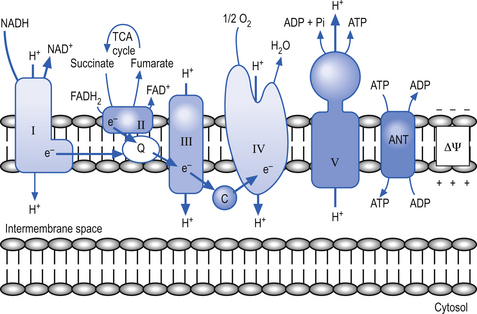
Complexes I, III, and IV use the energy from electron transfer to pump protons across the inner mitochondrial membrane, thus setting up a proton gradient. Complex V then uses the energy generated by the proton gradient to form ATP from ADP and inorganic phosphate (Pi) (Nelson & Cox 2000) (Fig. 3.11). During this process, about 90–95% of the oxygen delivered to the neuron is reduced to H2O; however, about 1–2% is converted to oxygen radicals by the direct transfer of reduced quinoids and flavins. This activity produces superoxide radicals at the rate of 107 molecules per mitochondria per day. Superoxide radicals are part of a chemical family called reactive oxygen species or free radicals. They are extremely reactive molecules because they contain an oxygen molecule with an unpaired electron (Del Maestro 1980). Although production of free radicals can occur during specialised cellular process such as in lysozyme production in neutrophils the vast majority of all free radical production occurs in the mitochondria.
Oxygen free radicals can also bind to iron–sulphur-containing proteins, releasing ferrous iron moieties that react with hydrogen peroxides to form an extremely reactive and damaging hydroxyl radical that can overwhelm the neuron’s normal biochemical supplies of antioxidants and result in oxidative stress (Jacobson 1996).
A unique characteristic of the genetics of the respiratory chain enzyme complexes is that the genes that code for each enzyme complex are composed of some from the mitochondrial DNA (mtDNA) and some from the host neuron DNA (Hatefi 1985; Birky 2001). Another fact that complicates the genetics of mitochondria is that the vast majority of the mtDNA comes directly from the mother. This is because very little mtDNA is carried or transferred by the sperm at fertilisation (Giles et al 1980; Sutovsky et al. 1999).
Particular mutations in mtDNA are responsible for specific patient presentations
MtDNA mutations cause a range of movement disorders including ataxias, dystonias, myoclonic epilepsy with lactic acidosis and stroke-like episodes (MELAS), and Kearn’s-Sayer (KS) syndrome, and more and more evidence points to their involvement in Parkinson’s disease. MELAS and KS syndrome will be briefly discussed here, while ataxias, dystonias, and Parkinson’s disease are covered in Chapter 11.
Conservative treatment of OxPhos disorders
CoQ10 supplementation given at 2–4 mg/kg/day has been effective for improving symptoms in patients with OxPhos disorders. The same treatment has been effective for increasing the mitochondrial respiration rate, which declines with age naturally by approximately 1% per year after the age of 40 (Bresolin et al. 1988; Ihara et al. 1989).
Apoptosis is a controlled, preprogrammed, process of neuron death
Apoptosis, which differs from necrotic cell death, involves a complex set of specific preprogrammed activities that result in the death of the neuron. This type of activity is actually an important part of normal embryological development of the nervous system which has been linked to the absence or lack of appropriate concentrations of nerve growth factors. The involvement of the mitochondria in apoptosis is well documented (Green & Reed 1998; Desagher & Martinou 2000). When activated by cellular damage or other proapoptotic signals, apoptogenic molecules that normally remain dormant in the membrane of the mitochondria become activated. These molecules then activate aspartate-specific cysteine protease (caspase), a major effector in apoptosis in neurons (Schulz et al. 1999). The caspase pathway is also activated by other cellular insults such as DNA damage and anoxia. The processes involved in apoptosis result in neuron shrinkage, condensation of chromatin, cellular fragmentation, and eventual phagocytosis of cellular remnants.
When the FOF of presynaptic neurons is decreased, as would occur because of subluxation, the following events may take place:
• ↓ CIEGr (cellular Immediate Early Gene responses)
• ↓ Cellular respiration (via mitochondrial electron transport chain)
• ↑ Resting membrane potential (RMP)
• Further inhibition of cellular respiration (electron transport chain) in the mitochondria
FOF—Frequency of firing (action potential generation)
Transneural degeneration
• Decreases in cellular immediate early gene responses (CIEGr);
• Decreases in protein production;
• Decreases in cellular respiration (via mitochondrial electron transport chain);
• Increases in resting membrane potential (RMP) in initial stages;
• Hyperpolarisation of membrane potential in the late stages of degeneration;
• Increased free radical formation; and
• Further inhibition of cellular respiration (electron transport chain) in the mitochondria.
