Bacterial Genetics, Metabolism, and Structure
1. Describe the basic structure and organization of prokaryotic (bacterial) chromosomes, including number, relative size, and cellular location.
2. Outline the basic processes and essential components required for genetic information transfer in replication, transcription, translation, and regulatory mechanisms.
3. Define mutation, recombination, transduction, transformation, and conjugation.
4. Describe how genetic alterations and diversity provide a mechanism for evolution and survival of microorganisms.
5. Differentiate environmental oxygenation and final electron acceptors (aerobes, facultative anaerobes, and strict anaerobes) in the formation of energy.
6. Compare and contrast the key structural elements, cellular organization, and types of organisms classified as prokaryotic and eukaryotic.
7. State the functions and biologic significance of the following cellular structures: the outer membrane, cell wall, periplasmic space, cytoplasmic membrane, capsule, fimbriae, pili, flagella, nucleoid, and cytoplasm.
8. Differentiate the organization and chemical composition of the cell envelope for a gram-positive and a gram-negative bacterium.
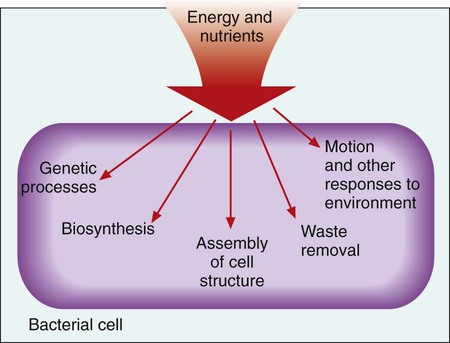
Microbial genetics, metabolism, and structure are the keys to microbial viability and survival. These processes involve numerous pathways that are widely varied, often complicated, and frequently interactive. Essentially, survival requires energy to fuel the synthesis of materials necessary to grow, propagate, and carry out all other metabolic processes (Figure 2-1). Although the goal of survival is the same for all organisms, the strategies microorganisms use to accomplish this vary substantially.
• The mechanism or mechanisms by which microorganisms cause disease
• Developing and implementing optimum techniques for microbial detection, cultivation, identification, and characterization
• Understanding antimicrobial action and resistance
• Developing and implementing tests for the detection of antimicrobial resistance
Bacterial Genetics
• The structure and organization of genetic material
• Replication and expression of genetic information
• The mechanisms by which genetic information is altered and exchanged among bacteria
Nucleic Acid Structure and Organization
Nucleotide Structure and Sequence
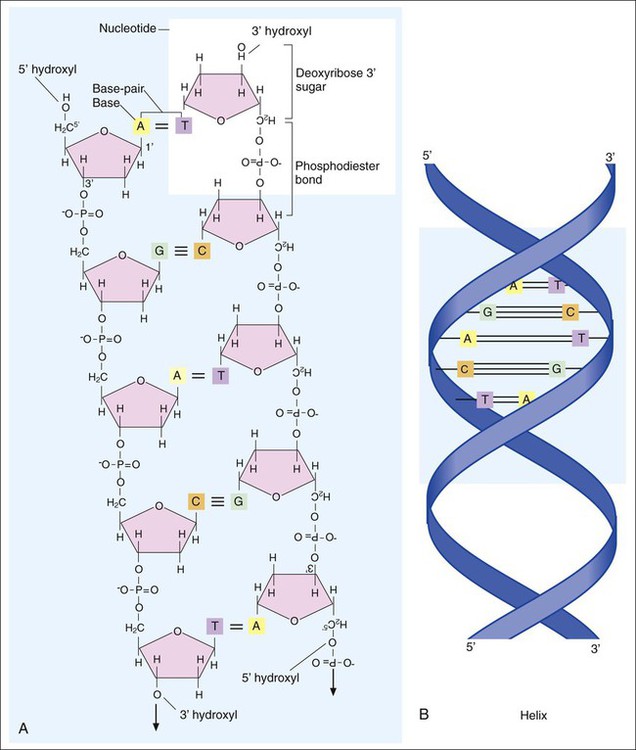
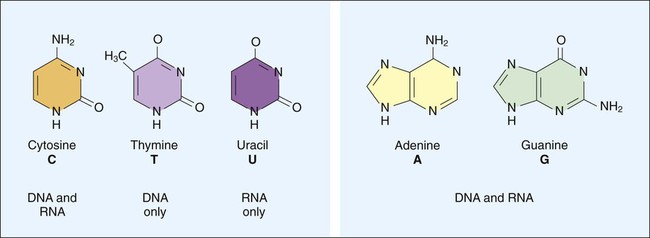
DNA consists of deoxyribose sugars connected by phosphodiester bonds (Figure 2-2, A). The bases that are covalently linked to each deoxyribose sugar are the key to the genetic code within the DNA molecule. The four bases include two purines, adenine (A) and guanine (G), and the two pyrimidines, cytosine (C) and thymine (T) (Figure 2-3). In RNA, uracil replaces thymine. The combined sugar, phosphate, and a base form a single unit referred to as a nucleotide (adenosine triphosphate [ATP], guanine triphosphate [GTP], cytosine triphosphate [CTP], and thymine triphosphate [TTP]). DNA and RNA are nucleotide polymers (i.e., chains or strands), and the order of bases along a DNA or RNA strand is known as the base sequence. This sequence provides the information that codes for the proteins that will be synthesized by microbial cells; that is, the sequence is the genetic code.