Changes in the FOF and fuel delivery can have a deleterious effect on the central integrated state (CIS) of a neuronal pool. The CIS determines the integrity of a neuronal pool and its associated functions and dictates the presence of neurological signs and symptoms.
Case 3.1
3.1.1
The reduced activation of the movement receptors in the joints of his arm will result in a decreased afferent input or reduced synaptic activity of the dorsal root ganglion cells responsible for detection of proprioception in his arm. The number of synapses firing at any one time (spatial summation) and the frequency of firing of any one synapse (temporal summation) are integral in determining the central integrative state of a neuron at any given moment. When a neuron is exposed to a decrease in synaptic activity over a period of time it may undergo a change in central integrative state that results in transneuron degeneration. Once normal activity is attempted in the arm the neurons that have been subject to a lack of synaptic activity, low glucose supplies, low oxygen supplies, decreased adenosine triphosphate (ATP) supplies, etc., may not be able to respond to a sudden synaptic barrage of restored movement in the appropriate manner and the overfunctional integrity of the system becomes less than optimal. In neurons that have been exposed to a decreased frequency of synaptic activation a number of responses can be found in the neurons including:
• Decreases in cellular immediate early gene responses;
• Decreases in protein production;
• Decreases in cellular respiration (via mitochondrial electron transport chain);
• Increases in resting membrane potential in initial stages;
• Hyperpolarisation of membrane potential in the late stages of degeneration;
• Increased free radical formation; and
• Further inhibition of cellular respiration (electron transport chain) in the mitochondria.
3.1.3
A variety of things could be done to prevent any serious damage from occurring:
1. The arm could be gently moved several times per day to stimulate proprioceptors.
2. Transepithelial electrical nerve stimulation (TENS) could be applied on a daily basis to maintain action potential frequency in the system.
3. Appropriate rehabilitation needs to be carried out before return to full activity.
Case 3.2
3.2.2
Third messengers are groups of nuclear proteins known as translational factors induced by a variety of extracellular signals. These proteins bind to specific nucleotide sequences in the promoter and enhancer regions of genes. Fine-tuning of the effects of third messengers is accomplished through a complex network of controls. Since there are now over a hundred IEGs and corresponding proteins composing third messengers a very complex matrix of interactivity which would allow complicated but minor variances in linear and temporal combinations of third messengers for various functions can be developed. Activation of a gene requires that the chromatin or nucleosomal structure be modified to allow the binding of regulatory proteins to the appropriate subset of genes. This is accomplished by a specialised group of proteins referred to as activator proteins that remodel the chromatin and expose core promoter sites on the appropriate genes. This permits the binding of yet another complex of proteins called general transcription factors to the core promoter site on the DNA. This complex of general transcription factors can then recruit and bind with RNA polymerase to enter the transcription initiation phase of the replication process. Several families of transcription factors have been identified as well as several modes of activation or inhibition of these factors. For example, the cAMP response element binding protein (CREB) family of transcription factors activate transcription of genes to which they are linked when they are phosphorylated by cAMP-dependent protein kinase (protein kinase A). Protein kinase A is activated in the presence of cAMP.
References
Barres B.A. New roles for glia. J. Neurosci.. 1991;11:3685-3694.
Berridge M.J., Irvine R.F. Inositol phosphates and cell signalling. Nature. 1989;341:197-205.
Black M.M., Baas P.W. The basis of polarity in neurons. Trends Neurosci.. 1989;12:211-214.
Brooks V.B. The Neural Basis for Motor Control. Oxford: Oxford University Press, 1984.
Choi D.W. Glutamate neurotoxicity and diseases of the nervous system. Neuron. 1988;1:623-634.
Green D.R., Reed J.C. Mitochondria and apoptosis. Science. 1998;281:1309-1312.
Hess P. Calcium channels in vertebrate cells. Annu. Rev. Biochem.. 1990;13:337-356.
Hollmann M., Heinemann S. Cloned glutamate receptors. Annu. Rev. Neurosci.. 1994;17:31-108.
Huang K.P. The mechanism of protein kinase C activation. Trends Neurosci.. 1989;12:425-432.
Kimelberg H.K., Norenberg M.D. Astrocytes. Sci. Am.. 1989;260:44-52.
Komuro H., Rakic P. Modulation of neuronal migration by NMDA receptors. Science. 1993;260:95-97.
Langer S.Z. Presynaptic regulation of catecholamine release. Br. J. Pharmacol.. 1974;60:481-497.
Nelson D.L., Cox M.M. Lelninger Principles of Biochemistry. New York: Worth, 2000.
Ptashne M., Gann A.A.F. Activators and targets. Nature. 1990;346:329-331.