DNA Molecular Structure
The intact DNA molecule is composed of two nucleotide polymers. Each strand has a 5’ (prime) phosphate and a 3’ (prime) hydroxyl terminus (see Figure 2-2, A). The two strands run antiparallel, with the 5’ of one strand opposed to the 3’ terminal of the other. The strands are also complementary, because the adenine base of one strand always binds to the thymine base of the other strand by means of two hydrogen bonds. Similarly, the guanine base of one strand always binds to the cytosine base of the other strand by means of three hydrogen bonds. As a result of the molecular restrictions of these base pairings, along with the conformation of the sugar-phosphate backbones oriented in antiparallel fashion, DNA has the unique structural conformation often referred to as a “twisted ladder” or double helix (see Figure 2-2, B). Additionally, the dedicated base pairs provide the format essential for consistent replication and expression of the genetic code. In contrast to DNA, which carries the genetic code, RNA rarely exists as a double-stranded molecule. The three major types of RNA (messenger RNA [mRNA], transfer RNA [tRNA], and ribosomal RNA [rRNA]) play key roles in gene expression.
Genes and the Genetic Code
Certain genes are widely distributed among various organisms while others are limited to particular species. Also, the base pair sequence for individual genes may be highly conserved (i.e., show limited sequence differences among different organisms) or be widely variable. As discussed in Chapter 8, these similarities and differences in gene content and sequences are the basis for the development of molecular tests used to detect, identify, and characterize clinically relevant microorganisms.
Replication and Expression of Genetic Information
Replication
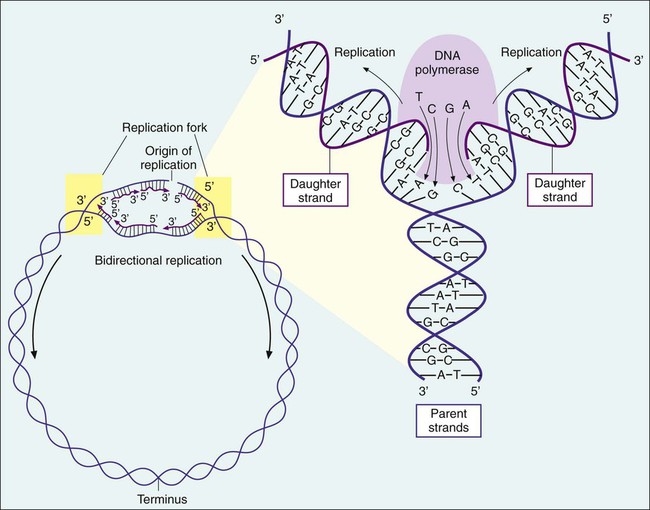
Bacteria multiply by cell division, resulting in the production of two daughter cells from one parent cell. As part of this process, the genome must be replicated so that each daughter cell receives an identical copy of functional DNA. Replication is a complex process mediated by various enzymes, such as DNA polymerase and cofactors; replication must occur quickly and accurately. For descriptive purposes, replication may be considered in four stages (Figure 2-4):
1. Unwinding or relaxation of the chromosome’s supercoiled DNA
2. Separation of the complementary strands of the parental DNA so that each may serve as a template (i.e., pattern) for synthesis of new DNA strands
3. Synthesis of the new (i.e., daughter) DNA strands
4. Termination of replication, releasing two identical chromosomes, one for each daughter cell
Expression of Genetic Information
Transcription.
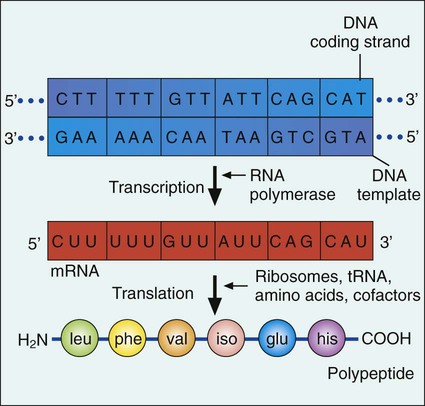
Gene expression begins with transcription. During transcription the DNA base sequence of the gene (i.e., the genetic code) is converted into an mRNA molecule that is complementary to the gene’s DNA sequence (Figure 2-5). Usually only one of the two DNA strands (the sense strand) encodes for a functional gene product. This same strand is the template for mRNA synthesis.
Translation.
The next phase in gene expression, translation, involves protein synthesis. Through this process the genetic code in mRNA molecules is translated into specific amino acid sequences that are responsible for protein structure and function (see Figure 2-5).
Before addressing the process of translation, a discussion of the genetic code that is originally transcribed from DNA to mRNA and then translated from mRNA to protein is warranted. The code consists of triplets of nucleotide bases, referred to as codons; each codon encodes for a specific amino acid. Because there are 64 different codons for 20 amino acids, an amino acid can be encoded by more than one codon (Table 2-1). Each codon is specific for a single amino acid. Therefore, through translation, the codon sequences in mRNA direct which amino acids are added and in what order. Translation ensures that proteins with proper structure and function are produced. Errors in the process can result in aberrant proteins that are nonfunctional, underscoring the need for translation to be well controlled and accurate.
TABLE 2-1
The Genetic Code as Expressed by Triplet-Base Sequences of mRNA*
| Codon | Amino Acid | Codon | Amino Acid | Codon | Amino Acid | Codon | Amino Acid |
| UUU | Phenylalanine | CUU | Leucine | GUU | Valine | AUU | Isoleucine |
| UUC | Phenylalanine | CUC | Leucine | GUC | Valine | AUC | Isoleucine |
| UUG | Leucine | CUG | Leucine | GUG | Valine | AUG (start)† | Methionine |
| UUA | Leucine | CUA | Leucine | GUA | Valine | AUA | Isoleucine |
| UCU | Serine | CCU | Proline | GCU | Alanine | ACU | Threonine |
| UCC | Serine | CCC | Proline | GCC | Alanine | ACC | Threonine |
| UCG | Serine | CCG | Proline | GCG | Alanine | ACG | Threonine |
| UCA | Serine | CCA | Proline | GCA | Alanine | ACA | Threonine |
| UGU | Cysteine | CGU | Arginine | GGU | Glycine | AGU | Serine |
| UGC | Cysteine | CGC | Arginine | GGC | Glycine | AGC | Serine |
| UGG | Tryptophan | CGG | Arginine | GGG | Glycine | AGG | Arginine |
| UGA | None (stop signal) | CGA | Arginine | GGA | Glycine | AGA | Arginine |
| UAU | Tyrosine | CAU | Histidine | GAU | Aspartic | AAU | Asparagine |
| UAC | Tyrosine | CAC | Histidine | GAC | Aspartic | AAC | Asparagine |
| UAG | None (stop signal) | CAG | Glutamine | GAG | Glutamic | AAG | Lysine |
| UAA | None (stop signal) | CAA | Glutamine | GAA | Glutamic | AAA | Lysine |
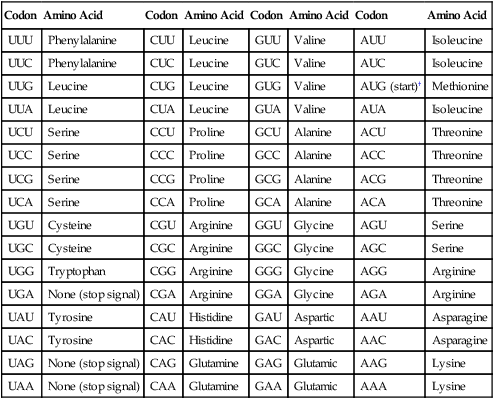
*The codons in DNA are complementary to those given here. Thus, U is complementary to the A in DNA, C is complementary to G, G to C, and A to T. The nucleotide on the left is at the 5’-end of the triplet.
†AUG codes for N-formylmethionine at the beginning of messenger ribonucleic acid (mRNA) in bacteria.
Modified from Brock TD et al, editors: Biology of microorganisms, Upper Saddle River, NJ, 2009, Prentice Hall.
To accomplish the task of translation, intricate interactions between mRNA, tRNA, and rRNA are required. Sixty different types of tRNA molecules are responsible for transferring different amino acids from intracellular reservoirs to the site of protein synthesis. These molecules, which have a structure that resembles an inverted t, contain one sequence recognition site (anticodon) for binding to specific 3-base sequences (codons) on the mRNA molecule (Figure 2-6). A second site binds specific amino acids, the building blocks of proteins. Each amino acid is joined to a specific tRNA molecule through the enzymatic activity of aminoacyl-tRNA synthetases. Therefore, tRNA molecules have the primary function of using the codons of the mRNA molecule as the template for precisely delivering a specific amino acid for polymerization. Ribosomes, which are compact nucleoproteins, are composed of rRNA and proteins. They are central to translation, assisting with coupling of all required components and controlling the translational process.
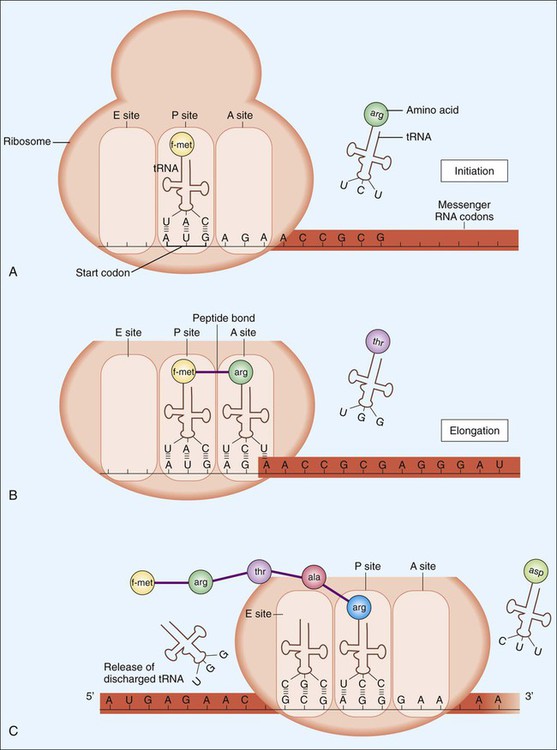
Translation, diagrammatically shown in Figure 2-6, involves three steps: initiation, elongation, and termination. Following termination, bacterial proteins often undergo posttranslational modifications as a final step in protein synthesis.
Initiation begins with the association of ribosomal subunits, mRNA, formylmethionine tRNA ([f-met] carrying the initial amino acid of the protein to be synthesized), and various initiation factors (see Figure 2-6, A). Assembly of the complex begins at a specific 3- to 9-base (Shine-Dalgarno sequence) on the mRNA about 10 bp upstream of the AUG start codon. After the initial complex has been formed, addition of individual amino acids begins.
Elongation involves tRNAs mediating the sequential addition of amino acids in a specific sequence that is dictated by the codon sequence of the mRNA molecule (see Figure 2-6, B and C, and Table 2-1). As the mRNA molecule threads through the ribosome in a 5’ to 3’ direction, peptide bonds are formed between adjacent amino acids, still bound by their respective tRNA molecules in the P (peptide) and A (acceptor) sites of the ribosome. During the process, the forming peptide is moved to the P site, and the most 5’ tRNA is released from the E (exit) site. This movement vacates the A site, which contains the codon specific for the next amino acid, so that the incoming tRNA−amino acid can join the complex (see Figure 2-6, C).
Termination, the final step in translation, occurs when the ribosomal A site encounters a stop or nonsense codon that does not specify an amino acid (i.e., a “stop signal”; see Table 2-1). At this point, the protein synthesis complex disassociates and the ribosomes are available for another round of translation. After termination, most proteins must undergo modification, such as folding or enzymatic trimming, so that protein function, transportation, or incorporation into various cellular structures can be accomplished. This process is referred to as posttranslational modification.
Regulation and Control of Gene Expression
In general, genes that encode anabolic enzymes for the synthesis of particular products are repressed (i.e., are not transcribed and therefore are not expressed) in the presence of the gene end product. This strategy prevents waste and overproduction of products that are already present in sufficient supply. In this system, the product acts as a co-repressor that forms a complex with a repressor molecule. In the absence of co-repressor product (i.e., gene product), transcription occurs (Figure 2-7, A). When present in sufficient quantity, the product forms a complex with the repressor. The complex then binds to a specific base region of the gene sequence known as the operator region (Figure 2-7, B). This binding blocks RNA polymerase progression from the promoter sequence and inhibits transcription. As the supply of product (co-repressor) dwindles, an insufficient amount remains to form a complex with the repressor. The operator region is no longer bound to the repressor molecule. Transcription of the genes for the anabolic enzymes commences and continues until a sufficient supply of end product is again available.
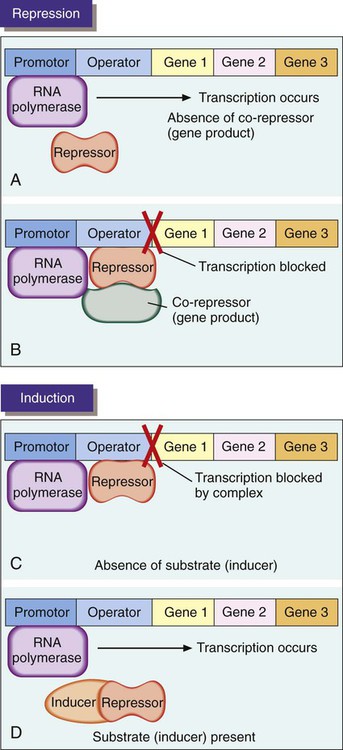
In contrast to repression, genes that encode catabolic enzymes are usually induced; that is, transcription occurs only when the substrate to be degraded by enzymatic action is present. Production of degradative enzymes in the absence of substrates would be a waste of cellular energy and resources. When the substrate is absent in an inducible system, a repressor binds to the operator sequence of the DNA and blocks transcription of the genes for the degradative enzymes (Figure 2-7, C). In the presence of an inducer, which often is the target substrate for degradation, a complex is formed between inducer and repressor and results in the release of the repressor from the operator site, allowing transcription of the genes encoding the specific catabolic enzymes (Figure 2-7, D).
Genetic Exchange and Diversity
Genetic Recombination
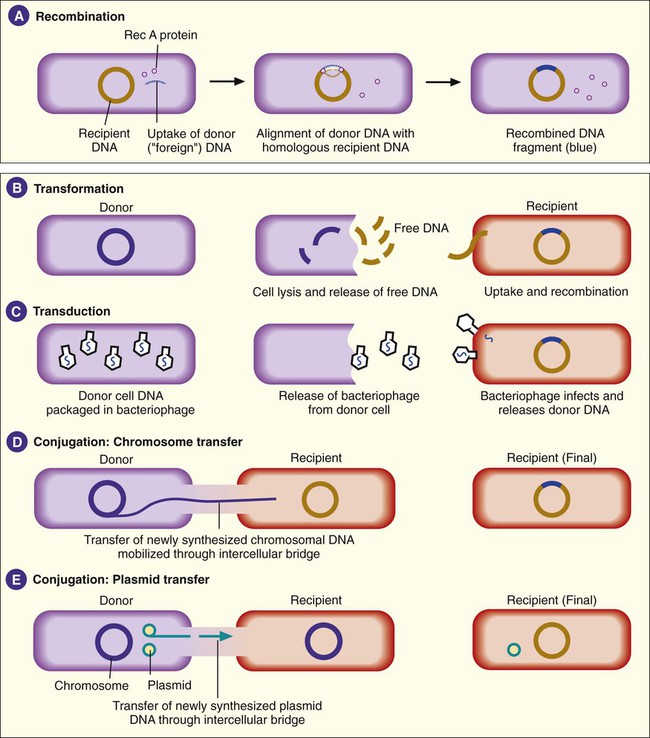
Besides mutations, bacterial genotypes can be altered through recombination. In this process, some segment of DNA originating from one bacterial cell (i.e., donor) enters a second bacterial cell (i.e., recipient) and is exchanged with a DNA segment of the recipient’s genome. This is also referred to as homologous recombination, because the pieces of DNA that are exchanged usually have extensive homology or similarities in their nucleotide sequences. Recombination involves a number of binding proteins, with the RecA protein playing a central role (Figure 2-8, A). After recombination, the recipient DNA consists of one original, unchanged strand and a second strand from the donor DNA fragment that has been recombined.
Genetic Exchange
Transformation.
Transformation involves recipient cell uptake of naked (free) DNA released into the environment when another bacterial cell (i.e., donor) dies and undergoes lysis (see Figure 2-8, B). This genomic DNA exists as fragments in the environment. Certain bacteria are able to take up naked DNA from their surroundings; that is, they are able to undergo transformation. Such bacteria are said to be competent. Among the bacteria that cause human infections, competence is a characteristic commonly associated with members of the genera Haemophilus, Streptococcus, and Neisseria.
Transduction.
Transduction is a second mechanism by which DNA from two bacteria may come together in one cell, thus allowing for recombination (see Figure 2-8, C). This process is mediated through viruses capable of infecting bacteria (i.e., bacteriophages). In their “life cycle,” these viruses integrate their DNA into the bacterial cell’s chromosome, where viral DNA replication and expression occur. When the production of viral products is complete, viral DNA is excised (cut) from the bacterial chromosome and packaged within a protein coat. This virion contains bacterial and viral DNA. The newly formed recombinant virion, along with the additional multiple virions (virus particles), is released when the infected bacterial cell lyses. In transduction, the recombinant virion incorporates its own DNA but may also pick up a portion of the donor bacterium’s DNA.
Conjugation.
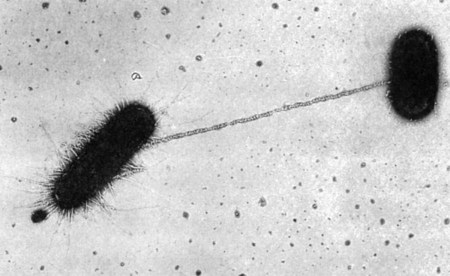
The third mechanism of DNA exchange between bacteria is conjugation. This process occurs between two living cells, involves cell-to-cell contact, and requires mobilization of the donor bacterium’s chromosome. The nature of intercellular contact is not well characterized in all bacterial species capable of conjugation. However, in E. coli, contact is mediated by a sex pilus (Figure 2-9). The sex pilus originates from the donor and establishes a conjugative bridge that serves as the conduit for DNA transfer from donor to recipient cell. With intercellular contact established, chromosomal mobilization is undertaken and involves DNA synthesis. One new DNA strand is produced by the donor and is passed to the recipient (see Figure 2-8, D). The amount of DNA transferred depends on how long the cells are able to maintain contact, but usually only portions of the donor molecule are transferred. In any case, the newly introduced DNA is then available to recombine with the recipient’s genome.
In addition to chromosomal DNA, genes encoded in extrachromosomal genetic elements, such as plasmids and transposons, may be transferred by conjugation (see Figure 2-8, E). Not all plasmids are capable of conjugative transfer, but for those that are, the donor plasmid usually is replicated so that the donor retains a copy of the plasmid transferred to the recipient. (See the discussion of the F plasmid in the section Cellular Appendages, later in the chapter.) Plasmid DNA may also become incorporated into the host cell’s chromosome.
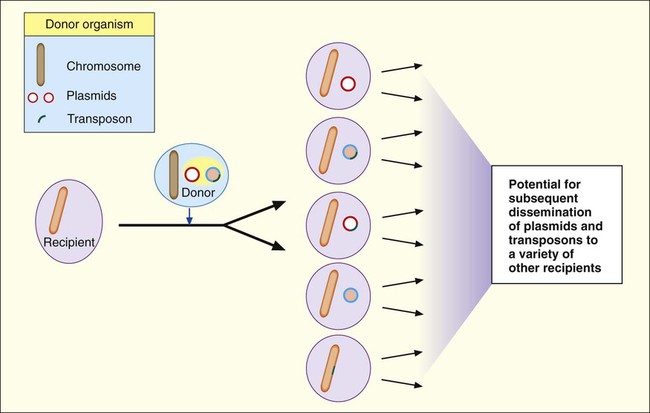
Plasmids and transposons play a key role in genetic diversity and dissemination of genetic information among bacteria. Many characteristics that significantly alter the activities of clinically relevant bacteria are encoded and disseminated on these elements. Furthermore, as shown in Figure 2-10, the variety of strategies that bacteria can use to mix and match genetic elements provides them with a tremendous capacity to genetically adapt to environmental changes, including those imposed by human medical practices. A good example of this is the emergence and widespread dissemination of resistance to antimicrobial agents among clinically important bacteria. Bacteria have used their capacity for disseminating genetic information to establish resistance to many of the commonly prescribed antibiotics. (See Chapter 11 for more information about antimicrobial resistance mechanisms.)
Bacterial Metabolism
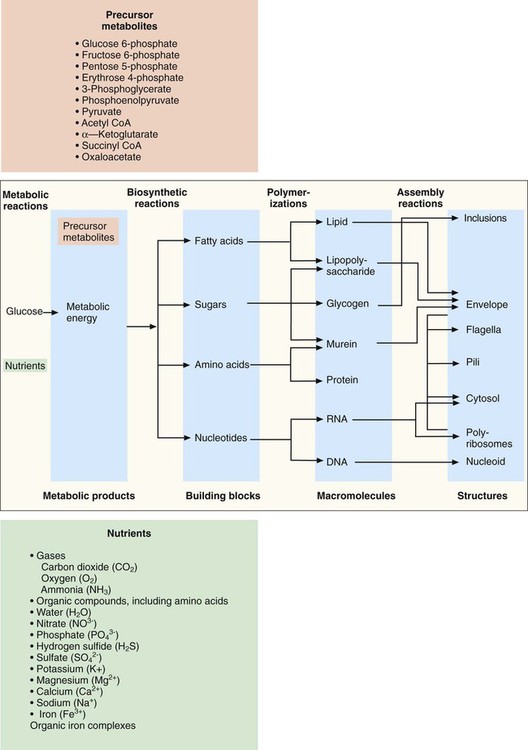
For the sake of clarity, metabolism is discussed in terms of four primary, but interdependent, processes: fueling, biosynthesis, polymerization, and assembly (Figure 2-11).
Fueling
Production of Precursor Metabolites
Once inside the cell, many nutrients serve as the raw materials from which precursor metabolites for subsequent biosynthetic processes are produced. These metabolites, listed in Figure 2-11, are produced through three central pathways; the Embden-Meyerhof-Parnas (EMP) pathway, the tricarboxylic acid (TCA) cycle, and the pentose phosphate shunt. These pathways and their relationship to one another are shown in Figure 2-12; not shown are the several alternative pathways (e.g., the Entner-Douder off pathway) that play key roles in redirecting and replenishing the precursors as they are used in subsequent processes.
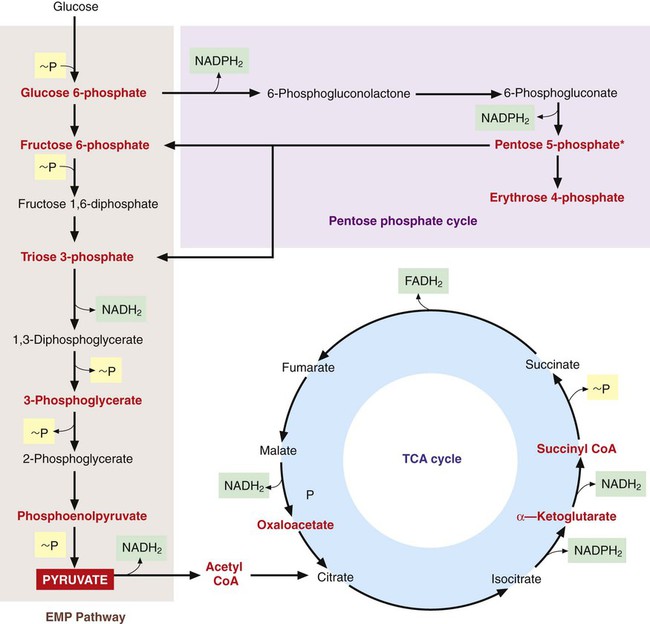
Energy Production
The two general mechanisms for ATP production in bacterial cells are substrate-level phosphorylation and electron transport, also referred to as oxidative phosphorylation. In substrate-level phosphorylation, high-energy phosphate bonds produced by the central pathways are donated to adenosine diphosphate (ADP) to form ATP (see Figure 2-12). Additionally, pyruvate, a primary intermediate in the central pathways, serves as the initial substrate for several other pathways to generate ATP by substrate level phosphorylation. These other pathways constitute fermentative metabolism, which does not require oxygen and produces various end products, including alcohols, acids, carbon dioxide, and hydrogen. The specific fermentative pathways and the end products produced vary with different bacterial species. Detection of these products is an important basis for laboratory identification of bacteria. (See Chapter 7 for more information on the biochemical basis for bacterial identification.)
Oxidative Phosphorylation.
Oxidative phosphorylation involves an electron transport system that conducts a series of electron transfers from reduced carrier molecules such as NADH2 and NADPH2, produced in the central pathways (see Figure 2-12), to a terminal electron acceptor. The energy produced by the series of oxidation-reduction reactions is used to generate ATP from ADP. When oxidative phosphorylation uses oxygen as the terminal electron acceptor, the process is known as aerobic respiration. Anaerobic respiration refers to processes that use final electron acceptors other than oxygen.
Biosynthesis
The fueling reactions essentially bring together all the raw materials needed to initiate and maintain all other cellular processes. The production of precursors and energy is accomplished through catabolic processes and the degradation of substrate molecules. The three remaining pathways for biosynthesis, polymerization, and assembly depend on anabolic metabolism. In anabolic metabolism, precursor compounds are joined for the creation of larger molecules (polymers) required for assembly of cellular structures (see Figure 2-11).
Biosynthetic processes use the precursor products in dozens of pathways to produce a variety of building blocks, such as amino acids, fatty acids, sugars, and nucleotides (see Figure 2-11). Many of these pathways are highly complex and interdependent, whereas other pathway are completely independent. In many cases, the enzymes that drive the individual pathways are encoded on a single mRNA molecule that has been transcribed from contiguous genes in the bacterial chromosome (i.e., an operon).
Structure and Function of the Bacterial Cell
Eukaryotic and Prokaryotic Cells
• Endoplasmic reticulum—process and transport proteins
• Golgi body—modification of substances and transport throughout the cell, including internal delivery of molecules and exocytosis or secretion of other molecules
• Mitochondria—generate energy (ATP)
• Lysosomes—provide environment for controlled enzymatic degradation of intracellular substances
Bacterial Morphology
Most clinically relevant bacterial species range in size from 0.25 to 1 µm in width and 1 to 3 µm in length, thus requiring microscopy for visualization (see Chapter 6 for more information on microscopy). Just as bacterial species and genera vary in their metabolic processes, their cells also vary in size, morphology, and cell-to-cell arrangements and in the chemical composition and structure of the cell wall. The bacterial cell wall differences provide the basis for the Gram stain, a fundamental staining technique used in bacterial identification schemes. This staining procedure separates almost all medically relevant bacteria into two general types: gram-positive bacteria, which stain a deep blue or purple, and gram-negative bacteria, which stain a pink to red (see Figure 6-3). This simple but important color distinction is due to differences in the constituents of bacterial cell walls that influence the cell’s ability to retain differential dyes following treatment with a decolorizing agent.
Common bacterial cellular morphologies include cocci (circular), coccobacilli (ovoid), and bacillus (rod shaped), as well as fusiform (pointed end), curved, or spiral shapes. Cellular arrangements are also noteworthy. Cells may characteristically occur singly, in pairs, or grouped as tetrads, clusters, or in chains (see Figure 6-4 for examples of bacterial staining and morphologies). The determination of the Gram stain reaction and the cell size, morphology, and arrangement are essential aspects of bacterial identification.
Bacterial Cell Components
Cell Envelope
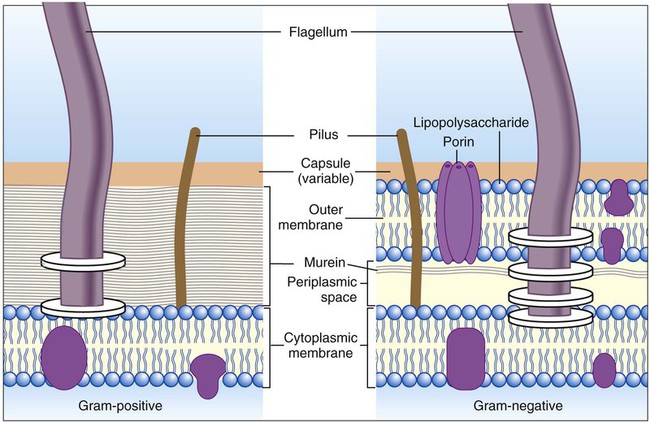
As shown in Figure 2-13, the outermost structure, the cell envelope, comprises:
• An outer membrane (in gram-negative bacteria only)
• A cell wall composed of the peptidoglycan macromolecule (also known as the murein layer)
• Periplasm (in gram-negative bacteria only)
• The cytoplasmic or cell membrane, which encloses the cytoplasm
Cell Wall (Murein Layer).
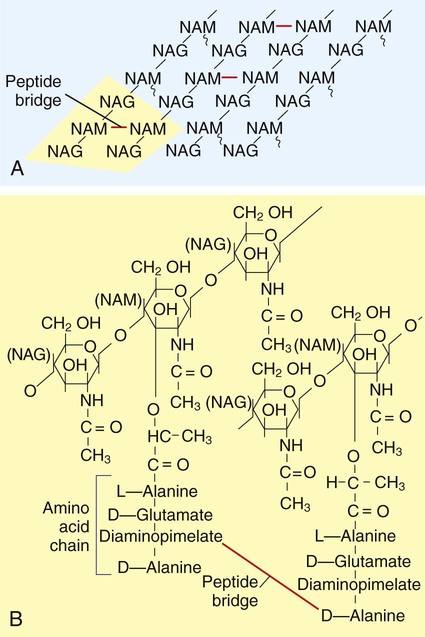
The structure of the cell wall is unique and is composed of disaccharide-pentapeptide subunits. The disaccharides N-acetylglucosamine and N-acetylmuramic acid are the alternating sugar components (moieties), with the amino acid chain linked to N-acetylmuramic acid molecules (Figure 2-14). Polymers of these subunits cross-link to one another by means of peptide bridges to form peptidoglycan sheets. In turn, layers of these sheets are cross-linked with one another, forming a multilayered, cross-linked structure of considerable strength. Referred to as the murein sacculus, or sack, this peptidoglycan structure surrounds the entire cell.
A notable difference between the cell walls of gram-positive and gram-negative bacteria is the substantially thicker peptidoglycan layer in gram-positive bacteria (see Figure 2-13). Additionally, the cell wall of gram-positive bacteria contains teichoic acids (i.e., glycerol or ribitol phosphate polymers combined with various sugars, amino acids, and amino sugars). Some teichoic acids are linked to N-acetylmuramic acid, and others (e.g., lipoteichoic acids) are linked to the next underlying layer, the cellular membrane. Other gram-positive bacteria (e.g., mycobacteria) have waxy substances within the murein layer, such as mycolic acids. Mycolic acids make the cells more refractory to toxic substances, including acids. Bacteria with mycolic acid in the cell walls require unique staining procedures and growth media in the diagnostic laboratory.
Cytoplasmic (Inner) Membrane.
• Transport of solutes into and out of the cell
• Housing of enzymes involved in outer membrane synthesis, cell wall synthesis, and the assembly and secretion of extracytoplasmic and extracellular substances
• Generation of chemical energy (i.e., ATP)
• Mediation of chromosomal segregation during replication
• Housing of molecular sensors that monitor chemical and physical changes in the environment
Cellular Appendages.
The capsule is immediately exterior to the murein layer of gram-positive bacteria and the outer membrane of gram-negative bacteria. Often referred to as the “slime layer,” the capsule is composed of high-molecular-weight polysaccharides, the production of which may depend on the environment and growth conditions surrounding the bacterial cell. The capsule does not function as an effective permeability barrier or add strength to the cell envelope, but it does protect bacteria from attack by components of the human immune system. The capsule also facilitates and maintains bacterial colonization of biologic (e.g., teeth) and inanimate (e.g., prosthetic heart valves) surfaces through the formation of biofilms. A biofilm consists of a monomicrobic or polymicrobic group of bacteria housed in a complex polysaccharide matrix. (See Chapter 3 for further discussion of microbial biofilms.)